Clear Sky Science · en
Structural basis for fork reversal and RAD51 regulation by the SCF ubiquitin ligase complex of F-box helicase 1
How cells rescue a stalled DNA copy machine
Every time a cell divides, it must copy billions of DNA letters quickly and accurately. But this copying machinery often runs into trouble—DNA damage, hard-to-copy sequences, or drug treatments can all make the process stall. When that happens, cells rely on emergency repair teams to stabilize and restart the copying forks. This paper reveals, in unprecedented structural detail, how one such team leader—a protein complex built around the helicase FBH1—reshapes stalled DNA and keeps another key repair protein, RAD51, under tight control. Understanding this choreography helps explain how cells avoid dangerous genome breakage and why disruptions in these factors are linked to cancer.
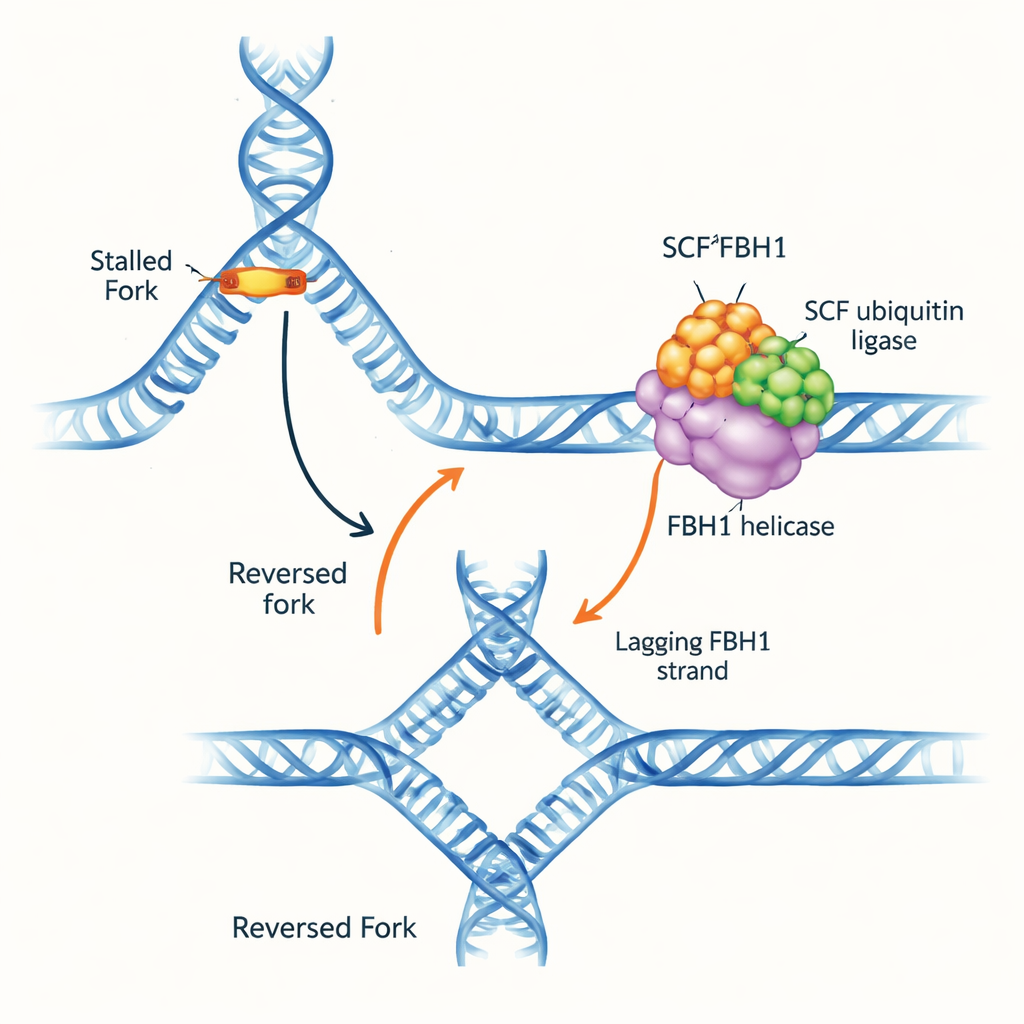
A traffic jam on the DNA highway
DNA replication forks are Y-shaped structures where the double helix is unwound and copied. When the fork encounters obstacles, it can reverse direction: the parental strands re-zip, and the newly made strands pair with each other, forming a four-way DNA junction. This “fork reversal” can buy time to repair damage and restart replication safely, but if mismanaged it also creates fragile DNA that can break. Several molecular motors are known to push forks backward by gripping the double-stranded DNA ahead of the fork. FBH1, however, stood out because it behaves differently and is frequently altered in tumors, suggesting it operates via a distinct and medically important pathway.
