Clear Sky Science · en
NatA engages in multi-factor complexes at the ribosomal polypeptide tunnel exit
How cells fine-tune new proteins as they are born
Every second, your cells manufacture thousands of new proteins on tiny machines called ribosomes. As each protein chain emerges, it must be trimmed, tagged and folded correctly, or it can misbehave and contribute to disease. This study looks at one of the key tagging systems, a modification called N-terminal acetylation, and shows how a central enzyme complex, NatA, works together with several partners right where new proteins exit the ribosome. Understanding this choreography helps explain how cells keep their protein factories running smoothly.
The tiny chemical tag that matters
Most proteins start life with the same first building block, the amino acid methionine. Often, this initial methionine is removed and replaced with a small chemical tag called an acetyl group. This tag, added to the protein’s very tip (its N-terminus), can influence how long the protein survives, where it goes in the cell and how it functions. Two main types of enzymes sit at the ribosome’s exit tunnel to manage this first makeover: methionine aminopeptidases (MAPs), which cut off the starting methionine, and N-terminal acetyltransferases (NATs), which add the acetyl tag. Among the NATs, NatA is the workhorse in human cells, potentially modifying almost 40% of all proteins. Because many proteins are made at once, the cell must organize these enzymes so that trimming and tagging happen quickly and in the right order.
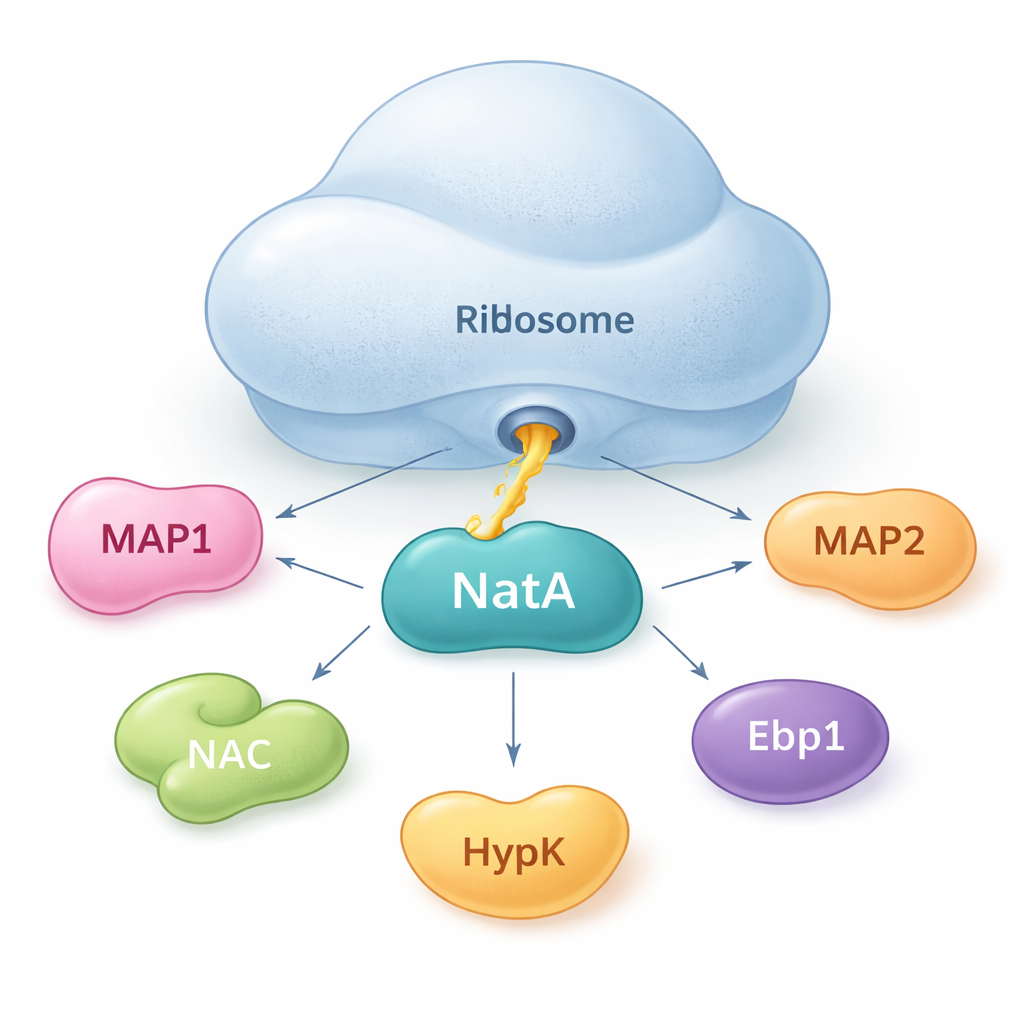
NatA as a meeting point for protein helpers
The authors show that NatA does not act alone. Instead, it behaves like a hub that brings together multiple helper proteins, both on and off the ribosome. Using sensitive binding measurements in solution, they found that NatA can form tight complexes with MAP1, the nascent polypeptide–associated complex (NAC), the regulatory protein HypK, and