Clear Sky Science · en
Click-based determination of accumulation of molecules in Escherichia coli
Why tiny drugs struggle to reach dangerous bacteria
Modern medicine depends on antibiotics, yet some of the most dangerous bacteria are now shielded by tough outer walls that keep many drugs out. Gram‑negative bacteria such as Escherichia coli have an especially effective outer membrane barrier, making it hard to design medicines that actually get inside and reach their targets. This paper introduces a new lab test, called the CHAMP assay, that lets scientists quickly measure how well thousands of different small molecules slip past these defenses and accumulate inside living E. coli cells.
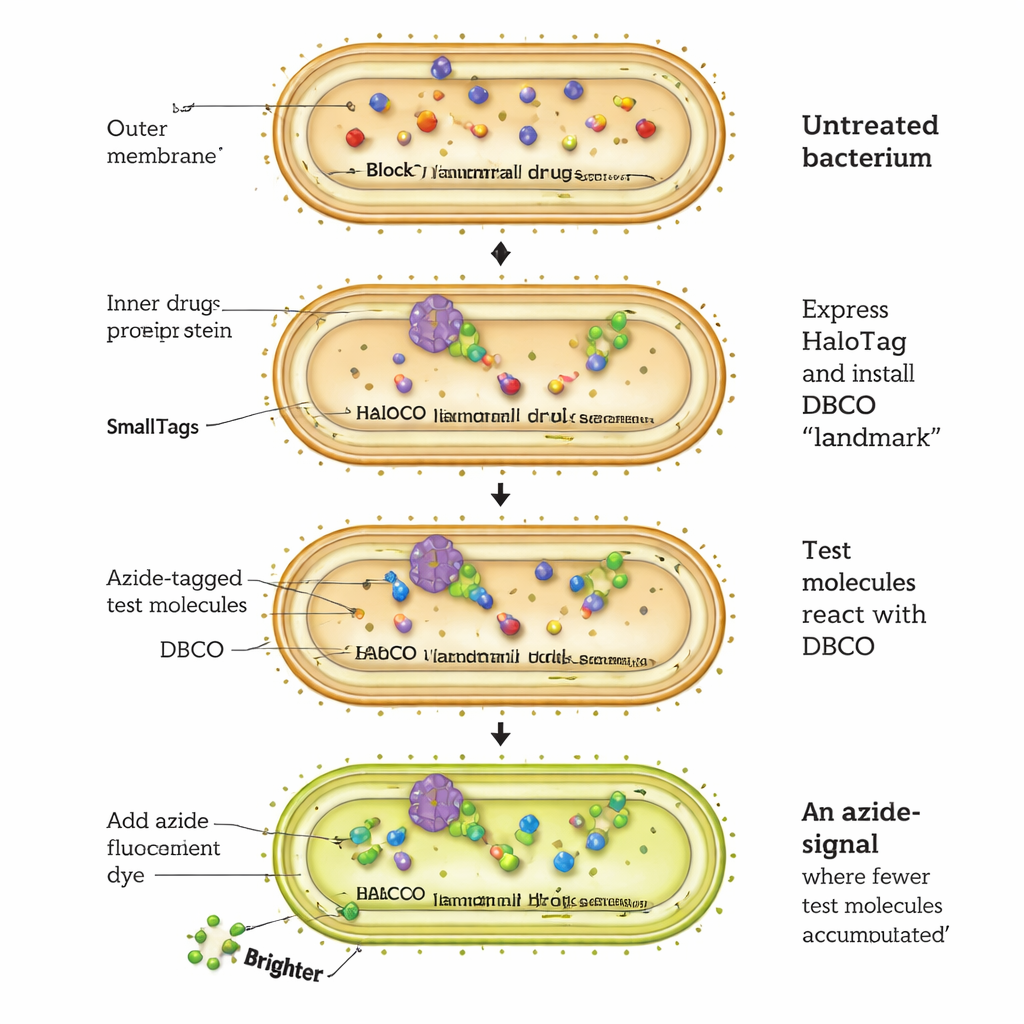
A new way to watch drugs enter bacterial cells
Traditional antibiotic discovery relied on naturally occurring compounds and simple growth tests to see whether bacteria lived or died. Those tests do not reveal how much drug actually made it into the cell, or where it went once inside. The authors set out to build a direct, widely usable method for tracking drug entry into the bacterial cytosol—the fluid interior where many antibiotic targets reside. Their goal was to move beyond crude readouts like minimum inhibitory concentration, which mix together effects of drug uptake, target binding, and downstream biology, and instead get a clean measurement of intracellular accumulation.
