Clear Sky Science · en
A highly dynamic mononuclear non-heme iron enzyme for the two-step isonitrile biosynthesis
How Bacteria Build Exotic Chemical Tools
Isonitriles are small but powerful chemical groups that behave like Swiss Army knives: they can grab metals, react in many different ways, and often act as the “warhead” of potent natural antibiotics and toxins. Some disease-causing bacteria, including the tuberculosis pathogen Mycobacterium tuberculosis, stitch isonitriles onto fatty molecules to help them steal metals from their host. This article explores how one of their enzymes, called Rv0097, pulls off this unusually demanding bit of chemistry in two carefully choreographed steps.
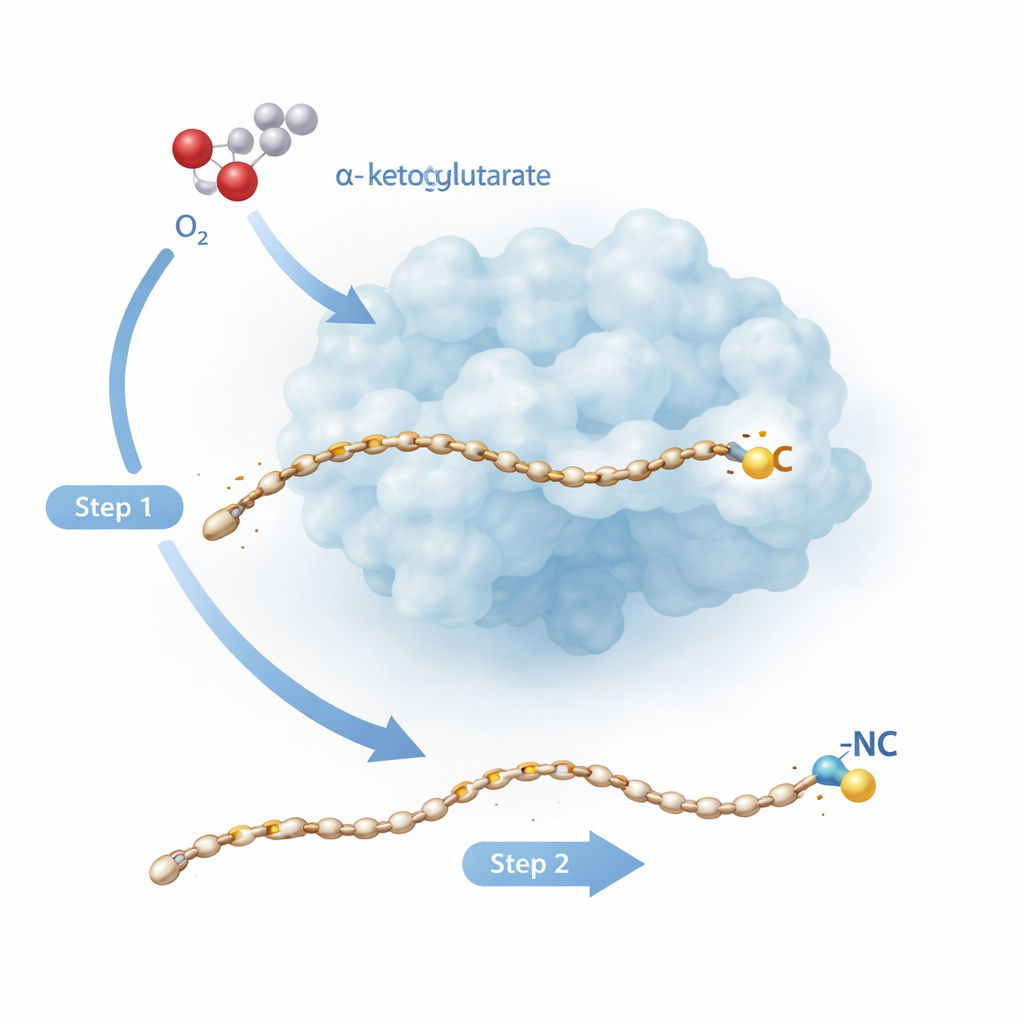
A Two-Step Molecular Makeover
Rv0097 belongs to a large family of iron-dependent enzymes that normally carry out simpler tasks like adding oxygen to a single spot on a molecule. Here, the task is tougher: starting from a “glycyl–fatty acid” building block, the enzyme must create a brand‑new isonitrile group, with both of its key atoms—the carbon and the nitrogen—coming from the same small glycine fragment. Earlier work on a related enzyme, ScoE, suggested that this cannot be done in one shot. Instead, two separate reaction “half‑cycles” are needed, each powered by an iron–oxygen species formed when the enzyme tears apart a helper molecule called α‑ketoglutarate and combines it with oxygen from the air. The new study turns to the tuberculosis enzyme Rv0097 to watch, at atomic resolution, how an enzyme can string these two radical steps together on the same substrate without losing control.
