Clear Sky Science · en
NeoPrecis: enhancing immunotherapy response prediction through integration of qualified immunogenicity and clonality-aware neoantigen landscapes
Why some cancers respond to immunotherapy and others don’t
Immunotherapy has transformed cancer care, yet many patients still do not benefit and some face serious side effects. A big question is why some tumors are seen and destroyed by the immune system while others slip past. This study introduces NeoPrecis, a computational method that looks more closely at the “flags” tumors display to the immune system—called neoantigens—and uses that information to better predict which patients are likely to respond to modern immunotherapies.
New flags on cancer cells
Cancer cells accumulate DNA mutations that can change the proteins they make. Small fragments of these altered proteins, known as neoantigens, can be displayed on the cell surface and recognized as foreign by T cells, the immune system’s assassins. For years, doctors and researchers have relied on coarse measures such as tumor mutation burden—the total number of mutations—to estimate how likely a cancer is to respond to immune checkpoint inhibitors. But this is a blunt tool: not every mutation creates a visible or attractive target for T cells, and tumors can be patchworks of many different cell subtypes. NeoPrecis was designed to move beyond simply counting mutations and instead judge how many of them are genuinely promising targets across the whole tumor.
Looking at three key ingredients at once
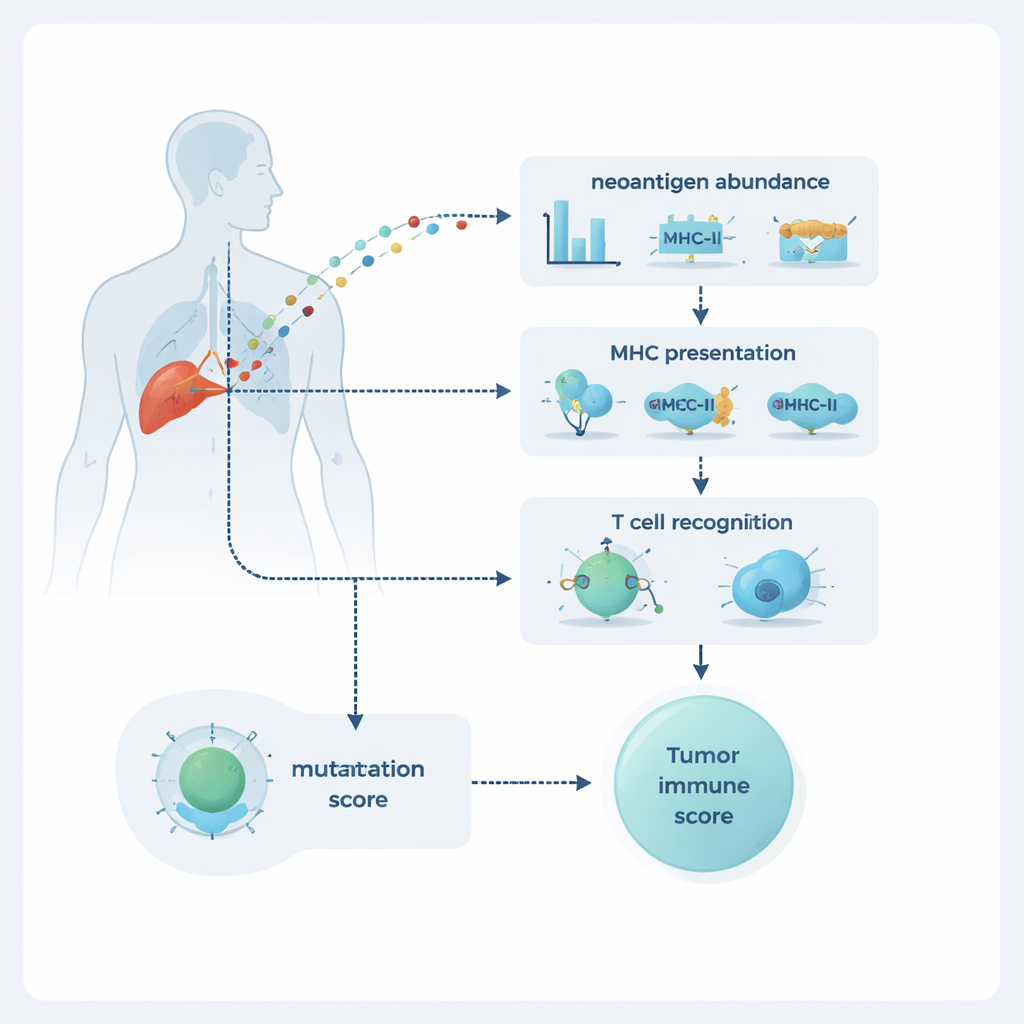
NeoPrecis evaluates each mutation along three dimensions: how abundant it is in the tumor, whether it is likely to be displayed on the cell surface, and how likely it is to be noticed by T cells. Abundance is estimated from DNA and RNA sequencing, which reveal how common the mutation is and how strongly it is expressed. Display is modeled through binding to molecules called MHC class I and class II, which act like billboards showing protein fragments to T cells. The most novel part is the T cell recognition component, NeoPrecis-Immuno. This model learns from large databases of known T cell–peptide interactions to gauge how different a mutated fragment is from its normal counterpart in a way that matters for T cell recognition, while also accounting for the specific MHC variants each person carries.
