Clear Sky Science · en
Large-scale capsid-mediated mobilisation of bacterial genomic DNA in the gut microbiome
Why tiny DNA shuttles in your gut matter
Your intestines are home to trillions of microbes whose genes help digest food, train the immune system and even influence mood. But those genes are not static. They can move from one bacterium to another, reshaping what the microbiome can do. This study uncovers a hidden highway for gene exchange in the human gut: microscopic protein shells, or capsids, that normally package viral DNA but frequently carry chunks of bacterial DNA instead. Understanding this traffic helps explain how our gut ecosystem adapts so quickly—to diet, drugs and disease.
Hidden couriers in the gut ecosystem
In the crowded gut, bacteria constantly swap genes, a process known as horizontal gene transfer. Some of this exchange is driven by viruses that infect bacteria (bacteriophages) and by virus-like particles called gene transfer agents. These structures are essentially tiny capsules that can ferry DNA from one cell to another. Until now, most evidence for this activity in the human gut came from indirect genetic patterns. It was hard to catch individual DNA-carrying particles in the act, and to tell real gene-transfer vehicles apart from random DNA debris released when cells burst.
To get a clearer picture, the researchers collected stool from three healthy adults and purified virus-like particles from each sample. They then used long-read nanopore sequencing, which can read out whole DNA molecules in one go. Because each type of capsid can only hold DNA up to a certain length, the sizes of these DNA fragments act like fingerprints for different transfer mechanisms. The team first validated their approach on well-studied lab systems in which the behavior of phages and gene transfer agents is already known, confirming that distinct length peaks in the data really do reflect DNA packaged inside intact particles.
