Clear Sky Science · en
ProteoAutoNet: high-throughput co-eluted protein analysis with robotics and machine learning
Why understanding protein partnerships matters
Inside every cell, proteins rarely work alone. They team up in shifting alliances to build structures, copy DNA, destroy damaged parts, and fuel growth. Many cancers hijack these partnerships, but mapping them in detail has been slow, painstaking work. This study introduces ProteoAutoNet, a robotics- and machine-learning–powered system that greatly speeds up how scientists discover protein partnerships in cells, and shows how this approach can reveal hidden weak points in thyroid cancers.
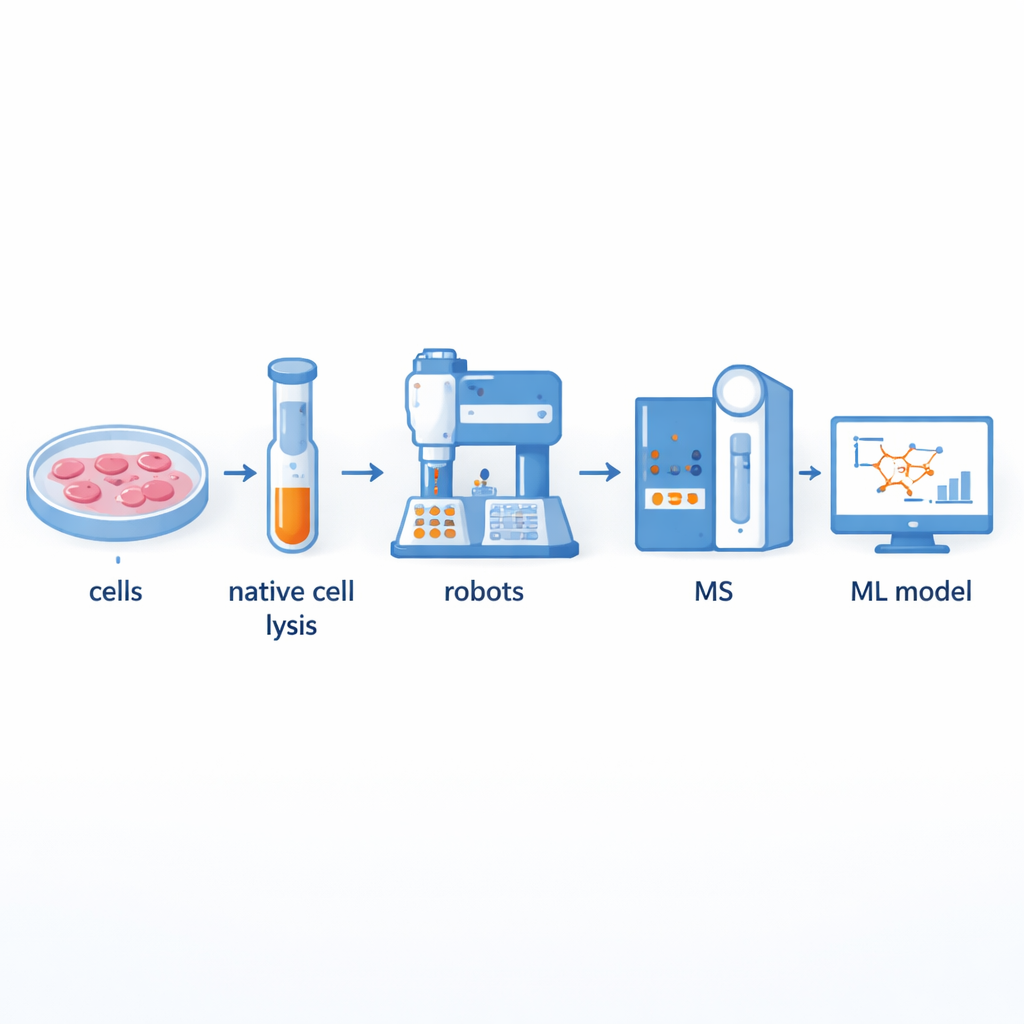
Building a faster protein partnership factory
Traditionally, scientists use a method called co-fractionation mass spectrometry to separate large protein complexes and then identify their components. While powerful, this approach is labor-intensive and low-throughput: preparing hundreds of fractions by hand can take many days. The authors built a robotics-assisted platform that automates most of this workflow. Cell contents are first gently broken open so that natural protein complexes remain intact, then passed through size-based columns to split them into dozens of fractions. Liquid-handling robots and robotic arms then take over, adding chemicals, digesting proteins into smaller pieces, cleaning up the samples, and delivering them to a mass spectrometer for measurement. This setup can process up to 540 fractions from multiple thyroid cell lines in just two to three days, roughly doubling throughput compared with previous semi-automated systems.