Teaching a computer what T cells “see”
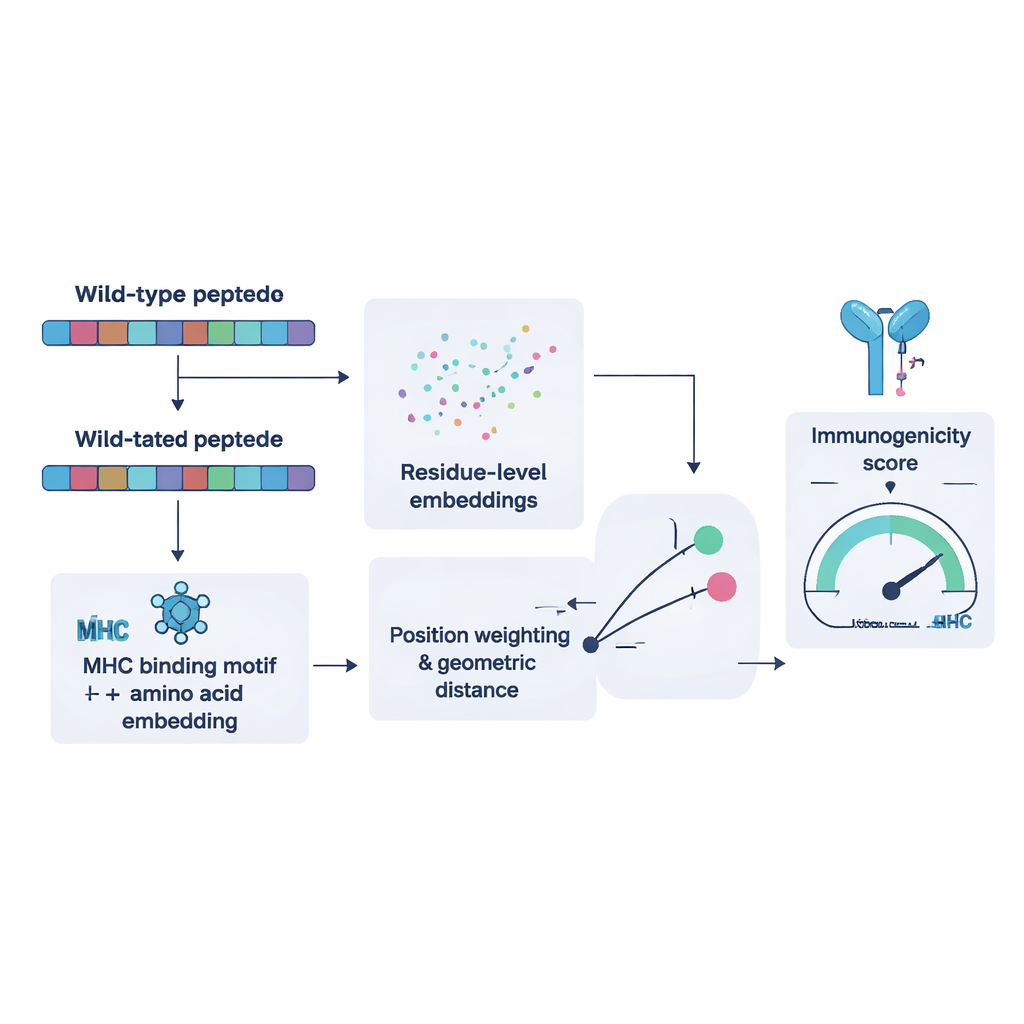
To train NeoPrecis-Immuno, the researchers first assembled thousands of examples where the same T cell recognizes multiple similar peptides and others it does not. They used these to teach the model that mutant fragments closely resembling normal ones are less likely to trigger an immune attack, because such self-like targets are usually filtered out during T cell development. The model represents each peptide as a point in a mathematical space shaped by both its amino acid sequence and the binding preferences of the person’s MHC molecules. It then measures how far the mutant peptide lies from the original. Larger, motif-informed distances correspond to a higher chance of being immunogenic. When tested against existing tools on independent cancer datasets, NeoPrecis-Immuno matched or outperformed them, especially when working with MHC class II, which presents to helper T cells that support and sustain anti-tumor responses.
From single mutations to the whole tumor
Individual mutations are only part of the story; how they are distributed within the tumor also matters. Some mutations are “clonal,” found in nearly every cancer cell, while others are “subclonal,” present only in certain pockets. NeoPrecis builds a “neoantigen landscape” by summing its immunogenicity scores across mutations and layering in information about which subclones they belong to and how common those subclones are. This produces tumor-level scores that highlight cancers rich in strong, widely shared neoantigens—especially those that can be presented on both MHC class I and class II molecules and potentially trigger coordinated helper and killer T cell responses. In patient groups with melanoma and non‑small cell lung cancer treated with checkpoint inhibitors, these NeoPrecis-based scores more accurately separated responders from non‑responders than standard mutation counts, and were particularly helpful in complex, heterogeneous lung tumors.
What this means for patients
For patients, the promise of NeoPrecis is more precise matching of immunotherapies to those most likely to benefit, and a clearer understanding of why some tumors resist treatment. By focusing on the quality and distribution of neoantigens—not just their quantity—the framework helps explain why tumors with similar mutation loads can behave very differently. In the future, such detailed maps of a tumor’s immune-visible landscape could guide not only checkpoint inhibitor use, but also the design of personalized cancer vaccines that prioritize the most potent and broadly shared targets within each person’s cancer.
Citation: Lee, KH., Sears, T.J., Zanetti, M. et al. NeoPrecis: enhancing immunotherapy response prediction through integration of qualified immunogenicity and clonality-aware neoantigen landscapes. Nat Commun 17, 1966 (2026). https://doi.org/10.1038/s41467-026-68651-6
Keywords: cancer immunotherapy, neoantigens, tumor heterogeneity, checkpoint inhibitors, computational oncology