Clear Sky Science · en
Single-cell exon deletion profiling reveals splicing events that shape gene expression and cell state dynamics
How tiny edits to RNA can change cell behavior
Our cells constantly read and splice RNA instructions to decide which proteins to make and how to respond to stress, grow, or divide. This study shows that changing very small pieces of those RNA messages—single segments called exons—can strongly reshape gene activity and even shift how cells move through the cell cycle. The work introduces a powerful tool that lets scientists scan many of these tiny RNA choices at once, cell by cell, opening new ways to understand disease and discover drug targets.
Cutting out selected words from the genetic script
Genes are written as long stretches of DNA, but cells do not read them straight through. Instead, they cut and paste together smaller blocks, called exons, to build an RNA message. By choosing which exons to keep, cells can make multiple protein versions from a single gene, much like producing different edits of a film from the same raw footage. Many diseases, including cancer and autism, are linked to mistakes in this process, yet for most exon choices we still do not know what they actually do. The authors set out to change that by building a large-scale system that can remove specific exons from many genes and then watch, in thousands of individual cells at once, how those edits affect the cell’s internal wiring.
A single-cell "search engine" for exon function
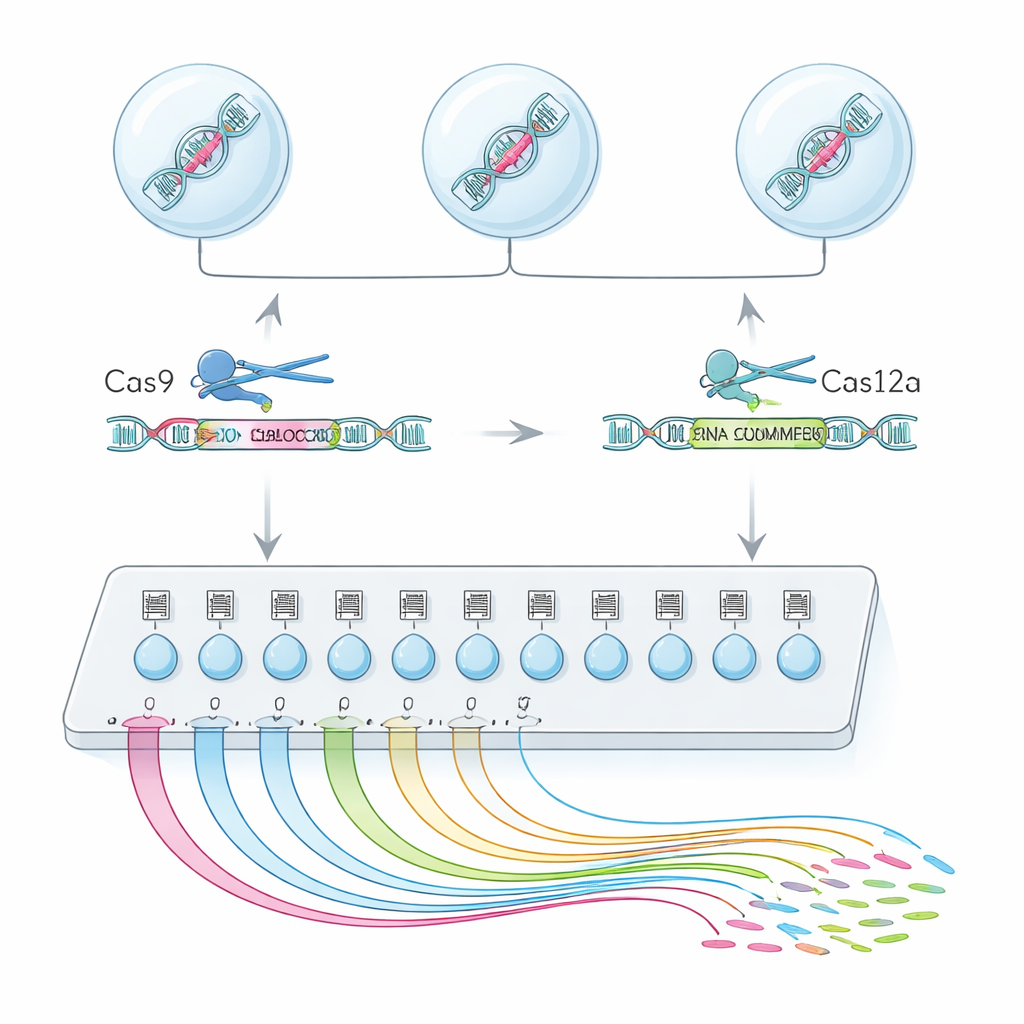
To achieve this, the team combined two genome-editing enzymes, Cas9 and Cas12a, into a hybrid platform called CHyMErA. Each enzyme is guided to DNA by short RNA “addresses.” By directing the two enzymes to cut just before and just after a chosen exon, they can neatly delete that segment while leaving the rest of the gene intact. The new method, scCHyMErA-Seq, links this precise cutting to single-cell RNA sequencing technology. Each cell’s RNA is captured in a droplet together with a barcode and the guide RNAs that specify which exon was removed. Sequencing then reveals, for tens of thousands of cells, both the edit each cell received and the detailed pattern of genes it turned on or off.
