Clear Sky Science · en
Engineering the MmeFz2-ωRNA system for efficient genome editing through an integrated computational-experimental framework
Smaller Gene Scissors for Big Medical Problems
Gene editing has already changed how scientists study diseases, but turning it into safe, practical medicine is still a major challenge. One big obstacle is that today’s most powerful DNA “scissors,” like CRISPR-Cas9, are physically large and hard to package into the tiny delivery vehicles used in patients. This study introduces a new, compact gene-editing tool built from a little-known family of enzymes called Fanzor2 and shows how artificial intelligence can rapidly redesign it into a strong candidate for future genetic therapies.
A New Class of Compact Gene Editors
Most current genome editors, such as Cas9 and Cas12a, are bulky proteins made of more than a thousand building blocks. That size makes them difficult to squeeze into adeno-associated viruses (AAVs), the leading delivery system for gene therapies. In contrast, Fanzor2 proteins are much smaller and are naturally found in eukaryotes, the branch of life that includes animals and humans. The authors focused on one member of this family, MmeFz2, which uses a short RNA molecule called ωRNA to find and cut specific DNA sequences. On paper, MmeFz2 looked ideal for therapy: compact and programmable. In practice, however, its activity in human cells was dismal, editing less than one percent of target sites. The team set out to transform this underperforming enzyme into a practical tool.
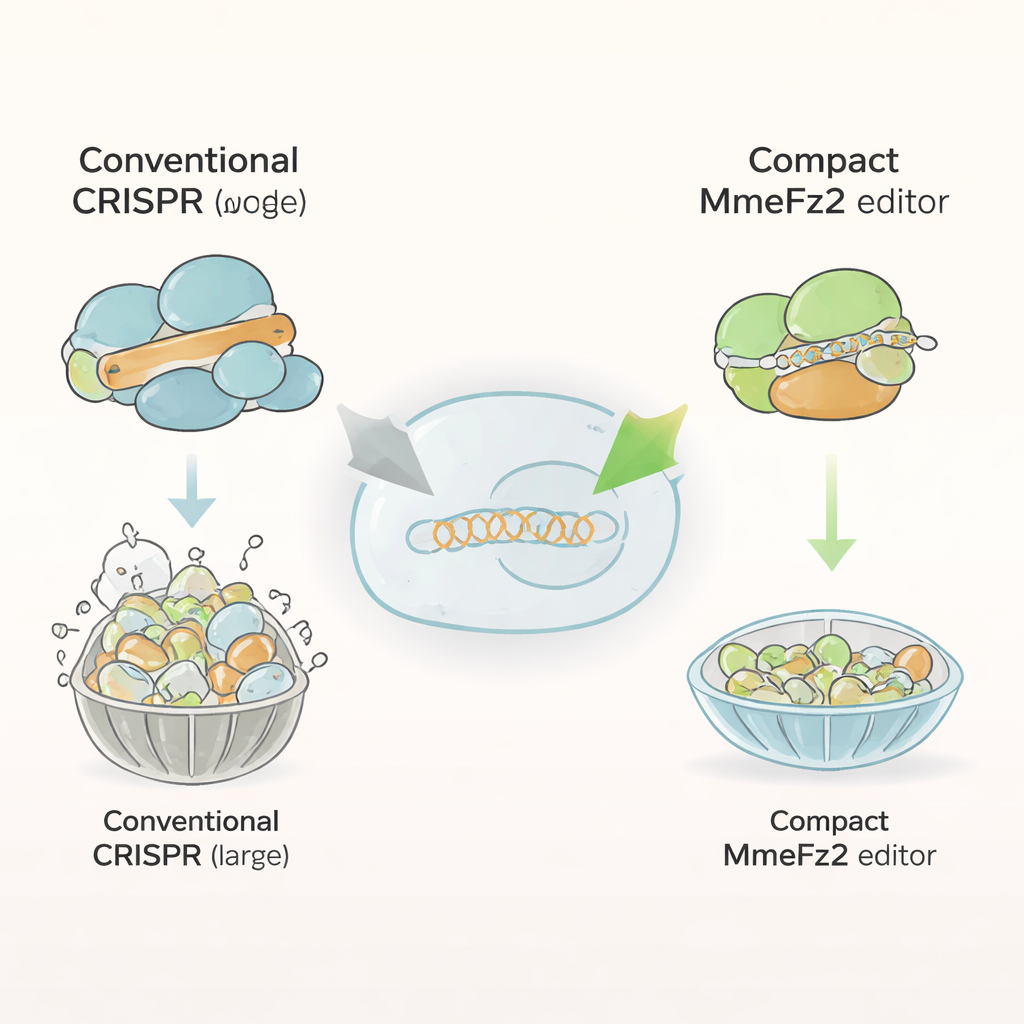
Using AI to Redesign the Guide Molecule
The first target was the ωRNA, the guide that tells MmeFz2 where to cut. Using AlphaFold3, a cutting-edge program that predicts 3D structures of protein–RNA–DNA complexes, the researchers built a model of MmeFz2 bound to its ωRNA and a DNA target. This virtual structure revealed that parts of the ωRNA were floppy, poorly paired, and barely touching the protein. Guided by this information, the team systematically replaced unstable links in the RNA and removed strings of uridine bases that can interfere with transcription. They then tested dozens of variants in human cells. By stacking the best changes and trimming an unnecessary stem region, they created a shortened ωRNA that was 30% smaller yet delivered close to a 20-fold boost in editing efficiency across many DNA sites.
