Clear Sky Science · en
Cycling molecular assemblies for Golgi imaging and disruption
Turning the Cell’s Shipping Center into a Strategic Target
Inside every animal cell sits the Golgi apparatus, a busy sorting station that finishes, packages, and ships thousands of proteins and fats. Many cancer cells depend heavily on this hub to modify growth signals and secrete factors that shape their surroundings. This paper introduces tiny designer molecules that home in on the Golgi, light it up for rapid imaging, and, when re‑engineered, can selectively jam its operations to cripple cancer cells while sparing certain healthy cells.
A New Way to Find and Watch the Golgi
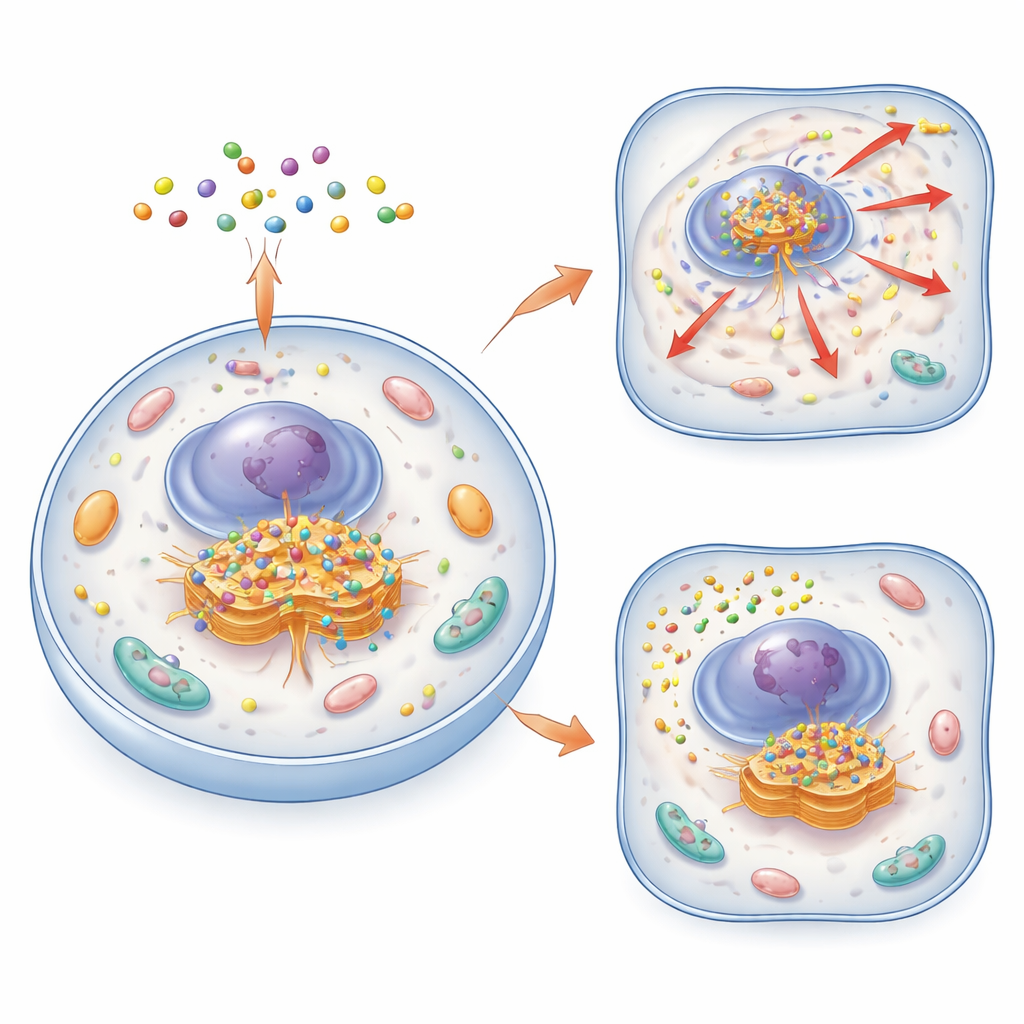
The authors created a family of small, smart molecules called cycling molecular assemblies, or CyMA. These molecules are built from short peptides that can both slip through cell membranes and stick together into tiny clusters. In their “imaging” version (CyMA‑i), the peptides carry a fluorescent dye that glows brightly when the molecules assemble. Once inside a cell, enzymes snip a protective cap from the CyMA precursor, exposing a reactive handle. Golgi‑resident enzymes then attach a fatty chain to this handle, making the molecules far more sticky and prone to self‑assemble exactly at the Golgi. Because this process is rapid and uses natural cellular machinery, the researchers can outline the shape of the Golgi in living cells within minutes and at very low probe concentrations.