Turning bacteria into tiny chemical recorders
The CHAMP assay works by rewiring a bacterial protein called HaloTag into a kind of molecular landing pad. E. coli cells are engineered to produce HaloTag in their cytosol. Researchers first attach a special chemical handle, a "strained" alkyne called DBCO, to HaloTag using a short chloroalkane linker. Next, they expose the bacteria to test molecules that each carry a very small azide tag. Only those test molecules that cross the outer membrane, pass the inner membrane, and reach the cytosol can react with the DBCO handle in a highly selective "click" reaction, permanently marking HaloTag. Finally, the cells are treated with an azide‑tagged fluorescent dye that can only bind to any remaining unoccupied DBCO. The brighter the fluorescence, the fewer test molecules arrived in the cytosol; dimmer cells indicate better accumulation.
Probing how chemistry and biology shape drug entry
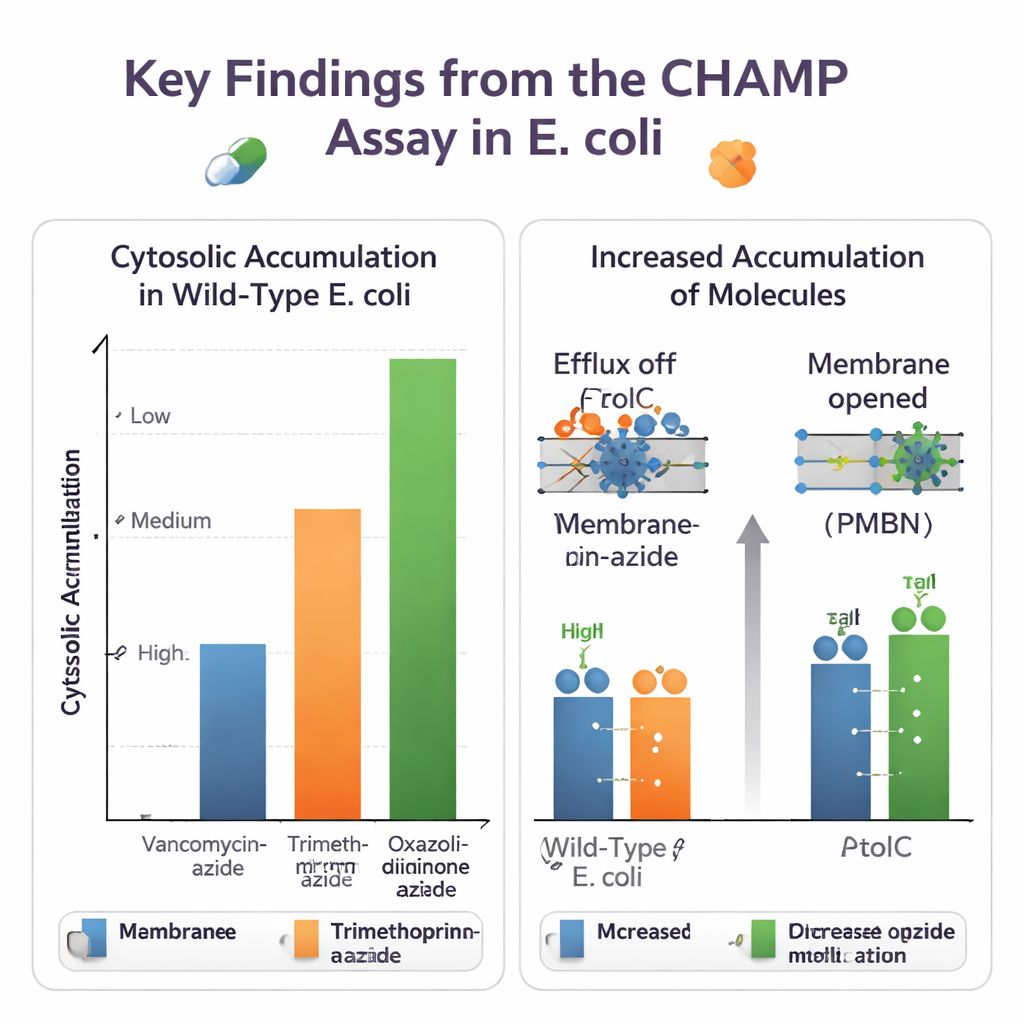
Using CHAMP, the team first fine‑tuned expression of HaloTag, the amount of DBCO anchor, and the choice of fluorescent dyes to get a large and reliable signal window. They then used sets of closely related small molecules to see how chemical tweaks affect entry. Converting carboxylic acids to amides, and adding or removing basic amino groups, could noticeably change cytosolic levels. In many cases, primary amines enhanced accumulation, echoing independent "eNTRy rules" developed by other groups. The assay also handled azide‑tagged versions of real antibiotics, revealing big differences: bulky drugs like vancomycin barely reached the cytosol at all, while smaller antibiotics such as trimethoprim derivatives and some oxazolidinones entered much more readily.
Unmasking bacterial defenses that pump drugs back out
Because the method runs in live, genetically programmable bacteria, it can untangle the roles of different cellular defenses. The authors compared normal E. coli with strains lacking TolC, a key component of a major multidrug efflux pump, or with cells treated with a small‑molecule TolC inhibitor. For novobiocin and several other azide‑tagged antibiotics, CHAMP showed clearly higher cytosolic accumulation when TolC was disabled, confirming these compounds as efflux substrates. They also tested ways of loosening the outer membrane barrier, either chemically with the peptide PMBN or genetically by overexpressing a wide open porin channel. Many molecules that previously accumulated poorly showed large gains when the membrane was permeabilized, highlighting how entry and efflux act together to set internal drug levels.