another enzyme, Naa50, even without the ribosome present. NAC can bridge NatA to MAP1, allowing both the cutting and tagging steps to be linked in a single assembly. However, HypK, which normally dampens NatA’s activity, can block NAC from binding. This suggests that cells may switch NatA between a more active, NAC-linked state and a restrained, HypK-bound state, depending on their needs.
A second docking spot right at the tunnel exit
By using cryo–electron microscopy, a technique that images large molecules in near-atomic detail, the researchers discovered that NatA can bind to the ribosome in two distinct positions. One site, previously known, sits a bit away from the tunnel exit. The newly discovered “proximal” site places NatA’s catalytic core very close to where the growing protein chain emerges, shortening the distance the chain must travel to be modified. Intriguingly, both positions can be occupied at the same time, meaning two NatA complexes can sit on the same ribosome. The more distant copy acts as a scaffold and anchor, while the proximal copy is optimally placed to tag new proteins. The two NatA molecules also touch each other, indicating that NatA can coordinate its own duplicates into multi-enzyme teams.
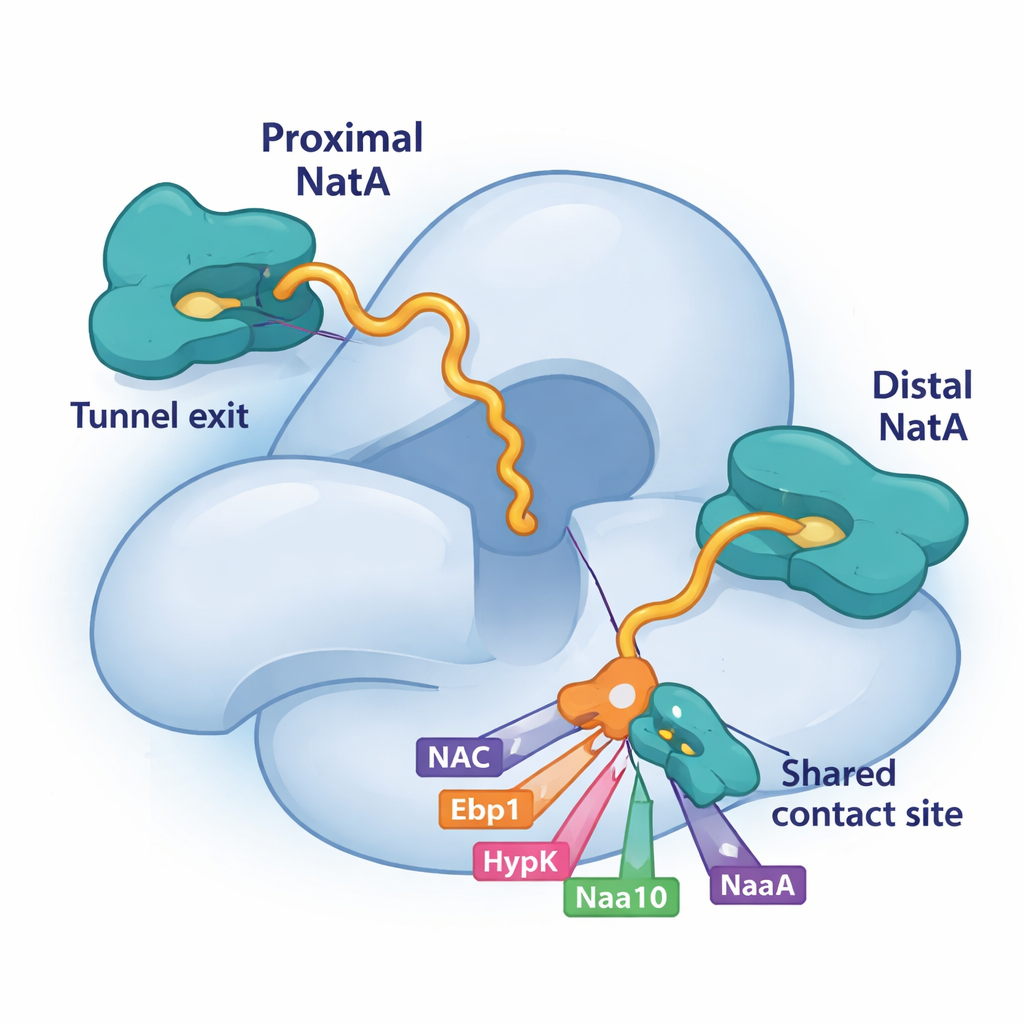
Competition for a shared landing pad on NatA
A key piece of this puzzle is a small groove on the large NatA subunit, Naa15. The study shows that four different ribosome-associated factors—NAC, HypK, Ebp1 and even a tail segment of NatA’s own catalytic subunit, Naa10—use short helices to latch onto exactly the same patch on Naa15. Because they all rely on this shared landing pad, they must compete for access. When HypK occupies the site, it not only silences NatA’s activity but also prevents NatA from pairing with NAC or forming NatA–NatA dimers. When NAC, the pseudo-enzyme Ebp1, or proximal Naa10 bind there instead, they can remodel NatA’s position or connections at the ribosome. This competitive binding gives the cell a flexible way to choose which partners assemble around NatA at any moment.