Freezing the Enzyme in the Act
Using X‑ray crystallography, the researchers captured sixteen high‑resolution snapshots of Rv0097 in different states: empty, bound to its fatty substrate (a ten‑carbon molecule called CADA), bound to the α‑ketoglutarate helper, and in combinations that mimic fleeting reaction intermediates. These structures show that Rv0097 has a split personality. At rest, its active site is mostly closed off, with a key amino acid (phenylalanine 102) flipping between two orientations that keep the long fatty tail pocket dry and ready for hydrophobic binding. When CADA binds, this residue snaps into a single orientation and the substrate’s polar “head” is anchored by charged side chains while its oily tail nestles into a snug tunnel. Mutating this gating pair of residues (F102 and a nearby glycine, G204) either blocks the pocket or lets in too much water, sharply reducing activity and changing which chain lengths the enzyme prefers.
Moving Lids and Dynamic Loops
Beyond the substrate pocket, Rv0097 behaves like a tiny machine with moving doors. The team observed “open” and “closed” conformations in which two lid‑like regions swing apart or together over the active site. Opening exposes the iron center and creates a pathway for α‑ketoglutarate and substrate to enter; closing shields the chemistry from the surrounding water once the reaction is under way. Two flexible surface loops, called the His loop and Arg loop after the histidine and arginine they carry, act as an additional gate. In some structures, these loops swing inward so that the arginine residue can grab α‑ketoglutarate; in others they swing outward, apparently creating a side channel through which spent helper fragments (succinate and carbon dioxide) can leave and a fresh α‑ketoglutarate can slip in for the second step—all while the substrate‑derived intermediate remains clamped in place.
