Clear Sky Science · en
Iterative discovery of potent polymeric antibiotics via multi-stage and multi-task learning against antimicrobial resistance
Why new antibiotics matter to everyone
Drug-resistant infections are rising worldwide, making familiar illnesses harder and sometimes impossible to treat. This study describes a new way to rapidly discover a different kind of antibiotic: tiny synthetic polymers that can both kill tough bacteria and help old drugs, like penicillin, work again. The work blends advanced artificial intelligence with chemistry and animal testing to search an enormous chemical space that would be impossible to explore by trial and error alone.
Hunting for new defenders in a crowded chemical universe
Traditional antibiotics usually target specific bacterial proteins, which bacteria can gradually evade. By contrast, the polymers in this study mimic our body’s own defense peptides, which physically damage bacterial membranes and are less prone to resistance. The challenge is that there are tens of thousands of possible polymer structures, and their behavior depends on a delicate balance of positive charge, water-loving and fat-loving components. Manually testing enough candidates to find safe and powerful ones would take years. The authors built a combinatorial library of about 100,000 related polymers called poly(β-amino esters), each designed to self-assemble into nanosized particles that can interact strongly with bacterial cell surfaces.
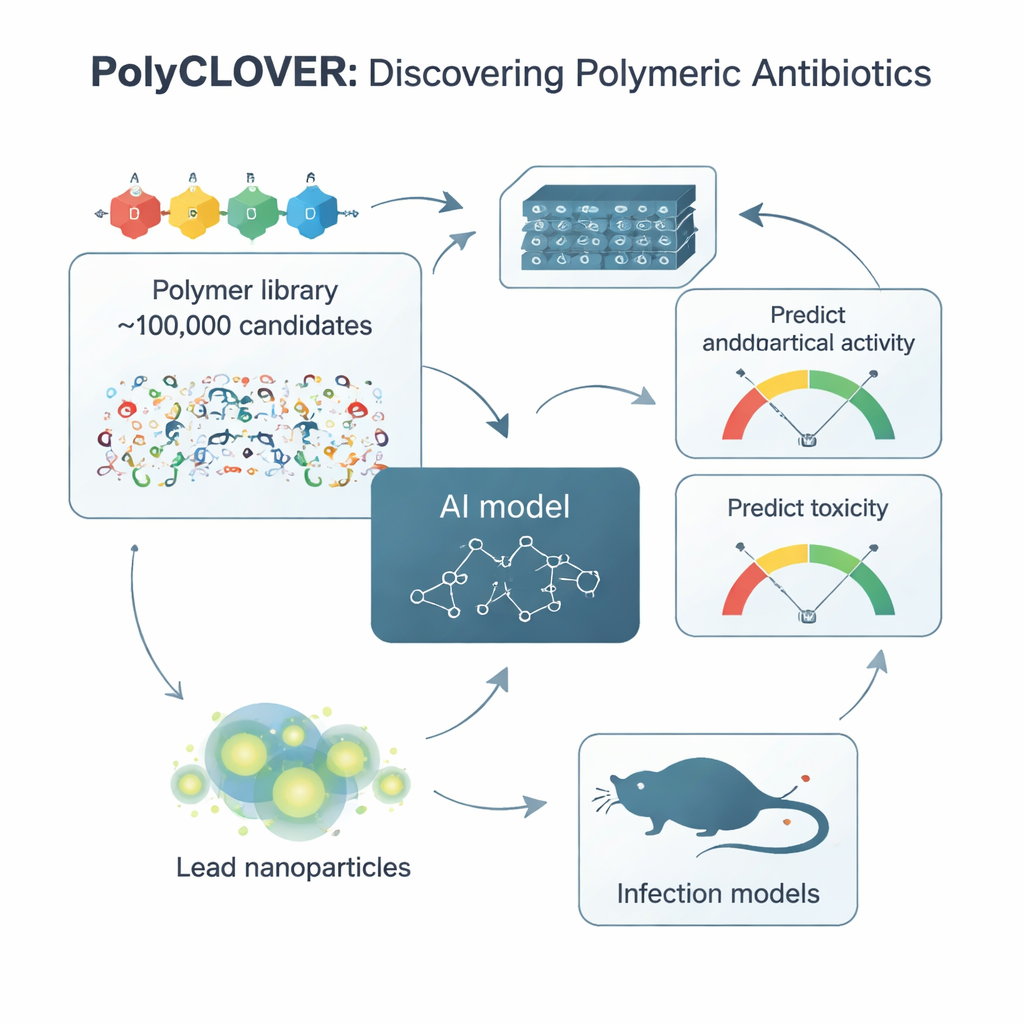
Teaching an AI to read polymer “body language”
To navigate this library, the team created a framework they call PolyCLOVER. At its heart is a graph-based neural network that treats each polymer like a network of connected atoms, allowing the model to sense subtle differences in structure. Because there were very few labeled examples of which polymers were good or bad antibiotics, the researchers used a multi-stage self-supervised learning strategy. First, the model was trained on about a million unlabeled polymer structures to capture