A pseudo-enzyme as a traffic controller
The protein Ebp1 is especially intriguing. It resembles one of the methionine-removing enzymes but lacks catalytic activity; it is a “pseudo-enzyme.” Earlier work showed that Ebp1 can sit on the ribosome near the tunnel exit and clamp down a long RNA segment. In this study, the authors find that when NatA joins the ribosome, Ebp1 shifts position, lets go of the RNA, and instead uses its own helix to occupy the same binding site on Naa15 that NAC and HypK use. This rearrangement suggests that Ebp1 can act as a placeholder or organizer: it can shape the local RNA environment, then hand off control to active enzymes like MAPs and NatA, without directly performing chemistry itself.
Why this molecular crowd control matters
To a non-specialist, the details of helices and binding sites may sound abstract, but the message is simple: cells run a highly choreographed, multi-step quality-control system right where proteins are born. NatA sits at the center of this system, able to recruit or release different partners depending on which factors win access to a crucial docking patch and which ribosomal site NatA occupies. This flexible network helps ensure that trimming and acetylation occur with the right timing and order for thousands of different proteins. Because errors in N-terminal processing are linked to developmental disorders, neurodegeneration and cancer, mapping out how NatA and its partners assemble gives researchers a clearer blueprint for how early protein processing is controlled—and how it might be targeted when it goes awry.
Citation: Klein, M., Wild, K., McTiernan, N. et al. NatA engages in multi-factor complexes at the ribosomal polypeptide tunnel exit. Nat Commun 17, 884 (2026). https://doi.org/10.1038/s41467-026-68787-5
Keywords: protein quality control, N-terminal acetylation, ribosome tunnel exit, NatA complex, co-translational modification