Engineering the tool to see both blades of the scissors
A key challenge was reliably detecting the Cas12a guides alongside Cas9 in the same single-cell experiment. Initial designs either failed to capture the Cas12a guide RNAs or damaged the editing efficiency. The researchers solved this by carefully redesigning the Cas12a handle sequence to remove stretches that prematurely halted transcription, and by adding a small stabilizing RNA element and a targeted amplification step. These adjustments boosted the ability to see both guides in about 90 percent of cells, while preserving strong exon deletion. With this optimized setup, the authors screened 224 alternative exons across 161 genes in human cells, profiling more than 200,000 high-quality single cells.
Discovering exons that control gene programs and cell cycles
When the team compared the gene activity of edited cells to controls, nearly half of the tested exons caused substantial shifts in hundreds of other genes. Exons from genes involved in RNA processing and transcription tended to cluster together, producing similar expression fingerprints and revealing shared biological pathways, such as ribosome building or RNA decay. In some cases, deleting a single exon had a clear and interpretable effect: removing an exon in the TAF5 or LSM11 genes disrupted proper processing of histone RNAs, leading to an unusual build-up of polyadenylated histone messages. The dataset also highlighted dozens of exons whose loss altered how many cells were found in different stages of the cell cycle, tying specific splicing events directly to decisions about when a cell pauses, copies its DNA, or prepares to divide.
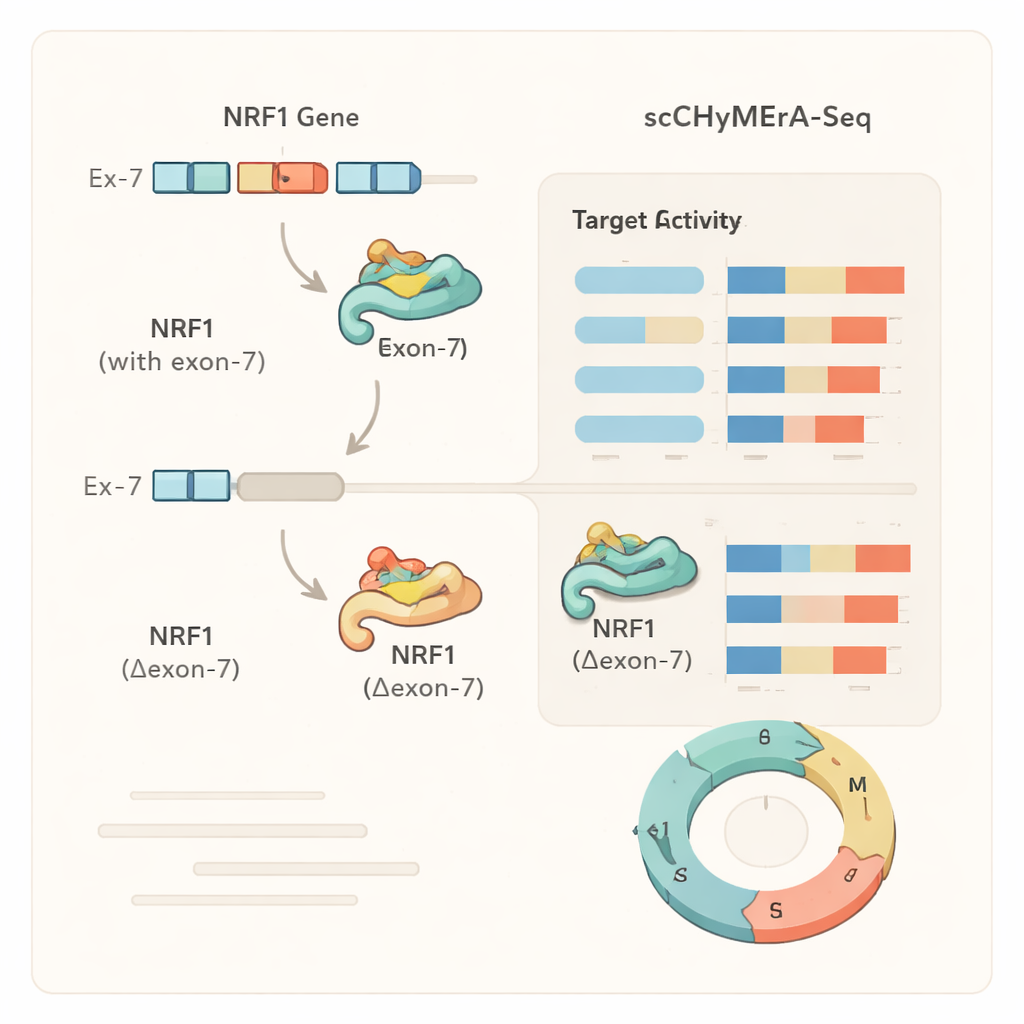
A case study: one exon that tunes a master regulator