general chemical patterns. Then it was further trained on the unlabeled 100,000-member library, and finally fine-tuned on a small set of 220 polymers that had been synthesized and tested for both antibacterial strength and damage to red blood cells. This staged learning dramatically improved the model’s ability to predict which new polymers would be both potent and safe.
Letting experiments and algorithms learn from each other
PolyCLOVER does not stop at a single round of predictions. It runs in a loop where the AI suggests the most promising and most informative candidates, chemists make and test them in high-throughput format, and the new results are fed back to refine the model. The selection step uses an “upper confidence bound” approach that balances exploitation (favoring polymers predicted to work well) with exploration (probing uncertain regions where the model might learn the most). Over about four iterative rounds, the suggested polymers steadily improved: they became more potent against methicillin-resistant Staphylococcus aureus (MRSA) while maintaining acceptable toxicity toward red blood cells. Within only 20 days of experimentation, the system converged on a small set of star performers.
Tiny self-assembling particles that punch holes in bacteria
The final three lead polymers, named H1, H2, and H3, spontaneously formed positively charged nanoparticles about 100 nanometers across. In lab tests, they killed MRSA as effectively as a standard antibiotic, streptomycin, and did so far faster—reducing bacterial counts by several orders of magnitude within minutes to hours, even in protein-rich serum. Electron microscopy and fluorescent probes showed that these particles rapidly latch onto the bacterial surface and disrupt the membrane’s integrity and electrical potential, causing the cell contents to leak out. Importantly, when MRSA was exposed to these polymers for nearly a month, the bacteria developed very little resistance, in stark contrast to streptomycin, whose required dose climbed dramatically over the same period.