A Futile Cycle that Pins Molecules to the Golgi
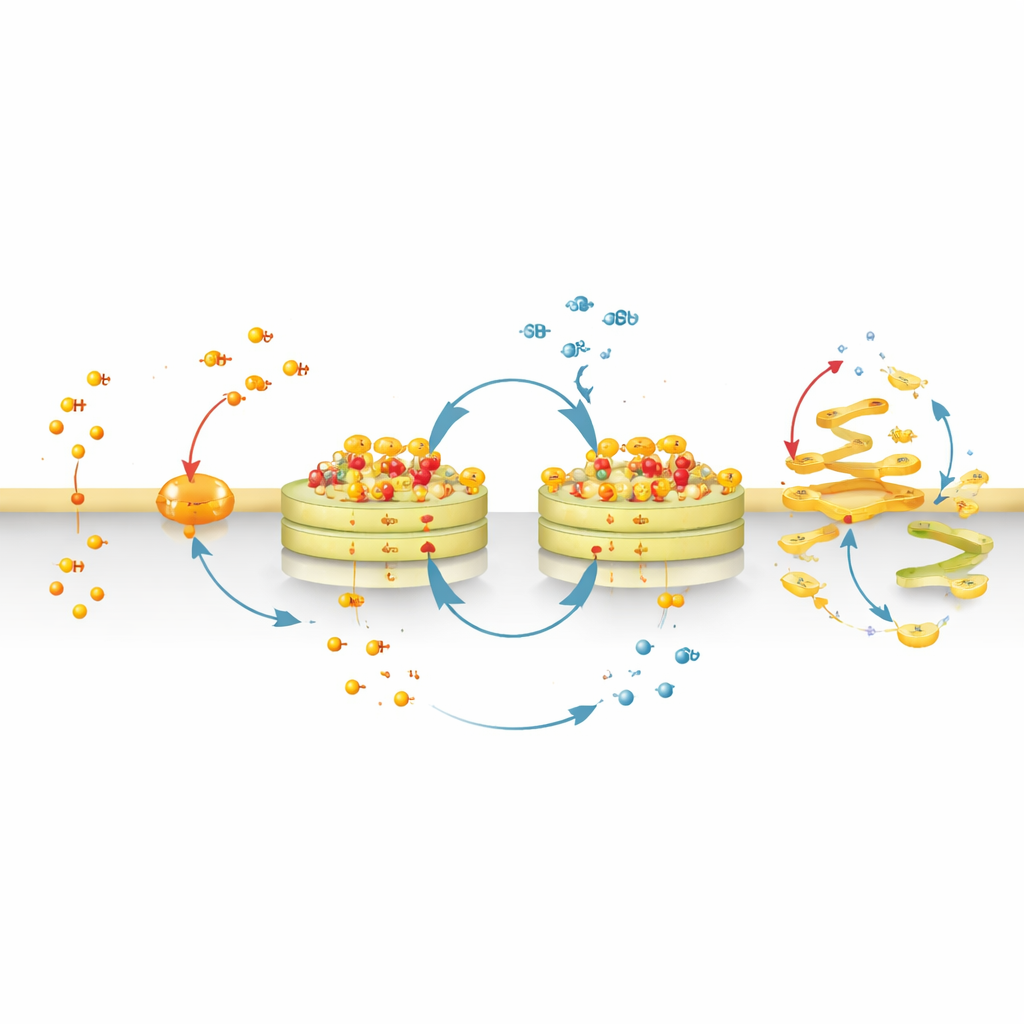
CyMA are not simply labels; they are designed to participate in a “futile cycle” at the Golgi. One group of enzymes attaches fatty chains to the peptide, while another group removes them. As the cell repeatedly adds and removes these chains, the modified peptides keep assembling and disassembling in place. This constant back‑and‑forth traps a solid‑like core of assemblies on Golgi membranes, yet keeps individual molecules cycling through. The cell must continually spend its own fatty‑acid building blocks to maintain this loop, effectively feeding the assemblies and helping them persist. At the same time, additional enzymes can slowly cut a distinct ester bond within CyMA, turning them into more water‑loving fragments that drift away. Because some healthy cells, such as liver cells and certain immune cells, are rich in such ester‑cleaving enzymes, they are naturally better at dismantling CyMA and are therefore less affected.