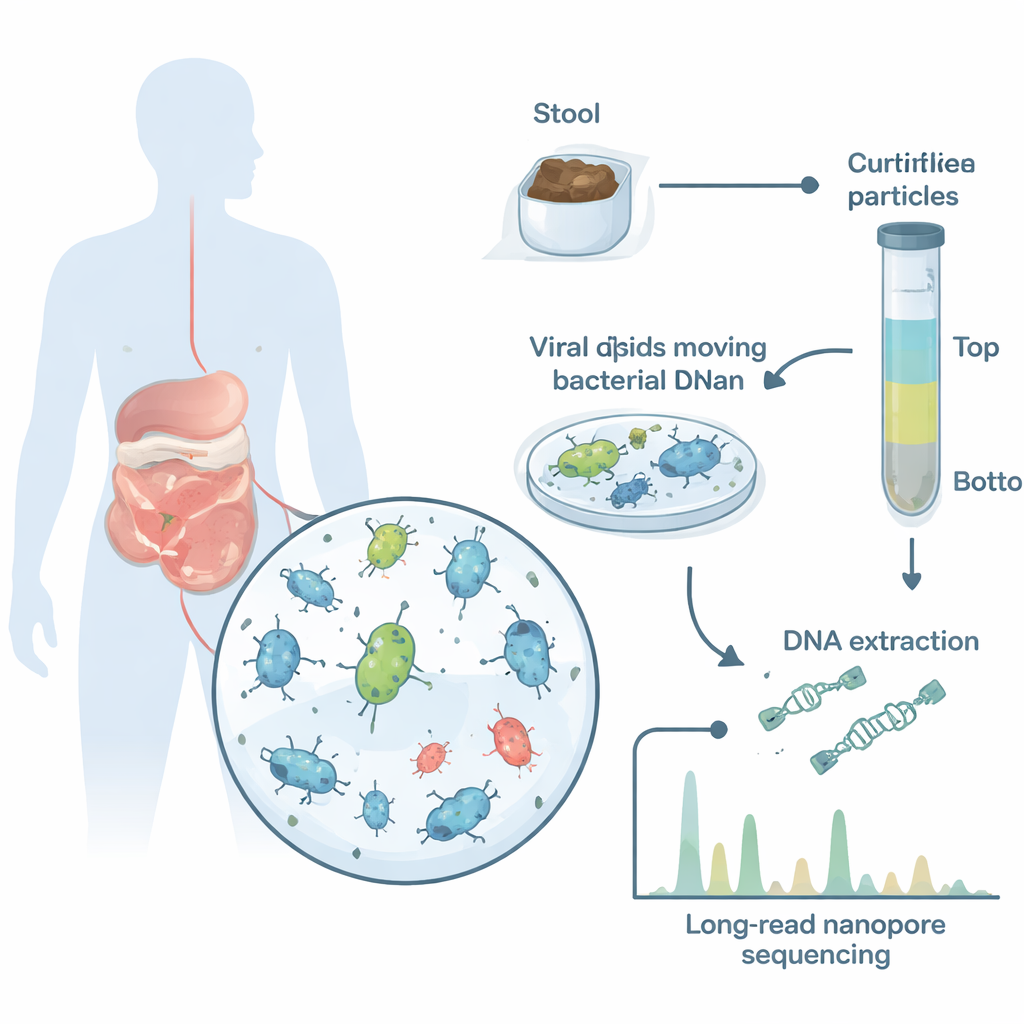
Measuring DNA packages one molecule at a time
When the gut samples were analyzed, the virus-like fractions showed clear peaks of DNA lengths from about 4,000 to 100,000 genetic letters, each peak representing a distinct population of particles. Up to 5.4% of all DNA inside these capsids came from bacterial, not viral, genomes—strong evidence that large-scale bacterial DNA packaging is common in the human gut. By combining long reads with conventional short-read sequencing, the scientists reconstructed many bacterial and viral genomes from the same samples and mapped each long DNA molecule back to its source. This allowed them to see exactly which bacterial groups were donating DNA, which regions of their chromosomes were packaged and how the packaging patterns looked.
The analysis revealed that not all bacteria contribute equally. While the overall bacterial community in stool was dominated by families such as Bacteroidaceae and Lachnospiraceae, the virus-like fractions were enriched for DNA from other groups, including Ruminococcaceae and Oscillospiraceae. In some cases, only narrow genomic regions near dormant viruses embedded in bacterial chromosomes were packaged, matching classic "prophage induction." In others, long runs of chromosome sequence extending away from these embedded viruses were captured, a hallmark of a powerful process called lateral transduction that can mobilize large swaths of bacterial DNA in a single event.
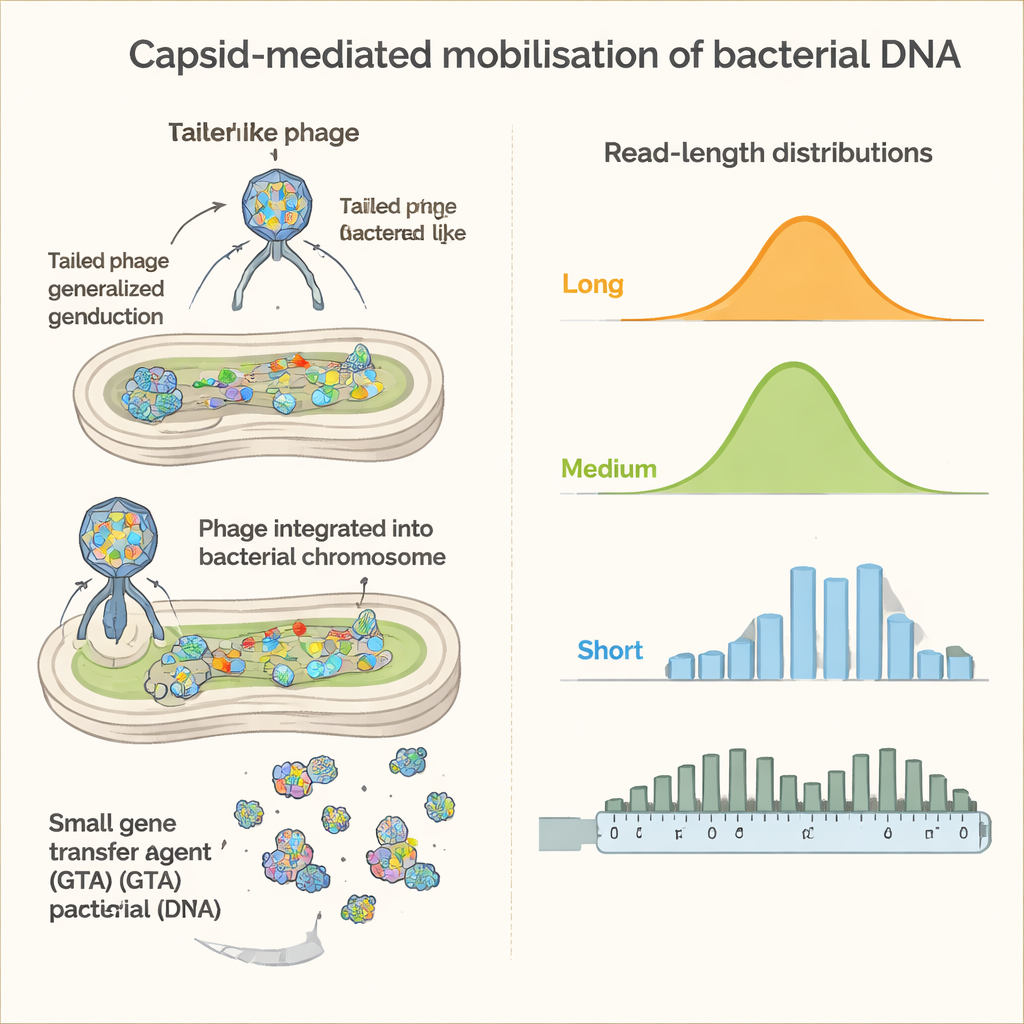
Discovery of busy gene-transfer hubs
Beyond these known mechanisms, one of the most striking findings was the prevalence of gene transfer agent–like behavior among certain gut bacteria. In members of the Ruminococcaceae and Oscillospiraceae families—including the important gut genus Faecalibacterium—the researchers observed huge numbers of particles that package many short, randomly scattered DNA fragments, typically 4,600 to 8,900 letters long. This pattern closely matches gene transfer agents described in other environments, which resemble domesticated viruses that have been repurposed by bacteria to distribute their own DNA.
