Clear Sky Science · en
Dual genomic localizations and gene regulatory functions of MBD-2 with and without NuRD in Caenorhabditis elegans which lacks DNA methylation
How tiny worms rewrite the rules of gene control
Our cells use chemical tags on DNA and proteins to remember which genes should be on or off. One of the most famous tags, DNA methylation, is missing in some animals—yet they still grow and reproduce normally. This paper explores how a small worm, Caenorhabditis elegans, manages gene control without DNA methylation, revealing an unexpectedly flexible backup system that may reshape how we think about epigenetics.
A missing DNA mark raises a big question
In mammals, a chemical mark called 5‑methylcytosine (5mC) helps silence genes during development, X‑chromosome inactivation, and defense against jumping genes. Proteins called MBD2 and MBD3 read these marks and recruit a large protein machine, the NuRD complex, which can reshape and de‑activate stretches of chromatin. Strangely, many invertebrates, including C. elegans and fruit flies, have lost 5mC and the enzymes that write it—but they kept an MBD2/3‑like protein. This raises a puzzle: why keep a “DNA methylation reader” in a genome that no longer uses DNA methylation, and what does it do instead?
A scaffold protein that worms cannot live without
The authors focused on the worm version of this protein, called MBD‑2. Unlike its mammalian cousins, worm MBD‑2 has lost the classic methyl‑binding domain that recognizes 5mC, but it retains flexible and coiled segments that can latch onto other proteins. By tagging MBD‑2 with fluorescent markers, the team showed that it sits in the nuclei of almost all cells throughout the worm’s life, consistent with a broad role in gene regulation. Using genetic engineering, they then created worms completely lacking MBD‑2, or lacking only its coiled‑coil region, which mediates contact with NuRD. Both mutations caused severe problems: animals were small, moved poorly, developed malformed reproductive structures, and were largely sterile. This demonstrated that MBD‑2 is essential for normal development and fertility, even in the absence of DNA methylation.
Rebuilding the NuRD machine in a methylation‑free world
To see whether worm MBD‑2 still works with NuRD, the authors pulled down tagged MBD‑2 from worm extracts and identified its partners by mass spectrometry. Most of the known NuRD components were present, confirming that MBD‑2 serves as a core scaffold in this complex, much as MBD2/3 does in mammals. When the coiled‑coil region was deleted, many of these interactions were lost, especially those with proteins that remodel