Clear Sky Science · en
High-speed blind structured illumination microscopy via unsupervised algorithm unrolling
Sharper Movies of Life Inside Cells
Modern biology often depends on watching living cells in action, but many key structures are simply too small and too fast for ordinary microscopes to capture clearly. This paper introduces a new way to turn blurry, quickly captured images into crisp, super‑detailed movies in real time, without needing perfectly tuned hardware. The method, called unrolled blind structured illumination microscopy (UBSIM), promises to make advanced, high‑speed cellular imaging more accessible to everyday biology labs.
Why Regular Microscopes Fall Short
Traditional light microscopes are limited by diffraction, a basic property of light that blurs fine details smaller than a few hundred nanometers. Structured illumination microscopy (SIM) tackles this by shining patterned light onto a sample and using the resulting interference to tease out extra detail, roughly doubling resolution. However, classic SIM demands precisely known illumination patterns and careful calibration, which can be expensive and fragile. A newer variant, blind-SIM, relaxes these hardware demands by allowing random patterns and solving for both the sample and the illumination from the data itself. The downside is that this solving process is slow and iterative, taking seconds to minutes per frame—far too sluggish for real-time movies of living cells.
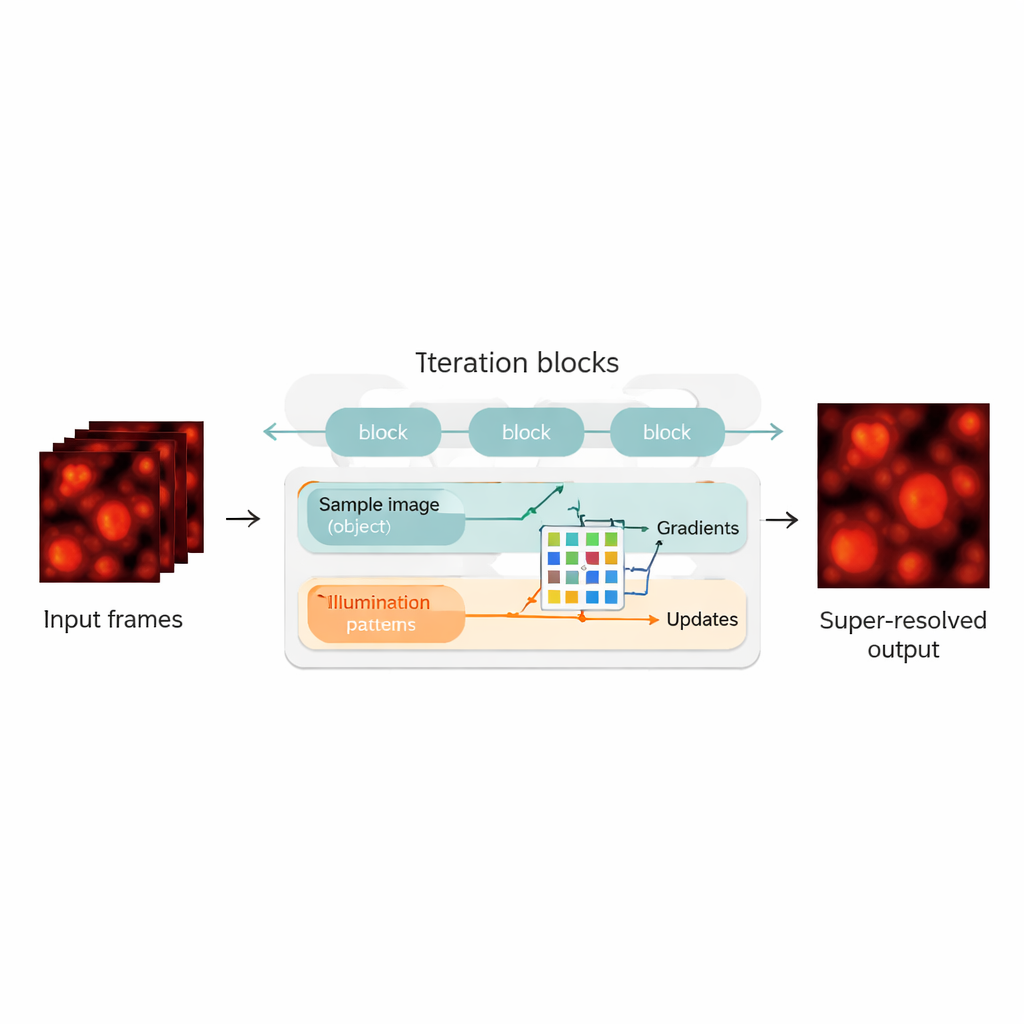
Blending Physics with Neural Networks
The authors bridge this gap by redesigning the blind-SIM reconstruction as a hybrid between a physics-based model and a neural network. They "unroll" the original iterative algorithm—each iteration becomes a layer in a neural network, forming a chain of update blocks. Within each block, the method computes how well the current guess of the sample and illumination explains the measured images, calculates gradients (directions of improvement), and then feeds these into a compact convolutional neural network. This network learns how to make smarter correction steps, playing a role similar to an automatically tuned accelerator for the original algorithm. Crucially, UBSIM is trained in an unsupervised way: instead of needing perfect example images as ground truth, it only needs the physical model of how light passes through the microscope. That reduces the risk of the network “hallucinating” plausible‑looking but incorrect structures.