Digging deeper into Faecalibacterium genomes, the team identified two clusters of genes that together appear capable of building such particles, packaging DNA and breaking open the host cell. In the lab, a Faecalibacterium strain carrying these clusters spontaneously produced capsid-like particles that held DNA fragments of the expected size. Electron microscopy showed small, roughly spherical shells, and protein analysis confirmed that the major components of these shells are encoded by the newly identified gene clusters. This strongly suggests that Faecalibacterium, one of the most abundant and health-associated bacteria in the human gut, is actively producing gene-transfer particles.
What this means for your microbiome
By reading the full lengths of DNA molecules inside virus-like particles, this work shows that capsid-mediated gene transfer is not a rare curiosity but a routine feature of the human gut. Many different mechanisms—classical viral transduction, lateral transduction and gene transfer agents—appear to be constantly moving bacterial DNA around, especially in key groups like Bacteroides and Faecalibacterium. For the microbiome, this means a built-in ability to rapidly reshuffle useful traits, from nutrient processing to drug resistance. For us, it underscores that our gut ecosystem is not just a static collection of species, but a highly dynamic genetic marketplace whose invisible couriers are hard at work every day.
Citation: Borodovich, T., Buttimer, C., Wilson, J.S. et al. Large-scale capsid-mediated mobilisation of bacterial genomic DNA in the gut microbiome. Nat Commun 17, 2046 (2026). https://doi.org/10.1038/s41467-026-68726-4
Keywords: gut microbiome, bacteriophages, horizontal gene transfer, gene transfer agents, viral capsids