Clear Sky Science · en
Multicountry genomic analysis underscores regional cholera spread in Africa
Why following germs across borders matters
Cholera still sickens and kills tens of thousands of people each year in Africa, yet basic questions remain about how outbreaks start, move between countries, and keep coming back. This study brings together scientists and public health teams from seven African nations to track the bacterium that causes cholera by reading its genetic code. By comparing hundreds of bacterial genomes, the researchers show how recent waves of cholera have flowed across borders, which kinds of strains are circulating where, and how this knowledge can sharpen efforts to stop future epidemics.
Taking a continent-wide look at cholera
To move beyond scattered case reports, the Africa Centres for Disease Control and Prevention launched a collaboration called the Cholera Genomics Consortium in Africa (CholGEN). Laboratories in Cameroon, the Democratic Republic of the Congo, Malawi, Mozambique, Nigeria, Uganda, and Zambia sequenced 763 high-quality genomes of the Vibrio cholerae O1 bacterium, mostly from 2019 to 2024. This represents the largest set of cholera genomes ever generated within Africa itself. By placing these new genomes alongside nearly 1,800 previously sequenced samples from Africa and Asia, the team could reconstruct how recent African outbreaks fit into the long-running global cholera pandemic.
Old strains, new routes
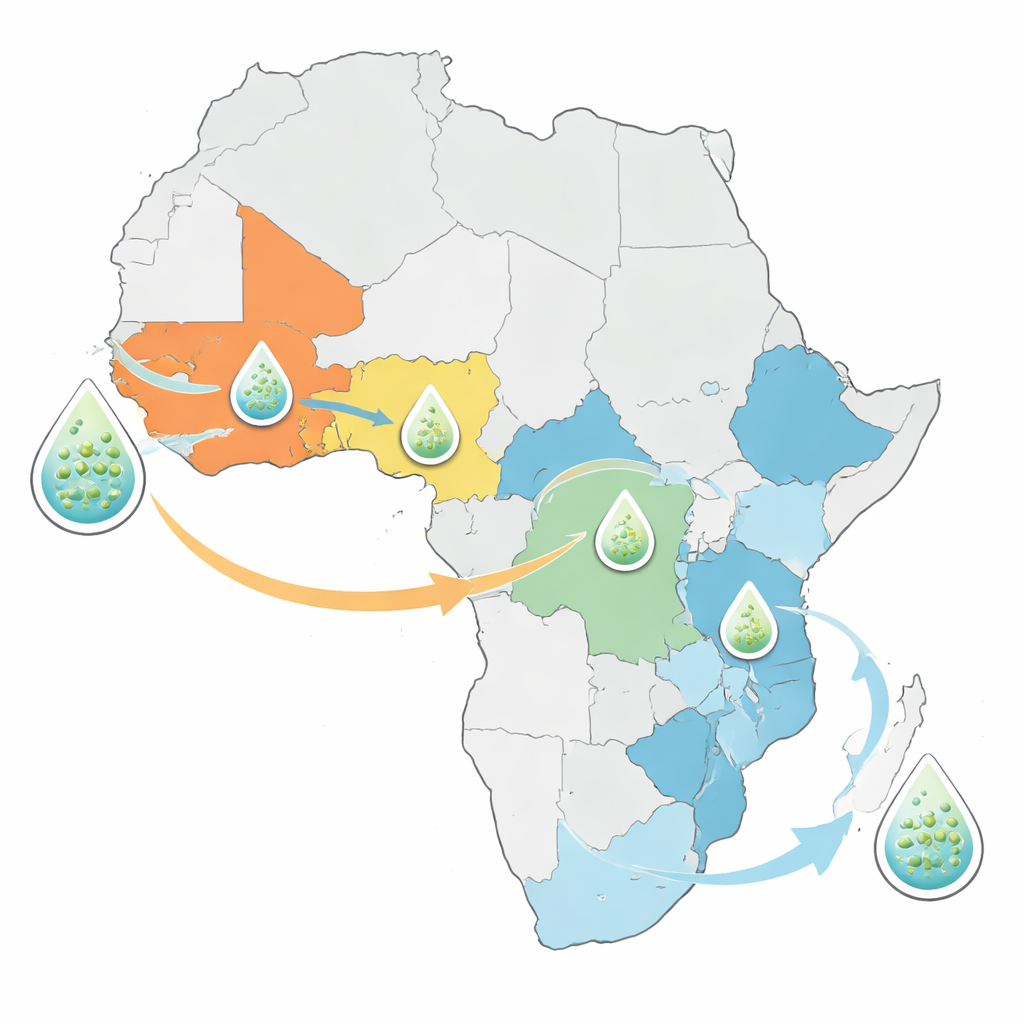
The analysis showed that the cholera strains causing today’s African outbreaks do not represent an entirely new threat. Instead, all of the newly sequenced bacteria descend from previously known introductions of the seventh-pandemic cholera lineage that first reached Africa from Asia in 1970. Western and Central African countries such as Nigeria, Cameroon, and the Democratic Republic of the Congo tend to be dominated by one or two long-lived lineages that have persisted for decades. Eastern and Southern African countries, by contrast, harbor a mix of several lineages at once. One lineage in particular, dubbed AFR15, has spread rapidly in recent years and is linked to unusually large outbreaks in Malawi, Zambia, and neighboring countries, as well as to epidemics in parts of the Middle East and South Asia.