Fine-Tuning the Protein with Machine Learning
Next, the team refined the protein itself. They again used AlphaFold3 to map where MmeFz2 touched RNA and DNA and designed more than a hundred single amino-acid changes aimed at strengthening those contacts or improving flexibility. After screening these in cells, they fed the performance data into EVOLVEpro, a machine-learning system built on protein language models trained from millions of natural sequences. This tool learned which features tended to increase activity and suggested new combinations of mutations predicted to work even better. Through a few rounds of this computational–experimental loop, the scientists arrived at two optimized enzyme variants that, when paired with the improved ωRNA, raised editing activity more than 60-fold relative to the starting system.
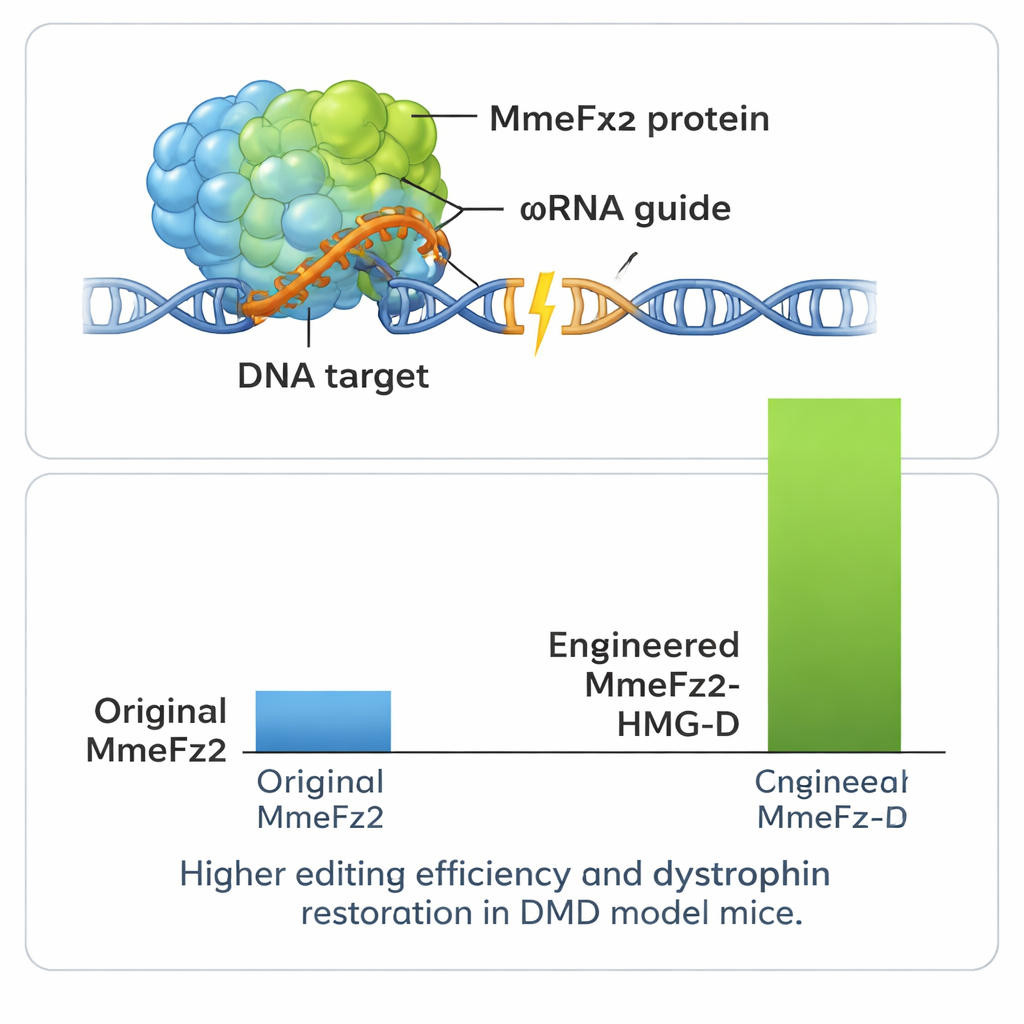
Boosting Power and Testing in a Disease Model
To squeeze even more performance out of the system, the authors fused MmeFz2 to small DNA-binding domains that help proteins latch onto genetic material. One of these, called HMG-D, proved especially effective when attached to the end of the engineered enzyme, producing editing levels above 80% at some sites. Importantly, tests at predicted off-target locations in the genome showed that this added power did not come with a major loss of precision. Because the entire gene editor—including protein and guide RNA—is so compact, it fits comfortably into a single AAV vector, something that is not possible for most CRISPR systems. The team put this advantage to use in a mouse model of Duchenne muscular dystrophy, a lethal muscle-wasting disorder caused by errors in the dystrophin gene. A one-time injection of the AAV-delivered editor into leg muscle led to detectable corrections in the gene’s RNA and restored dystrophin protein in a substantial fraction of muscle fibers.
What This Means for Future Gene Therapies
For non-specialists, the key message is that the authors have turned a weak, obscure gene-cutting enzyme into a compact, high-performance editor that can be delivered in a single viral package and can repair a serious genetic disease in a realistic animal model. Just as important is how they got there: by tightly combining AI-based structure prediction and machine learning with focused laboratory tests, they improved both the protein and its guide RNA far more efficiently than traditional trial-and-error approaches. This integrated strategy may accelerate the creation of many next-generation genome editors, bringing precise and safer gene therapies closer to patients with muscular dystrophy and other inherited conditions.
Citation: Li, S., Xu, K., Li, G. et al. Engineering the MmeFz2-ωRNA system for efficient genome editing through an integrated computational-experimental framework. Nat Commun 17, 1867 (2026). https://doi.org/10.1038/s41467-026-68644-5
Keywords: gene editing, CRISPR alternatives, Duchenne muscular dystrophy, viral gene therapy, artificial intelligence in biology