Clear Sky Science · en
Bridge helix of Cas12a is an allosteric regulator of R-loop formation and RuvC activation
Why this matters for gene editing
Many powerful gene-editing tools, including those behind emerging therapies and rapid diagnostic tests, rely on CRISPR proteins that cut DNA at chosen sites. But if these molecular scissors slice the wrong place, they can cause harmful side effects. This study dissects a tiny moving part inside the Cas12a editor, called the bridge helix, and shows how its shape changes act like an internal safety switch that links correct target recognition to DNA cutting. Understanding this switch offers a roadmap to engineer CRISPR tools that are both precise and safer for medical and biotechnology applications.
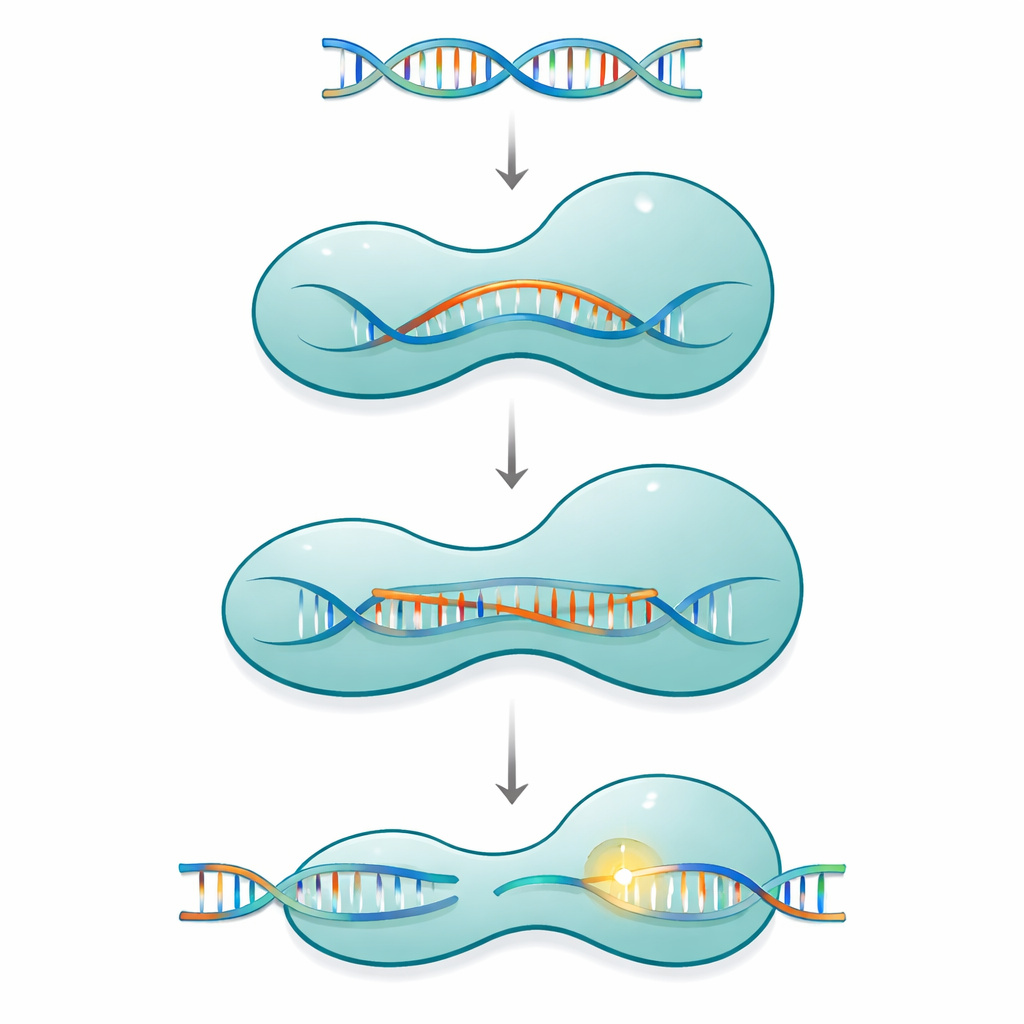
A molecular clamp with moving parts
Cas12a belongs to a family of single-protein CRISPR systems that use a short guide RNA to find and cut matching DNA in viruses or in a cell’s genome. The protein is shaped like a two-armed clamp: a recognition side that checks the sequence, and a cutting side that houses the chemical machinery. These two halves are connected by a slim, positively charged segment called the bridge helix. As Cas12a binds its RNA guide and then its DNA target, the protein does not act as a rigid tool. Instead, it moves through a series of large and small shape changes that gradually close around a growing RNA–DNA hybrid structure known as an R-loop, only fully activating the cutting site once a long enough stretch of correct base pairing has formed.
Probing the built-in safety switch
The researchers focused on a Cas12a from the bacterium Francisella novicida and a previously engineered version in which two amino acids in the bridge helix were replaced by prolines, a change known to stiffen or break helices. This variant, called FnoCas12aKD2P, cuts DNA more selectively but more slowly than the natural protein. Using cryo–electron microscopy, they captured five distinct structural snapshots of this variant bound to its guide RNA and a piece of target DNA. These snapshots show the protein at different stages along its activation pathway, from early initial DNA recognition to later states where part of the RNA–DNA hybrid is formed but full cutting readiness has not yet been achieved.