A fork specialist with a favorite side
The authors purified the human SCFFBH1 complex, which combines the FBH1 helicase with a ubiquitin-tagging module (SCF). They first asked what DNA shapes FBH1 prefers. Using binding and unwinding assays, they showed that SCFFBH1 strongly favors real fork-like structures, especially where there is a short stretch of single-stranded DNA on the “lagging” template strand—the side copied in small fragments. On these structures, FBH1 unwinds DNA hundreds of times faster than on a simple loose end, indicating that proper engagement at the fork junction supercharges its motor activity. Single-molecule magnetic tweezers confirmed that the complex can unwind and then travel along single-stranded DNA for thousands of bases without falling off, acting as a powerful, processive motor.
Reversing the fork by pulling from behind
To find out how this motor actually reverses forks, the team tested artificial fork substrates with or without single-stranded gaps. Unlike other fork remodelers, SCFFBH1 could not act on a fully paired fork; it required a gap on the lagging template and moved in the 3′–5′ direction along that single strand. This “pulling from behind” motion drives the parental strands to re-anneal while the enzyme stays clamped at the fork junction. A cryo–electron microscopy structure of SCFFBH1 bound to a stalled fork shows how: a special positively charged patch, a “junction binding motif” in one subdomain of FBH1, nestles right at the branch point where the three DNA arms meet. When the researchers mutated two key amino acids in this motif, fork reversal dropped about 25-fold in test tubes, yet the basic unwinding engine still worked reasonably well. In cells, the same mutation prevented the normal slowing of replication forks in response to a stress-inducing drug, a hallmark of failed fork reversal.
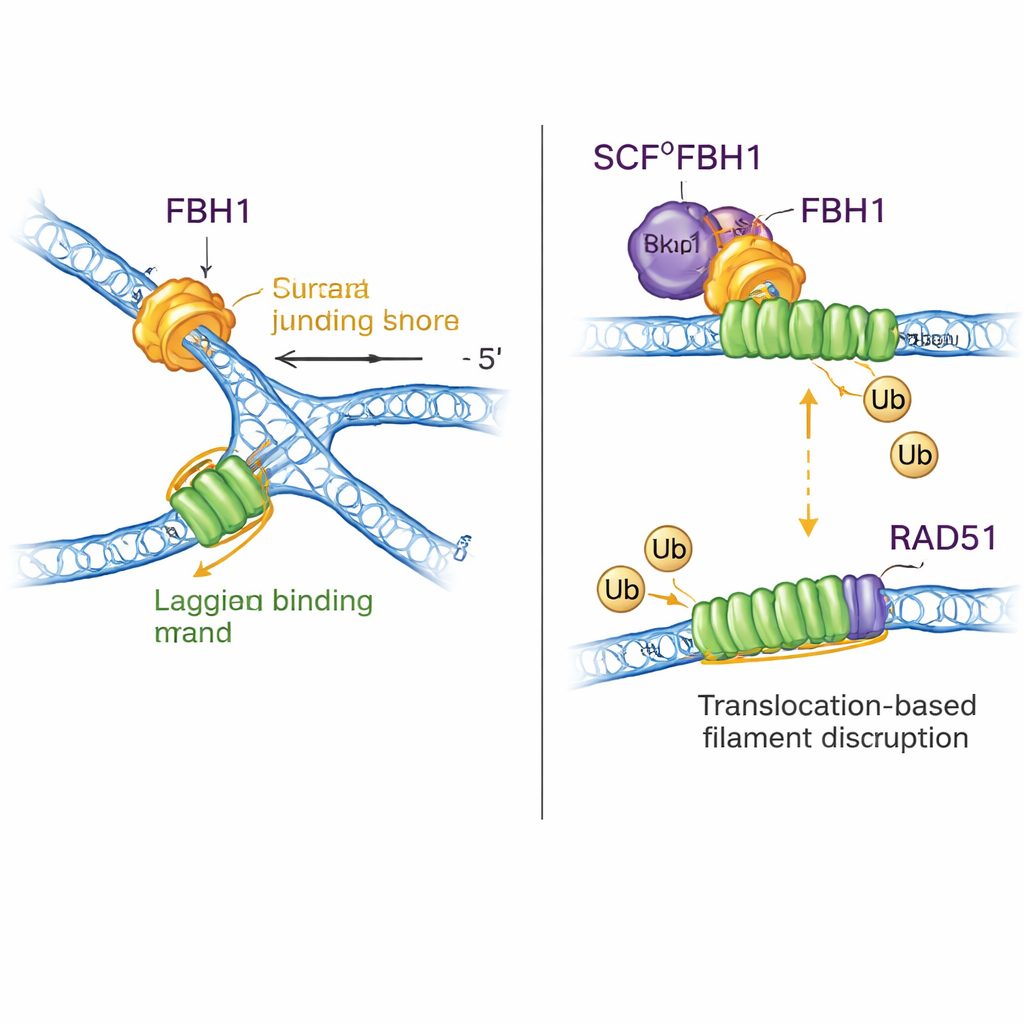
A dual role: reshaping DNA and removing RAD51