Scaling up to thousands of molecules for patterns and rules
To demonstrate true high throughput, the researchers screened a commercial set of 404 azide‑tagged compounds and a custom library of 1,152 azide‑bearing molecules. They measured accumulation in several bacterial contexts, including untreated cells, PMBN‑treated cells, hyperporinated cells, and hyperporinated cells lacking TolC. By comparing the same molecule across these conditions, they could see which changes were due to the cell biology rather than quirks of click‑reaction speed. Combining CHAMP data with computational analysis, they linked certain physicochemical features—such as polar surface area, hydrogen‑bonding capacity, and specific ring scaffolds—to stronger recognition by TolC‑based efflux. This kind of large‑scale profiling begins to outline design rules for molecules that either evade pumps or exploit porins more effectively.
What this means for future antibiotics
In simple terms, this work turns E. coli into a high‑throughput sensor that reports exactly how much of a test compound gets into the part of the cell where many drug targets live. CHAMP does not replace tests of whether bacteria are killed, but it fills a crucial gap by separating "getting in" from "doing damage." With the ability to measure more than a thousand molecules in different mutant or chemically altered strains within days, researchers can now systematically learn which chemical features promote entry, reduce efflux, or benefit from membrane‑disrupting partners. That knowledge should speed the design of new antibiotics and helper drugs that can breach the formidable defenses of Gram‑negative pathogens before resistance claims more of the antibiotic arsenal.
Citation: Ongwae, G.M., Liu, Z., Feng, S. et al. Click-based determination of accumulation of molecules in Escherichia coli. Nat Commun 17, 2008 (2026). https://doi.org/10.1038/s41467-026-68717-5
Keywords: antibiotic resistance, Escherichia coli, drug permeability, efflux pumps, click chemistry