One striking example was exon 7 in the gene NRF1, a transcription factor that controls many genes involved in cellular energy production. This exon partly overlaps NRF1’s DNA-binding region. When exon 7 was deleted, hundreds of NRF1-controlled genes changed their activity, and detailed experiments showed that the shortened NRF1 protein was much less able to bind its target promoters in the genome. The authors also identified a specific splicing regulator, SRSF3, that promotes inclusion of this exon, revealing a chain from a splicing factor, through an exon choice in NRF1, to broad changes in cellular gene programs and metabolism. Similar analyses showed that for some genes, exon deletion mimicked a full gene knockout, while for others it produced a more fine-tuned, state-dependent change, suggesting that alternative exons can subtly modulate protein behavior rather than simply turning it on or off.
Why this matters for health and future therapies
To a non-specialist, the key message is that cells rely on the inclusion or skipping of very short RNA segments to dial in how genes work, and that these fine adjustments can shift major traits such as how cells divide or respond to stress. The scCHyMErA-Seq platform acts like a high-throughput lab test for these splicing choices, pinpointing which exons have real functional impact and how they reshape cellular states. Because many cancers and neurological disorders involve mis-spliced exons or altered transcription factors, this approach could help prioritize which splice variants to target with future drugs or RNA therapies, and provide a roadmap for understanding how small edits in the genetic script ripple out to large changes in cell behavior.
Citation: Kumari, B., Damodaran, A.P., Guiblet, W.M. et al. Single-cell exon deletion profiling reveals splicing events that shape gene expression and cell state dynamics. Nat Commun 17, 1218 (2026). https://doi.org/10.1038/s41467-026-68774-w
Keywords: alternative splicing, single-cell RNA sequencing, CRISPR screening, gene regulation, cell cycle