FBH1 is not just a motor; through the SCF complex it also helps attach small ubiquitin tags to proteins. The new structures assemble nearly the entire SCFFBH1–DNA complex and show that the ubiquitin-transfer “head” of the ligase sits less than 45 angstroms from the lagging-strand DNA emerging from FBH1. By docking in known structures of other ubiquitin ligases, the authors propose that this geometry positions RAD51—an enzyme that forms filaments on DNA to promote recombination—right in the line of fire on the lagging strand of a stalled fork. Prior work showed FBH1 can mechanically strip RAD51 off DNA and also mark it with ubiquitin so it does not readily rebind. The new model unifies these ideas: as FBH1 pulls along the lagging strand to reshape the fork, the nearby SCF module can tag RAD51 subunits, helping dismantle or prevent excessive RAD51 filaments that might otherwise block proper fork processing.
What this means for genome stability and cancer
Put together, the study shows that SCFFBH1 reverses stalled DNA replication forks using a mechanism distinct from other known fork remodelers: it grabs the branch point tightly while translocating along the lagging template from behind, promoting safe rewinding of parental DNA without forming a long-lived, mobile four-way junction. At the same time, its built-in ubiquitin ligase is strategically positioned to remove RAD51 from the same strand, preventing runaway recombination during replication stress. Because FBH1 is frequently lost or mutated in cancers, these structural and mechanistic insights help explain how such tumors accumulate DNA damage and why they may respond differently to drugs that provoke replication stress or target related pathways.
Citation: Greer, B.H., Mendia-Garcia, J., Mullins, E.A. et al. Structural basis for fork reversal and RAD51 regulation by the SCF ubiquitin ligase complex of F-box helicase 1. Nat Commun 17, 2041 (2026). https://doi.org/10.1038/s41467-026-68752-2
Keywords: DNA replication stress, fork reversal, FBH1 helicase, RAD51 regulation, genome stability