How shape changes drive accurate cutting
Comparing the variant with previously determined structures of normal Cas12a revealed a clear pattern. In the natural protein, a segment of the bridge helix converts from a loose loop into a longer, straight helix and bends toward the growing RNA–DNA hybrid, coming into close contact with it. At the same time, a neighboring helix in the cutting domain loosens, and a small region called the “lid,” which normally blocks the active site, flips from a loop into a helix and swings open. Together, these coordinated motions open a pocket that allows a single DNA strand to enter and be cut. In the proline-containing variant, the bridge helix cannot fully carry out this loop-to-helix transition or bend properly. As a result, the RNA–DNA hybrid remains distorted and farther away, the lid stays in a closed, loop-like state, and the complex rarely reaches a fully pre-catalytic conformation. This mechanical bottleneck slows on-target cutting and makes it harder for mismatched DNA to sneak through the internal checkpoints.
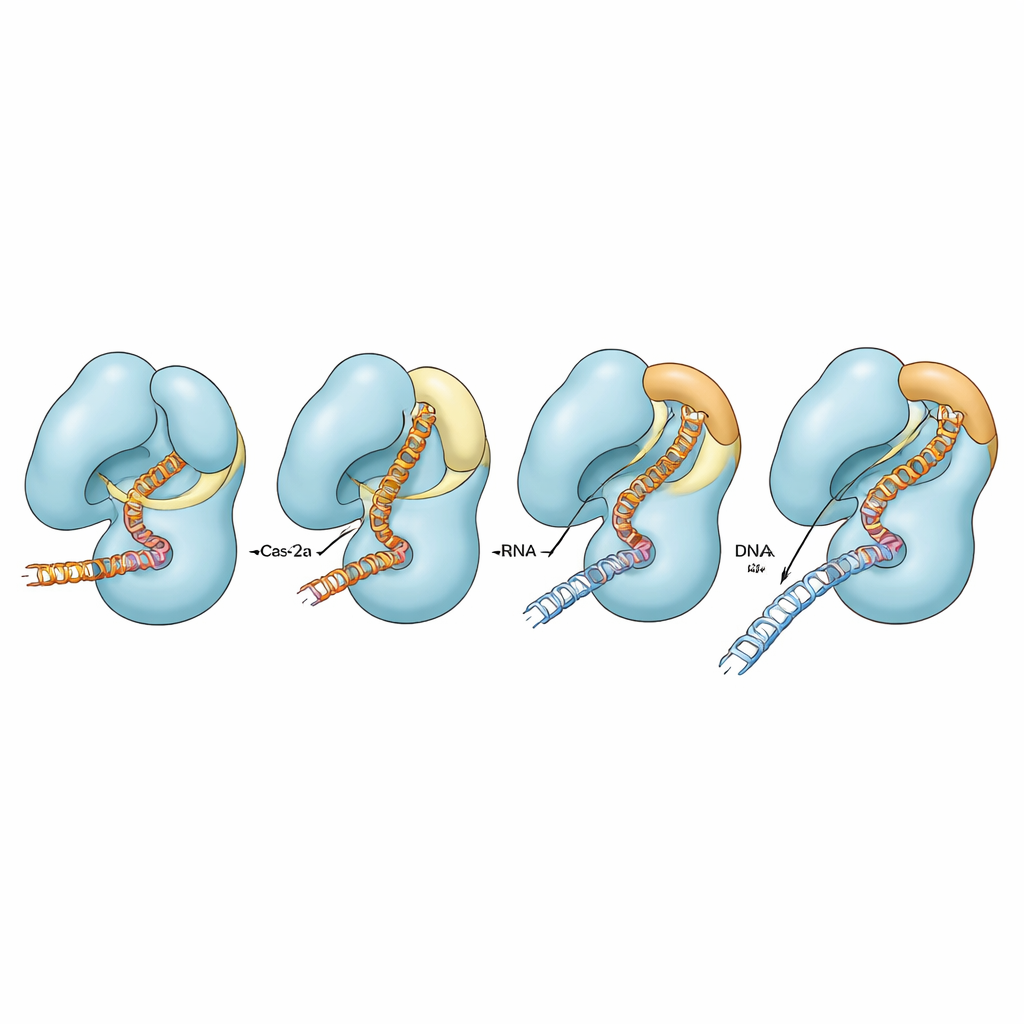
Fine-tuning fidelity by tweaking contacts
To test how different contacts around the lid and bridge helix contribute to activity, the authors changed specific charged residues that link these regions. In the normal Cas12a, breaking these contacts had only modest effects on cutting perfectly matched DNA, though some conditions affected how efficiently the second DNA strand was cleaved when mismatches were present. In the impaired bridge-helix background, however, the same substitutions severely reduced or almost abolished cutting of both strands, especially when guide and DNA did not match perfectly. Computer simulations supported these findings, showing that only when the bridge helix can fully adopt its helical form do the bridge, the adjacent helix, the lid, and the nucleic acids move in a strongly coordinated fashion. When the helix is impaired, these motions become partially or fully uncoupled, and the enzyme struggles to complete its catalytic cycle.
A design guide for next-generation CRISPR tools
Overall, the work reveals the bridge helix as an internal control lever that ties the length and quality of the RNA–DNA hybrid to opening of the cutting pocket in Cas12a. By forcing the protein to complete a specific sequence of shape changes before cleaving DNA, this mechanism naturally enforces mismatch discrimination. The findings help explain why targeted changes in this small region can yield Cas12a variants with reduced off-target cutting and diminished “collateral” DNA activity, properties desirable for safer genome editing and diagnostics. More broadly, the study highlights how subtle shifts in short helices can coordinate long-range communication in large molecular machines, offering general principles for engineering high-fidelity nucleic-acid-processing enzymes.
Citation: Ganguly, C., Aribam, S.D., dos Santos, A.M. et al. Bridge helix of Cas12a is an allosteric regulator of R-loop formation and RuvC activation. Nat Commun 17, 2126 (2026). https://doi.org/10.1038/s41467-026-68657-0
Keywords: CRISPR-Cas12a, gene editing fidelity, bridge helix, R-loop formation, allosteric regulation