Clear Sky Science · en
Automated mapping of DNA replication fork progression in human cells with ForkML
Why tracking DNA copying speed matters
Every time a human cell divides, it has to copy more than three billion DNA letters quickly and accurately. If this copying process slows down or stalls, it can damage the genome and contribute to cancer and developmental disorders. Yet, until now, scientists have lacked a simple way to see exactly how fast individual DNA "copying machines" move along specific stretches of human DNA. This article presents ForkML, a new technique that uses nanopore DNA sequencing and machine learning to automate this task at an unprecedented scale.
Watching the cell’s copying machines in real time
DNA is duplicated by tiny molecular machines called replication forks that move along the double helix, making new strands. ForkML lets researchers watch these forks indirectly by adding a harmless chemical tag, BrdU, into newly made DNA for two very short bursts separated by a fixed time. Because BrdU can be detected in single DNA molecules by nanopore sequencers, the scientists see two tagged "stripes" on each DNA strand where a fork passed during the two pulses. By measuring the distance between the stripes and dividing by the known time gap, they can calculate how fast each fork moved in that region of the genome. 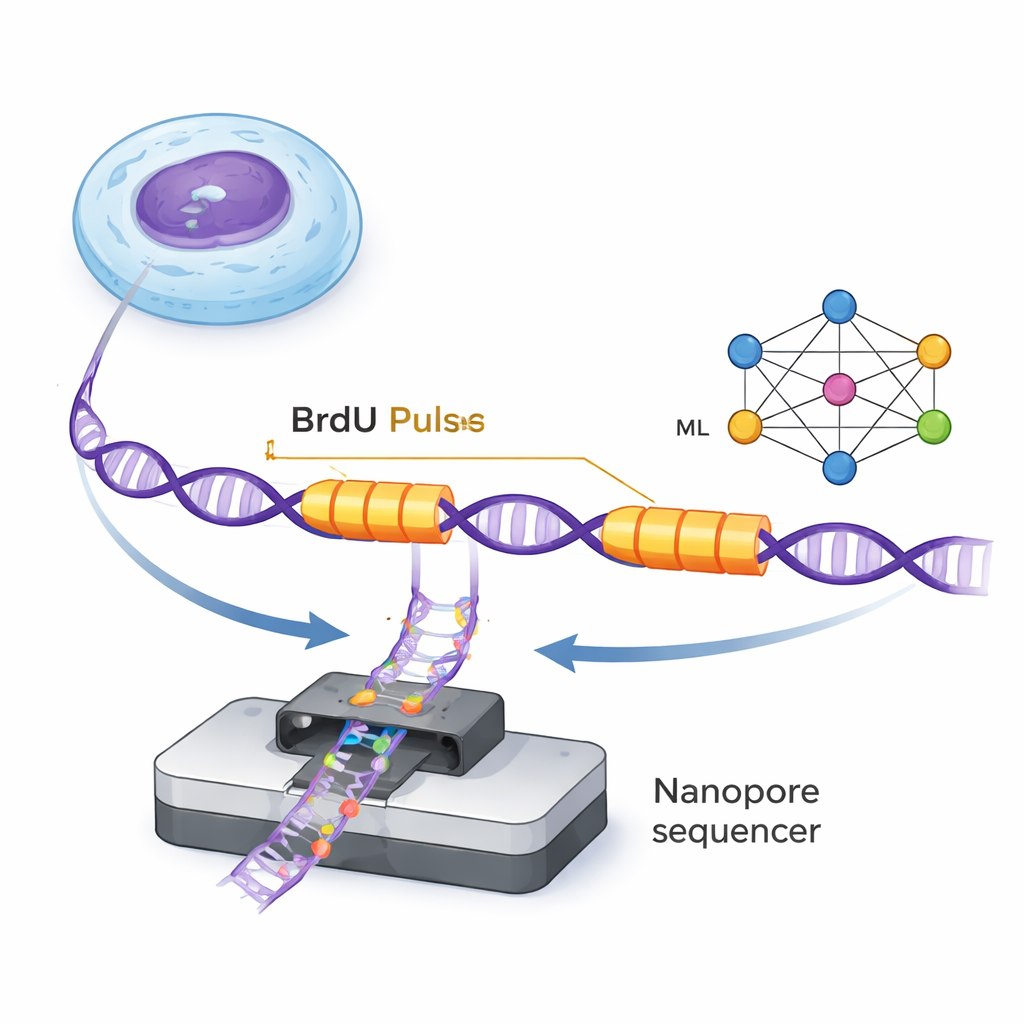
Teaching a computer to read the chemical tracks
In earlier work with yeast, the authors could spot these BrdU tracks using simple rules, but in human cells the signals are fainter and more complex. Human experts can still recognize the characteristic pattern—a sharp rise in BrdU as the pulse begins, followed by a gentle decline when it is washed out—but doing this by eye for millions of DNA fragments is impossible. ForkML solves this by training a neural network, a form of machine learning, on thousands of manually annotated examples. The model learns to classify each stretch of DNA as background or as a right- or left-moving fork, and to pinpoint the start of each BrdU pulse with high accuracy. This allows fully automated mapping of thousands of individual fork speeds from a single sequencing run.