When size of outbreaks is not in the genes
One might suspect that AFR15’s explosive spread is due to major genetic changes that make it more dangerous or better able to evade treatment. However, when the researchers compared the rate and pattern of mutations across several active lineages, they found no striking differences. The bacteria were evolving at similar speeds, and the types of mutations and affected genes looked much the same from one lineage to another. Overall profiles of antibiotic resistance genes were also mostly stable over time and between countries. The main exception was in Uganda, where bacteria acquired a large mobile DNA element called a plasmid that carries multiple resistance genes, likely imported along with strains linked to outbreaks in Yemen and Lebanon. Even so, the study did not find new genes that would explain the severity of recent African outbreaks on their own.
Hidden journeys revealed by better sampling
Because the bacterial genomes carry a record of where closely related strains were found, the team could infer how often cholera crosses borders. They detected many examples of international spread among neighboring countries, including repeated exchanges between Zambia and the Democratic Republic of the Congo. Yet when they looked more closely, they saw that statistical signals of cross-border jumps were strongest in years and places with intensive sampling. This suggests that real-world movements of cholera are more frequent than the current data reveal; many transmission events likely go unnoticed simply because no one is sequencing bacteria in those places or times. To address this, the authors developed a framework to estimate how much new information a country gains from sequencing additional samples, balancing genetic diversity, number of introductions, and existing data.
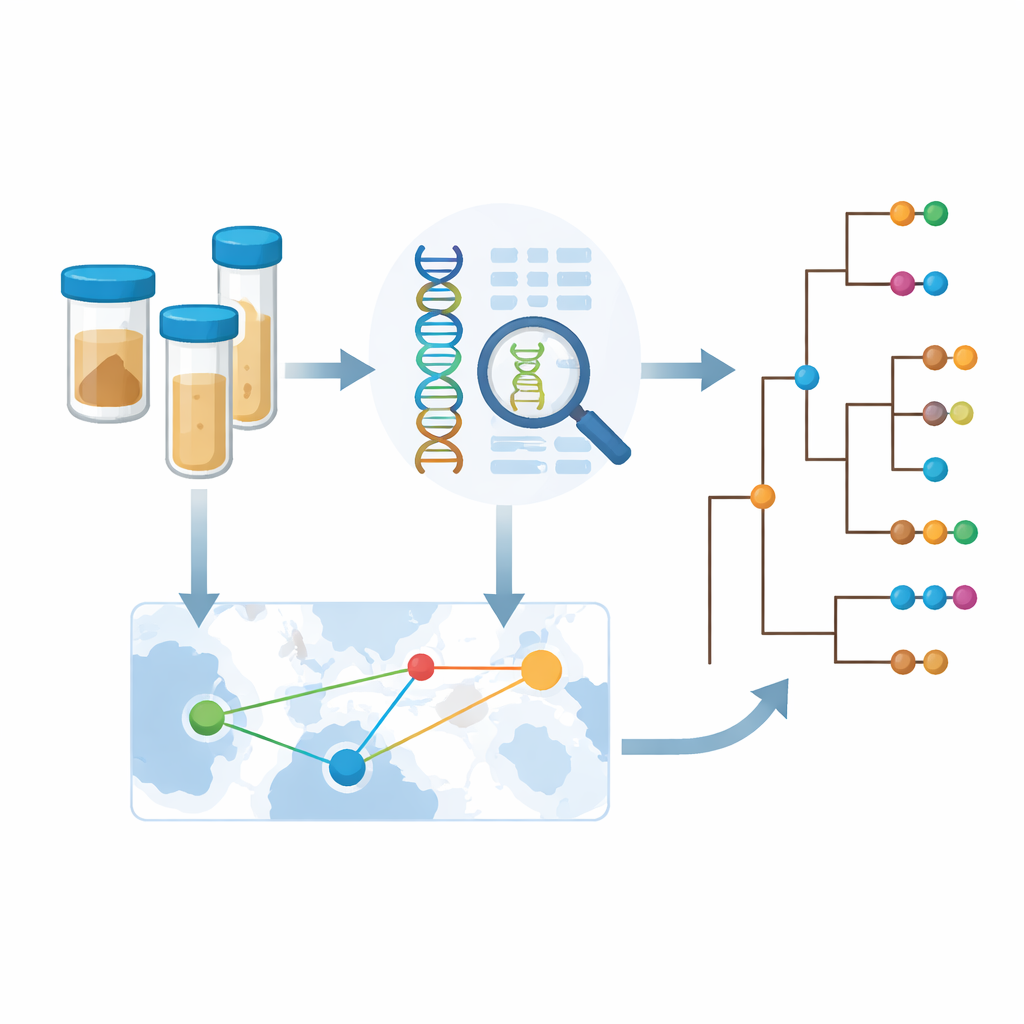
Using genomics to guide smarter control efforts
The study concludes that, for cholera in Africa today, how and where the disease is spreading matters more than any dramatic change in the bacterium itself. The findings argue for routine, regionally coordinated genomic surveillance so that neighboring countries can spot shared outbreaks early, trace their sources, and target interventions such as vaccination campaigns, water and sanitation improvements, and border-area responses more efficiently. By building sequencing capacity within African public health laboratories and sharing data across borders, initiatives like CholGEN provide a practical blueprint for using modern genetics to move closer to the ambitious goal of eliminating cholera as a public health threat by 2030.
Citation: Mboowa, G., Matteson, N.L., Tanui, C.K. et al. Multicountry genomic analysis underscores regional cholera spread in Africa. Nat Commun 17, 2539 (2026). https://doi.org/10.1038/s41467-026-68642-7
Keywords: cholera, genomic surveillance, Africa, cross-border transmission, antimicrobial resistance