Robots that are not just faster, but more reliable
Speed alone is not enough if results are noisy or inconsistent. The team carefully checked whether the robotic pipeline matched or exceeded the quality of traditional manual processing. Using quality-control samples, they showed that the automated system repeatedly identified nearly 3,000 proteins per thyroid cell line with very high overlap between replicates and strong agreement in measured protein amounts. When they directly compared robotic and manual processing of the same samples, both approaches detected similar numbers of proteins, but the robotic method produced slightly less variation in counts and more stable protein abundance measurements. This means the new platform not only saves time and labor, but also supports more reproducible experiments—a crucial requirement for large studies and clinical applications.
Teaching computers to recognize meaningful connections
Even with fast instruments, a central challenge remains: deciding which proteins truly interact and which merely appear together by chance. To tackle this, the authors combined curated protein-complex databases with a machine-learning model based on the XGBoost algorithm. They first cleaned and merged three major protein complex resources, ending up with 96,635 known protein–protein interactions. They then used profiles of how proteins appeared across the fractions as input features, and labeled pairs as likely partners or non-partners based on the databases. Because real, high-confidence partnerships are relatively rare, they used a targeted data-augmentation strategy: they made many slightly perturbed versions of known positive examples to teach the model to recognize robust patterns rather than memorize specific traces. Trained on tens of millions of such examples from three thyroid cell lines, the model achieved strong performance, correctly ranking true interactions well above random both in internal tests and in an independent validation cell line.
New views on cancer cell machinery
Armed with this workflow, the researchers charted interaction networks in a normal thyroid cell line and two cancerous ones: a papillary thyroid carcinoma line and a follicular carcinoma line that can spread to the lungs. Across these cells, they identified over 25,000 likely protein interactions and found strong signals from well-known cellular machines such as ribosomes (which build proteins) and proteasomes (which break them down), confirming that the method recovers established biology. By comparing cancers to the normal line, they uncovered networks that were dialed up in disease. In the metastatic follicular carcinoma cells, both proteasome components and a chaperone complex called prefoldin were markedly more connected and abundant. Several prefoldin subunits had previously been linked to other cancers, but global protein surveys had missed their coordinated behavior in thyroid cancer, possibly because these proteins are tightly controlled by degradation. The co-fractionation approach exposed their coordinated changes at the complex level.