From Gentle Imaging Tool to Golgi Disruptor
By swapping the fluorescent dye for a more strongly membrane‑seeking chemical group, the team converted CyMA‑i into CyMA‑d, a disruptive version that no longer glows but still cycles and assembles at the Golgi. These assemblies physically disturb the structure of the Golgi and the constant flow of vesicles to and from it. Proteins that normally travel from the endoplasmic reticulum to the Golgi and then out to the cell surface become stuck or misrouted. Incoming routes—from the cell surface back to the Golgi or from the Golgi to other organelles—are also blocked. As a result, key surface receptors and signaling molecules either fail to reach their destinations or accumulate in the wrong places inside the cell.
Scrambling Cell Signals and Secret Messages
The Golgi is where many proteins receive crucial finishing touches, such as sugar chains and lipid tails. CyMA‑d assemblies reduce the proper lipidation and glycosylation of numerous proteins, including well‑known cancer drivers like Ras and major receptor tyrosine kinases. These changes derail growth‑promoting pathways such as AKT and mTOR. At the same time, the cell’s secretory output is dampened: important factors like TGF‑β1 and VEGF, which tumors use to suppress immunity and stimulate blood vessel growth, are no longer efficiently released. The stress of this breakdown triggers tangled endoplasmic reticulum networks, altered mitochondria, stalled autophagy (the cell’s recycling system), and a surge in damaged, ubiquitin‑tagged proteins, together pushing cancer cells toward death.
Selective Pressure on Tumors and Promise for Therapy
Because CyMA‑d depend on a shared set of enzymes and metabolites that many cancer cells overuse, they can kill a wide array of tumor types, including drug‑resistant ovarian cancer models, at very low doses. Cells with high levels of specific esterases, however, dismantle CyMA‑d more readily and are less harmed, offering a built‑in route to selectivity. In three‑dimensional tumor spheroids, patient‑derived cultures, and mouse models, CyMA‑d shrink tumors, cut down metastases, and reduce fluid buildup in the abdomen. When combined with immune checkpoint therapies, they further boost tumor control and survival. In essence, this work shows that building dynamic materials inside the Golgi—rather than targeting a single protein—can shut down multiple cancer‑critical processes at once.
What This Means for Future Treatments
To a non‑specialist, the key idea is that the authors have turned the Golgi from a passive bystander into an active therapeutic target. By recruiting the cell’s own enzymes to assemble and recycle tiny molecules at this central hub, they can either harmlessly highlight it or gradually overwhelm its function. This dual‑use platform suggests a new kind of treatment: instead of designing one drug for one protein, we might engineer small precursors that cells transform into self‑organizing materials, capable of disrupting entire communication and transport networks that cancers rely on to grow and spread.
Citation: Tan, W., Zhang, Q., Liu, Z. et al. Cycling molecular assemblies for Golgi imaging and disruption. Nat Commun 17, 2102 (2026). https://doi.org/10.1038/s41467-026-68768-8
Keywords: Golgi apparatus, molecular assemblies, protein trafficking, cancer therapy, organelle targeting