Fast, Accurate, and Less Prone to Guesswork
To test UBSIM, the team first used simulated microscopy images where the true underlying structures are known. They showed that UBSIM recovers about twice the resolution of ordinary widefield images, comparable to standard blind-SIM, but runs two to three orders of magnitude faster—a 256×256 image can be reconstructed in about 10 milliseconds instead of seconds. Image quality scores, including error, similarity, and signal‑to‑noise measures, all improved markedly over conventional images. UBSIM also proved more trustworthy than popular deep-learning super‑resolution networks when confronted with unfamiliar data. While standard networks trained on one type of structure tended to imprint that pattern onto new, different samples—introducing subtle but misleading artifacts—UBSIM maintained consistent fidelity, because it is anchored to the underlying imaging physics rather than only to visual examples.
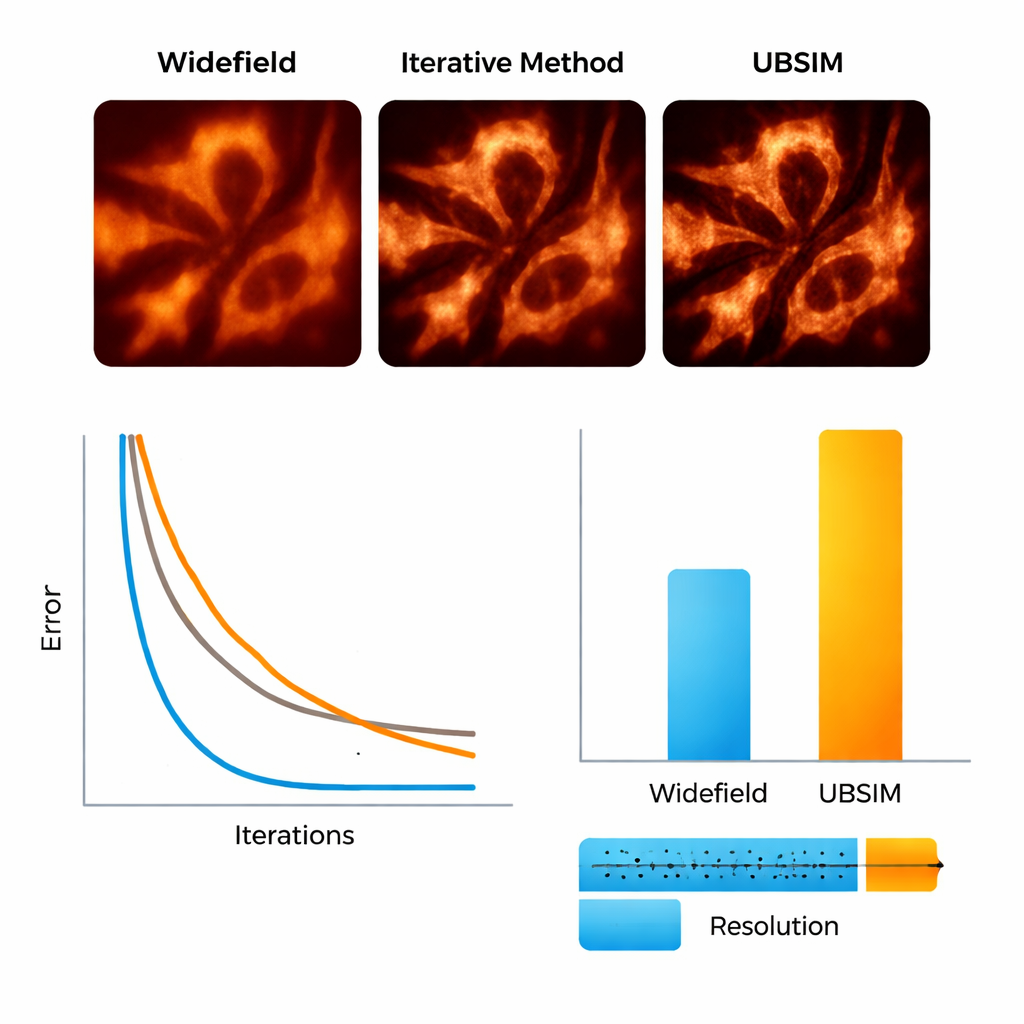
Seeing Cell Skeletons and Membranes in Motion
The researchers then moved to real biological samples. Using a flexible setup that projects random speckle patterns onto live cells, they imaged actin filaments—the protein “scaffolding” inside cells—and the endoplasmic reticulum (ER), a branching membrane network involved in protein production and cell stress responses. With UBSIM, actin fibers that appeared as fuzzy bands in ordinary images became sharply separated strands, with resolution improving from roughly 300 nanometers to about 150 nanometers. Most strikingly, UBSIM enabled video‑rate super‑resolution: by capturing raw data at up to 100 frames per second and reconstructing at up to 50 super‑resolved frames per second, the team could watch ER tubules grow, collapse, and reorganize in real time. These dynamics, occurring over fractions of a second to a few seconds, are normally hard to visualize with sufficient detail.
What This Means for Future Cell Imaging
For non‑specialists, the key takeaway is that UBSIM makes it far more practical to watch tiny cellular structures move, in real time, with clarity beyond the normal limits of light microscopes—all without demanding perfect hardware calibration or massive training datasets. By combining the reliability of physics‑based models with the speed of modern neural networks, this approach turns stacks of noisy, patterned images into trustworthy, ultra‑sharp movies fast enough for routine experiments. As the method is further refined and paired with better illumination strategies, it could help researchers probe how organelles like the ER respond to stress, how cell skeletons reorganize during movement or division, and how diseases alter cellular architecture at the nanoscale.
Citation: Burns, Z., Zhao, J., Sahan, A.Z. et al. High-speed blind structured illumination microscopy via unsupervised algorithm unrolling. Nat Commun 17, 1967 (2026). https://doi.org/10.1038/s41467-026-68693-w
Keywords: super-resolution microscopy, structured illumination, deep learning, live-cell imaging, endoplasmic reticulum dynamics