nucleosomes. At the same time, large‑scale RNA sequencing revealed that more than a quarter of all worm genes changed their activity in MBD‑2 mutants, with many more genes turning on than off. This pattern indicates that MBD‑2, often together with NuRD, generally acts as a repressor to keep inappropriate genes quiet, while also helping to maintain proper activity of a subset of highly expressed genes.
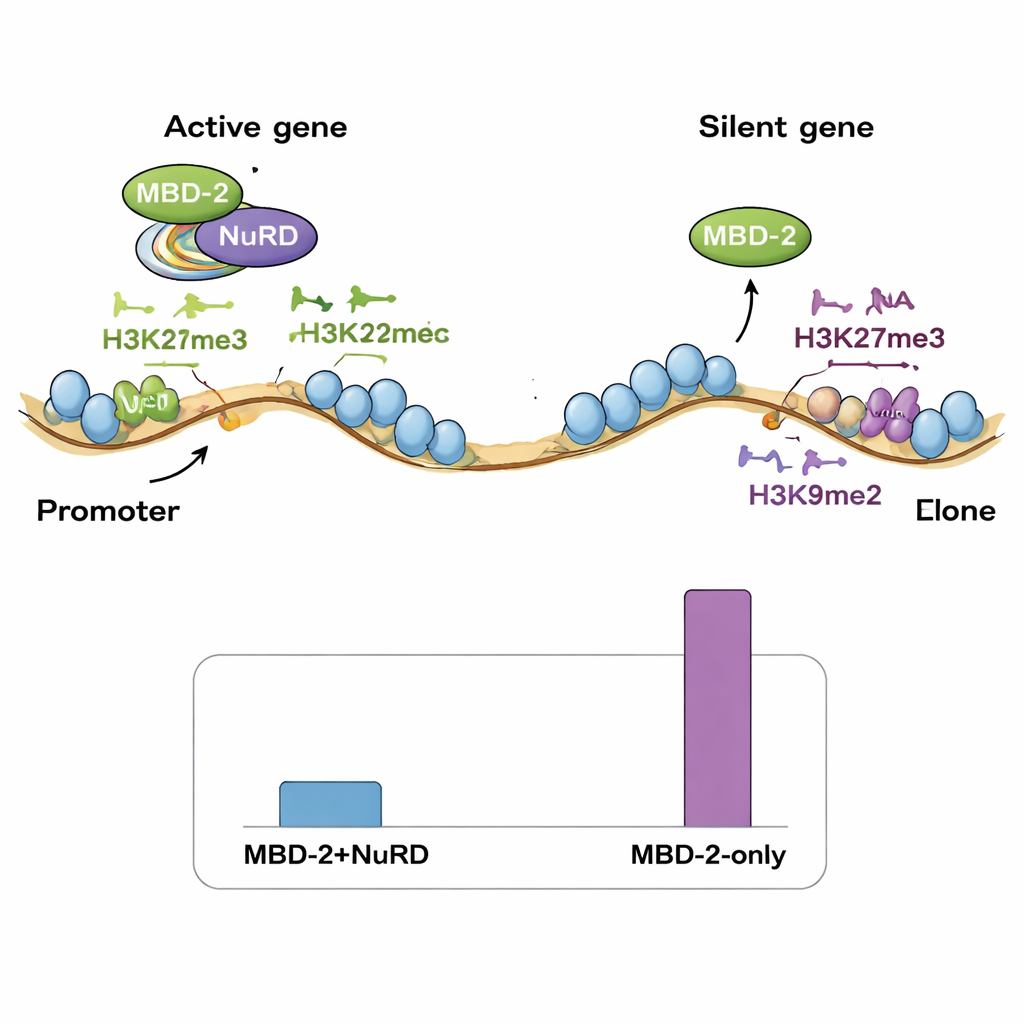
Two binding modes on the genome
Next, the team mapped where MBD‑2 sits along the genome using ChIP‑seq, and compared these locations with existing maps of NuRD components and various histone marks. Surprisingly, only a small fraction of MBD‑2 sites overlapped tightly with NuRD proteins and marks of open, active chromatin. The vast majority of MBD‑2 sites lay elsewhere, in chromosomal “arms” rich in repressive histone tags such as H3K27me3 and H3K9me2/3. In other words, MBD‑2 shows a dual behavior: at a minority of sites it travels with NuRD to fine‑tune active genes, but at many more sites it binds independently in regions that are already silent. The pattern of its binding within genes also matters—when MBD‑2 clusters near gene start sites, loss of the protein tends to activate those genes, whereas binding deeper in gene bodies is often linked to reduced expression when MBD‑2 is removed.
An evolutionary backup for missing DNA methylation
The authors propose that, in species that lost DNA methylation, repressive histone marks—especially H3K27me3—may have expanded to fill a similar regulatory niche. Worm MBD‑2 appears to have been repurposed: instead of reading 5mC on DNA, it now associates with chromatin regions marked by specific histone modifications, while still anchoring the NuRD complex through its conserved protein‑interaction domains. This work shows that an ancient gene‑silencing toolkit can be rewired over evolution: the chemical tag on DNA can disappear, but the associated protein machinery survives by switching to alternative signals. For non‑specialists, the key message is that gene regulation is remarkably adaptable—cells can lose a major epigenetic mark and yet maintain complex control of thousands of genes by leaning more heavily on other chromatin tags and the versatile scaffolding roles of proteins like MBD‑2.
Citation: Tsui, H.N., Wong, C.Y.Y., Zheng, C. et al. Dual genomic localizations and gene regulatory functions of MBD-2 with and without NuRD in Caenorhabditis elegans which lacks DNA methylation. Nat Commun 17, 1875 (2026). https://doi.org/10.1038/s41467-026-68592-0
Keywords: epigenetics, chromatin, gene regulation, C. elegans, histone modification