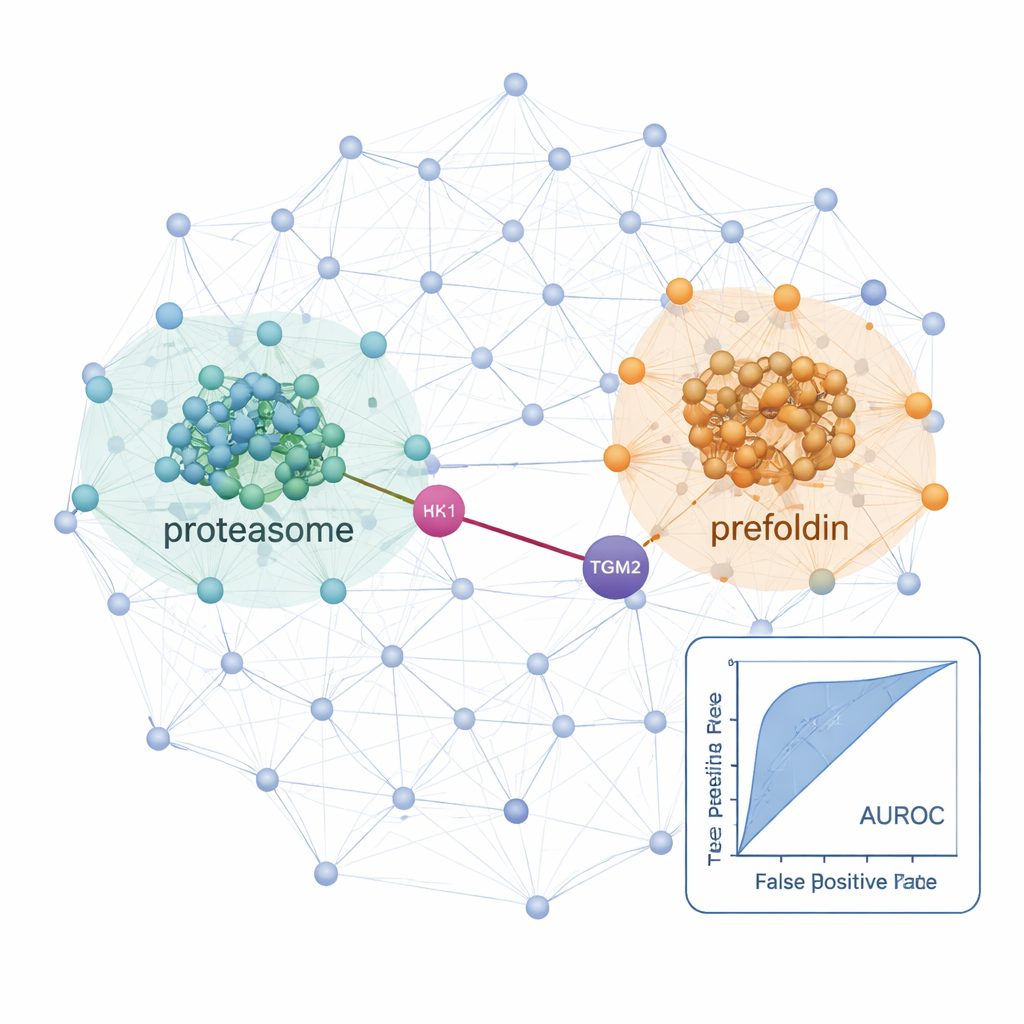
Hidden links that may guide future treatments
The study also highlighted specific interactions that could matter for how thyroid cancers grow and spread. One example is a predicted partnership between HK1, an enzyme that starts the cell’s main sugar-burning pathway, and TGM2, a protein known to encourage invasion and metastasis in thyroid tumors. This HK1–TGM2 connection, absent from existing interaction databases, was supported by structural modeling and appeared particularly active in the papillary carcinoma line, hinting that metabolic reprogramming and invasive behavior may be physically linked. Taken together, ProteoAutoNet shows how combining robotics and machine learning can turn slow, expert-only protein network mapping into a more scalable process. For non-specialists, the key message is that this technology can uncover both broad shifts in cellular machinery and unexpected protein partnerships that may one day help doctors better predict which thyroid cancers will behave aggressively and suggest new targets for therapy.
Citation: Lyu, M., Hu, P., Zhang, G. et al. ProteoAutoNet: high-throughput co-eluted protein analysis with robotics and machine learning. Nat Commun 17, 1949 (2026). https://doi.org/10.1038/s41467-026-68686-9
Keywords: protein interactions, mass spectrometry, machine learning in biology, thyroid cancer, proteasome and prefoldin