Clear Sky Science · en
High-throughput methods leveraging robotics and computer vision for the development of therapeutic phage cocktails
Why this matters for everyday infections
Antibiotic resistance is turning once-routine infections, like urinary tract infections (UTIs), into stubborn and sometimes life-threatening problems. This study describes how scientists are harnessing robots, cameras, and data analysis to build virus-based medicines, called phage cocktails, that can specifically attack drug-resistant bacteria causing UTIs. The work shows how an industrial-style, automated lab can sift through huge numbers of virus–bacteria pairings to design a single, off-the-shelf treatment that works for most patients.
Turning a common infection into a test case
UTIs are among the most frequent reasons people receive antibiotics, especially women, many of whom experience repeated infections. The main culprit is a form of E. coli that has become increasingly resistant to standard drugs. Traditional antibiotics act broadly and can disrupt the body’s helpful microbes, while also pushing bacteria to evolve resistance. Phages—viruses that infect bacteria—offer a more targeted option, but each phage tends to attack only certain bacterial strains. The challenge is to find the right mix of phages that can collectively handle the real-world diversity of UTI-causing E. coli.
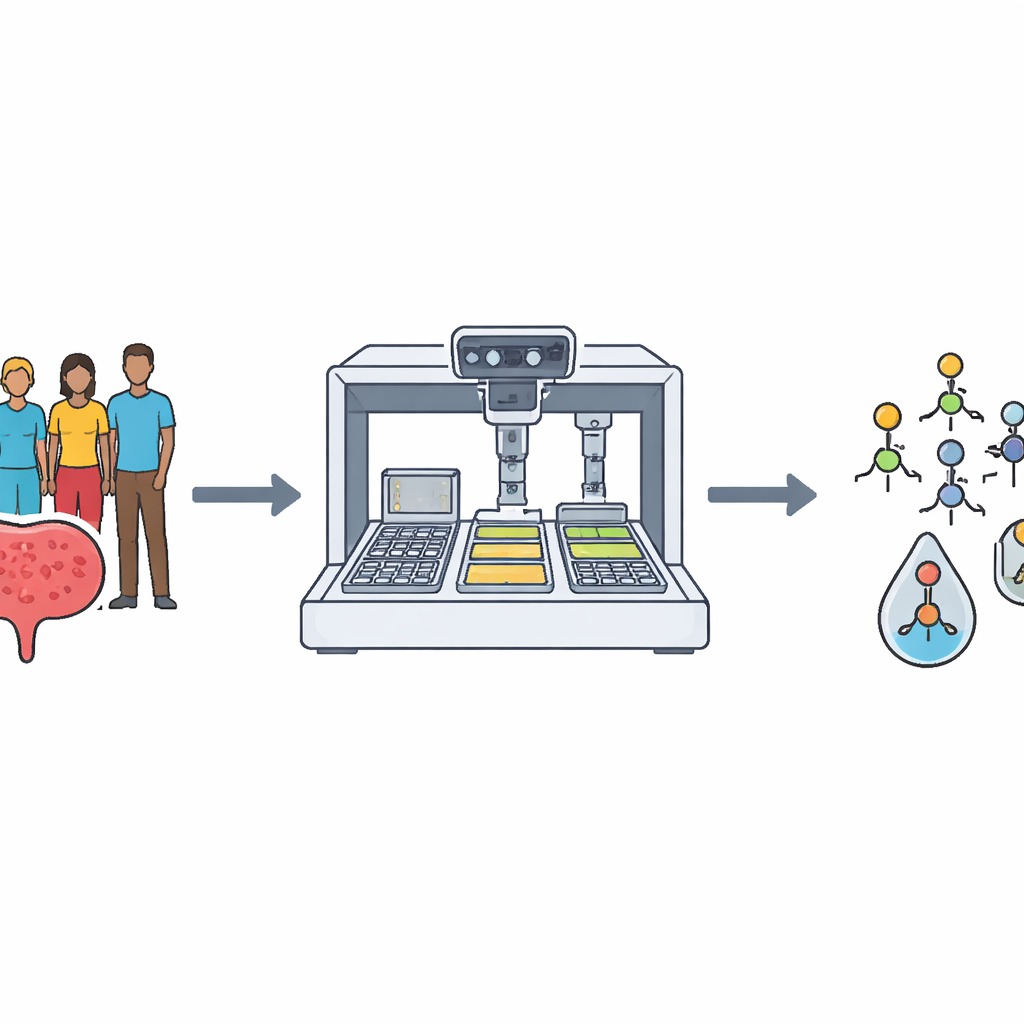
Building a realistic library of problem bacteria
The team first assembled a large and diverse panel of UTI-causing E. coli strains to mirror what is seen in clinics. From about 1,700 isolates collected from patients and commercial sources, and guided by genetic data from roughly 10,000 publicly available E. coli genomes, they selected 356 strains for a “Clinical Panel.” These strains captured major genetic lineages, came from patients across 39 U.S. states, and included many that were resistant to multiple antibiotic classes. Each strain was carefully grown, portioned, barcoded, and stored in automated freezers so robots could repeatedly and reliably pull them for testing.
Robots and cameras hunt for the best virus mixture
Next, the researchers turned to the other side of the equation: the phages. They collected over a thousand environmental samples, mainly from wastewater, and used their Clinical Panel to fish out 1,143 E. coli-targeting phages, of which 421 became the main workhorses. A custom robotic platform used multi-channel pipettes, incubators, and plate readers to mix bacteria, phages, and growth media in tiny wells and track how well bacteria grew over 20 hours. By comparing the growth of phage-treated wells to untreated controls, the system measured how effectively each phage—or combination of phages—suppressed bacterial growth. Over time, this produced about 1.5 million growth curves and more than 3.8 million phage–bacteria reaction records, which fed into a cocktail-prediction model that suggested new, promising combinations to test.