Measuring stress and differences across the genome
Using ForkML on a human colon cancer cell line, the team obtained over 2,000 fork speed measurements per experiment and found that the typical fork moves at about 1.2 kilobases per minute, consistent with earlier, lower-throughput methods. When they treated cells with drugs that are known to slow DNA replication, ForkML clearly detected the slowdown, proving it can sensitively measure replication stress. Because every fork is mapped back to its position in the reference genome, the authors could then relate speed to other features, such as when a region normally replicates during the cell cycle, how tightly its DNA is packed, and how actively it is being transcribed into RNA. 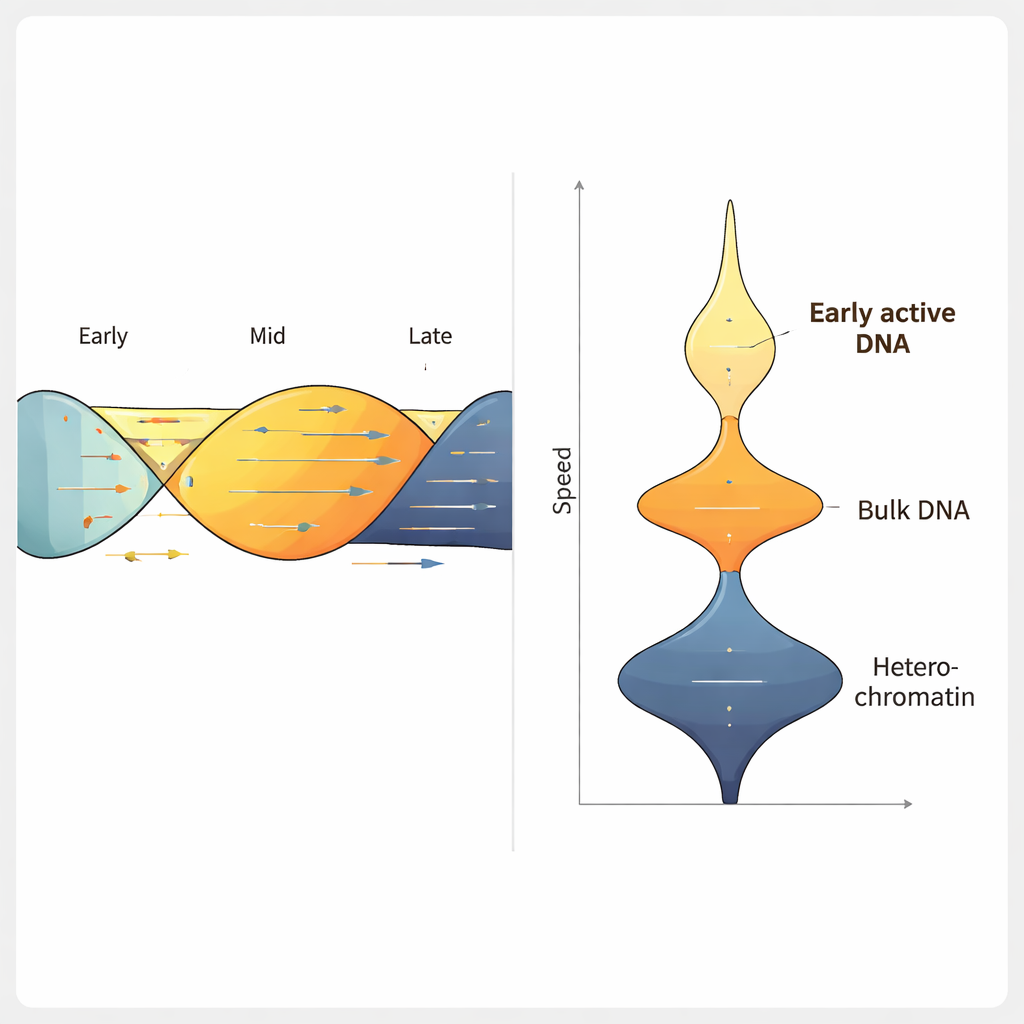
Revealing where DNA copying begins and how strands differ
Beyond speed, ForkML also identifies where DNA replication starts and stops, by spotting points where forks diverge or converge along the same molecule. Mapping more than 20,000 such start sites, the authors confirm that human cells use a mixed strategy: some copying begins in well-defined initiation zones, but most starts are scattered across the genome. By combining fork direction with which DNA strand was read by the sequencer, ForkML can also distinguish the rates of leading- and lagging-strand synthesis, something traditional fibre-based assays cannot do. Tests across six different human cell lines—both normal and cancerous—show that the same simple BrdU labeling conditions work broadly and yield robust speed estimates in each case.
A digital upgrade of a classic technique
To non-specialists, ForkML can be viewed as a modern, digital version of the classic DNA fibre assay: it uses a similar labeling scheme, but replaces manual microscopy with long-read sequencing and machine learning. This brings much higher throughput, direct placement of each measurement on the genome, and more detailed information on where and how quickly DNA is copied. Because the protocol is simple, compatible with current nanopore hardware, and adaptable to other organisms, ForkML is poised to become a standard tool for studying DNA replication. In practical terms, it offers researchers a powerful way to link local DNA copying speed—normal or stressed—to gene activity, chromatin state, and disease-related changes in the genome.
Citation: Rojat, V., Ciardo, D., Tourancheau, A. et al. Automated mapping of DNA replication fork progression in human cells with ForkML. Nat Commun 17, 1975 (2026). https://doi.org/10.1038/s41467-026-68750-4
Keywords: DNA replication, replication fork speed, nanopore sequencing, BrdU labeling, machine learning in genomics