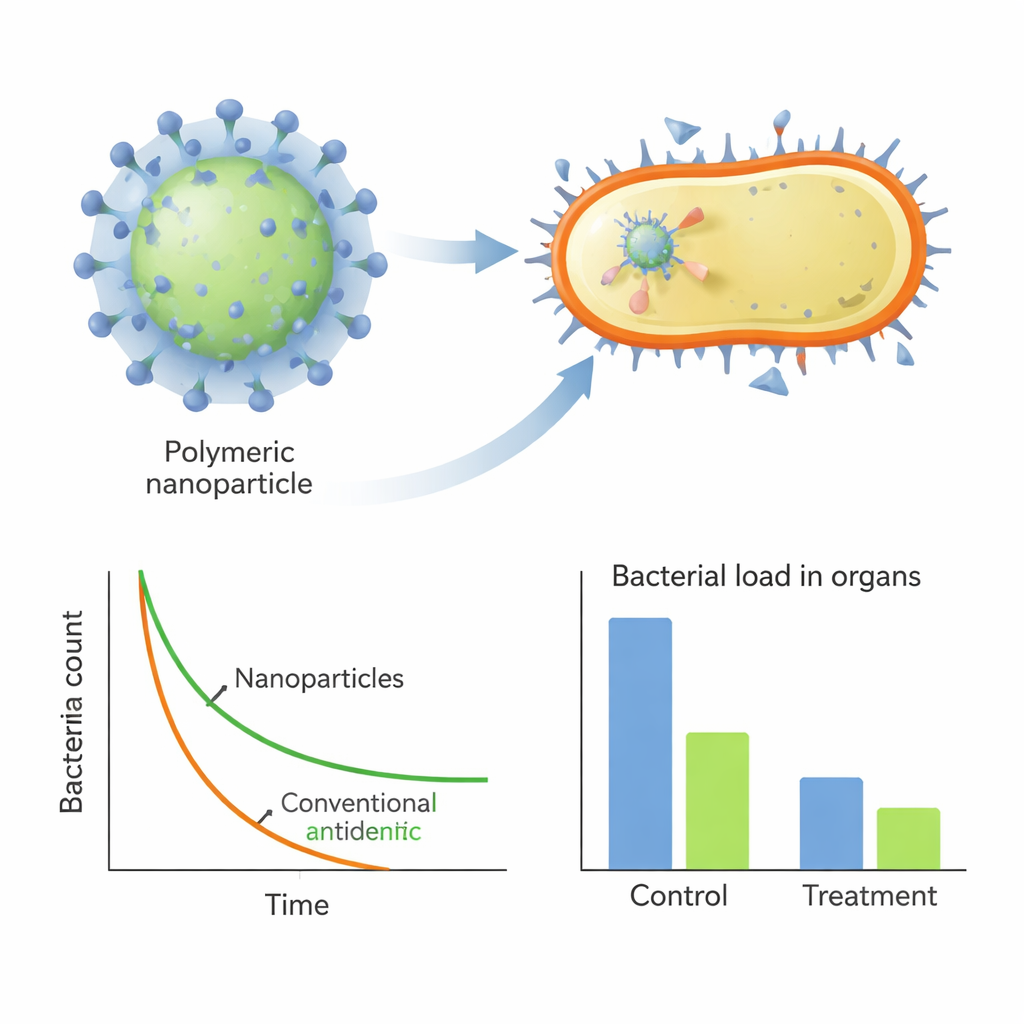
Turning old antibiotics back on in animal infections
Beyond working on their own, the nanoparticles also acted as powerful helpers for traditional drugs. When combined with penicillin G, to which MRSA is normally highly resistant, the best polymer (H2) formed nanocomposites that trapped the antibiotic inside the particle. These composites showed strong synergy in lab tests, slashing the penicillin dose needed to stop MRSA. In mouse models of lung infection and a severe abdominal infection, treatment with the lead polymers—either alone or in combination with penicillin—sharply reduced bacterial loads in organs, lowered inflammatory signals in the blood, and limited tissue damage, all without obvious toxicity.
A new playbook for smarter antibiotic discovery
For non-specialists, the key takeaway is that PolyCLOVER shows how AI and automated experiments can team up to discover entirely new types of antibiotics much faster than before. Instead of relying solely on chance and slow screening, this approach learns from each batch of results to home in on polymers that both attack drug-resistant bacteria and carry conventional antibiotics to them more effectively. While further work is needed before any of these materials reach the clinic, the study offers a promising route to replenish our dwindling arsenal against resistant infections and suggests a general strategy for designing many other smart biomaterials.
Citation: Wu, Y., Wang, C., Shen, X. et al. Iterative discovery of potent polymeric antibiotics via multi-stage and multi-task learning against antimicrobial resistance. Nat Commun 17, 1878 (2026). https://doi.org/10.1038/s41467-026-68682-z
Keywords: antimicrobial resistance, polymeric antibiotics, nanoparticles, machine learning, drug-resistant bacteria