Letting computer vision count the survivors
Optical readings show overall cloudiness but cannot easily detect when only a small number of bacteria remain. To quantify deep killing, the team added a second automated assay. Robots diluted samples, spotted them onto agar plates, and incubated them so individual bacterial survivors formed visible colonies. High-resolution cameras then imaged the plates. A custom image-analysis pipeline used three different algorithms to count colonies and phage-induced clearings, cross-checking results and flagging disagreements for human review. Across more than two million spots, the automated system matched or exceeded the accuracy of trained analysts while operating at far higher scale, providing reliable counts of live bacteria and phage particles.
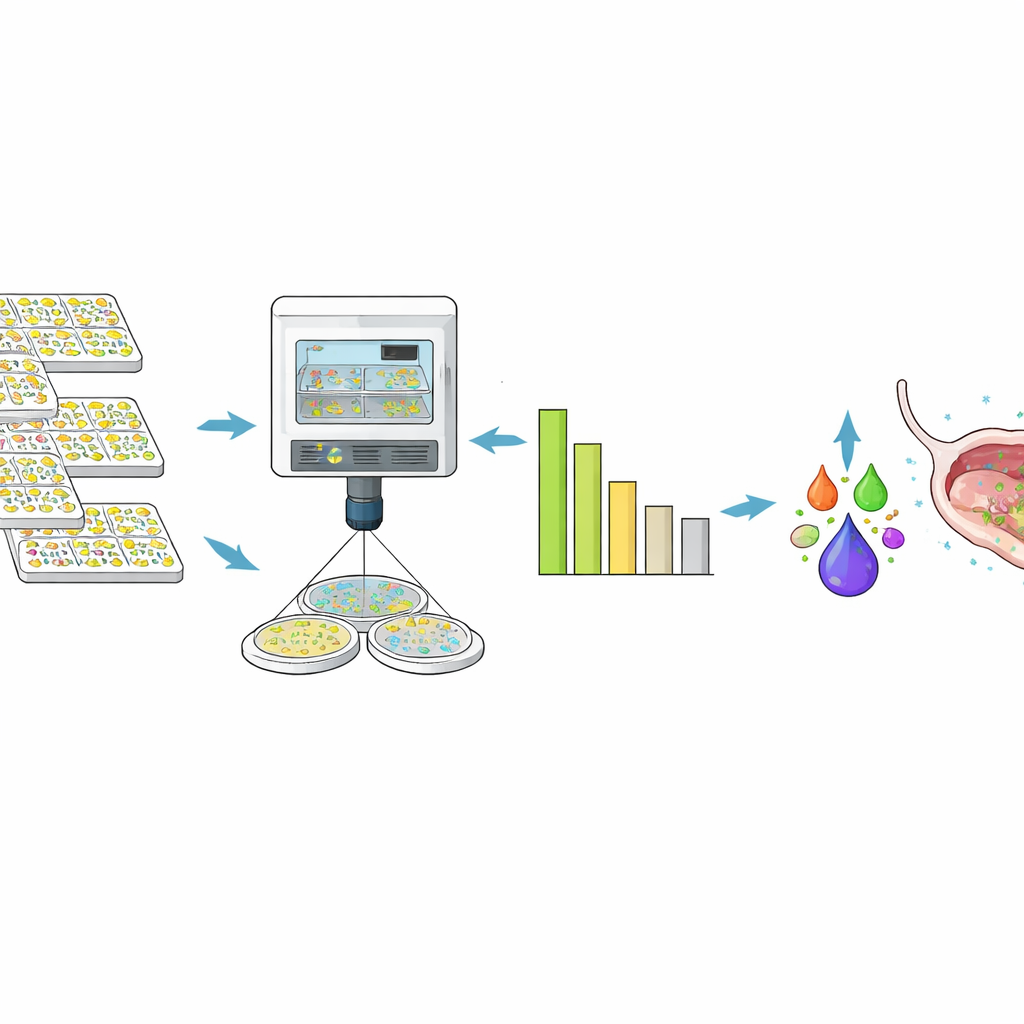
From millions of tests to one powerful cocktail
Using these high-throughput assays and prediction tools, the researchers narrowed thousands of possibilities down to a six-phage cocktail, LBP-EC01. Manufactured under pharmaceutical-grade conditions, this cocktail showed in-lab activity against 96.4% of the 356 Clinical Panel strains and cut bacterial numbers by at least 100,000-fold in the majority of cases. When the team tested E. coli isolates from the first part of an ongoing Phase 2 clinical trial in women with acute UTIs, 97% of unique strains were sensitive to the cocktail, and the pattern of killing closely matched what was seen in the pre-built panel. Importantly, they saw no evidence that patients’ bacteria evolved genetic resistance to the phages during the monitored period.
What this means for future treatments
Put simply, this work shows that carefully designed automation and imaging can take the guesswork out of phage therapy. By pairing realistic collections of patient bacteria with robotic testing and computer vision, the researchers created a broadly active phage cocktail that remains effective against real-world clinical isolates. While larger trials are needed to confirm clinical benefits, the approach offers a blueprint for developing fixed, scalable phage medicines to tackle multidrug-resistant infections and reduce reliance on traditional antibiotics.
Citation: Penke, T.J.R., Hammack, A.T., McMillan, L.J. et al. High-throughput methods leveraging robotics and computer vision for the development of therapeutic phage cocktails. Nat Commun 17, 2192 (2026). https://doi.org/10.1038/s41467-026-68684-x
Keywords: phage therapy, urinary tract infection, antibiotic resistance, laboratory automation, computer vision