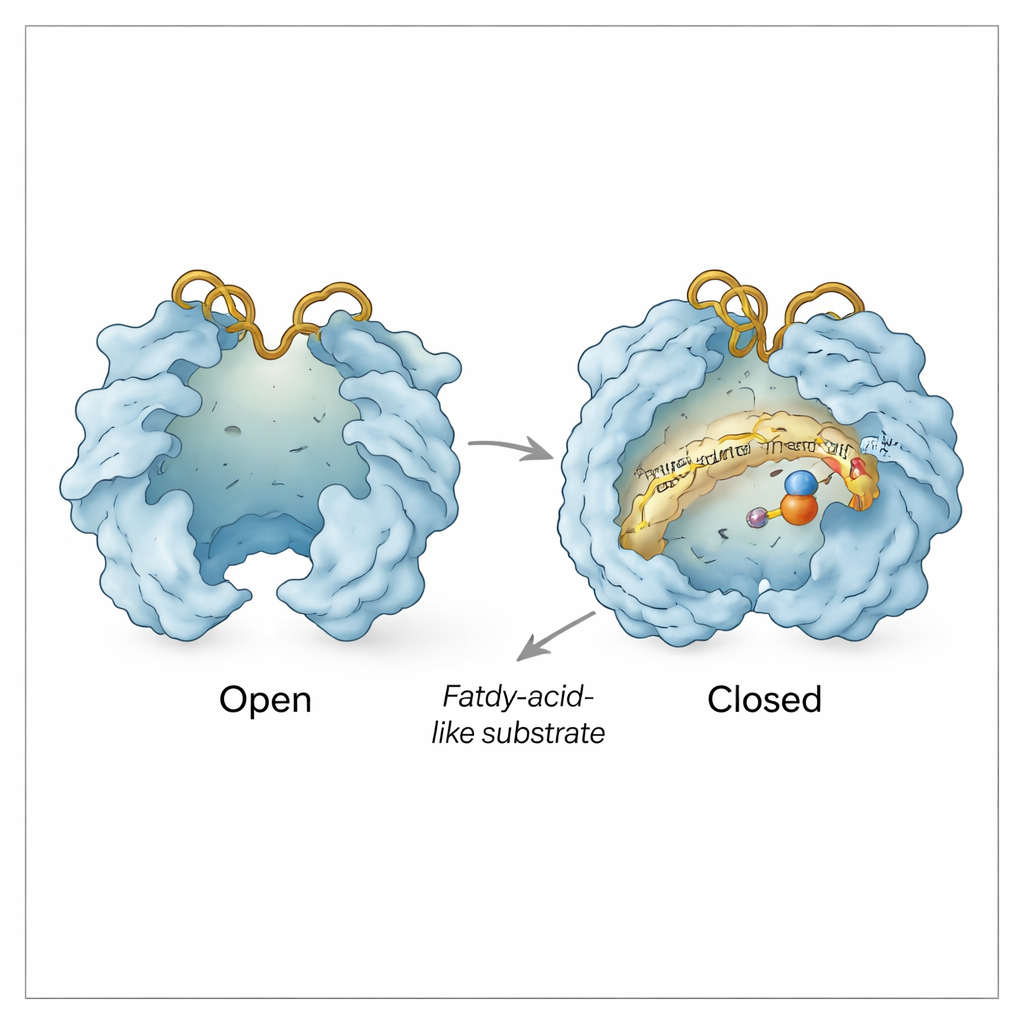
Protecting a Fragile Intermediate
Biochemical assays support this structural picture. The first half of the reaction converts CADA to a highly reactive imine intermediate that would quickly fall apart if it drifted into solution. By chemically trapping its breakdown product, the authors show that most of this intermediate is only detected when the enzyme is still present, implying it stays bound and protected inside Rv0097 between the two half‑reactions. The structures explain how: subtle repositioning of the His and Arg loops, along with small shifts of nearby residues, appear to seal the intermediate into a sheltered cavity while still allowing small molecules like α‑ketoglutarate, succinate, and carbon dioxide to exchange through controlled channels.
Why This Matters for TB and Beyond
Together, these results reveal Rv0097 as a highly dynamic, single‑protein assembly line that uses iron‑based chemistry twice in a row on the same substrate without ever releasing a dangerous halfway product. For tuberculosis bacteria, this precision underpins the manufacture of isonitrile‑decorated molecules that help them obtain essential metals inside the body, and genetic studies indicate the pathway is important for their survival during infection. For chemists and drug designers, the work offers a blueprint for engineering related enzymes to build new isonitrile‑containing compounds, and suggests that interfering with Rv0097’s moving lids and gates could be a way to develop new antibiotics.
Citation: Ye, N., Del Rio Flores, A., Zhang, W. et al. A highly dynamic mononuclear non-heme iron enzyme for the two-step isonitrile biosynthesis. Nat Commun 17, 2034 (2026). https://doi.org/10.1038/s41467-026-68588-w
Keywords: isonitrile biosynthesis, non-heme iron enzyme, Mycobacterium tuberculosis, enzyme dynamics, metal-chelating natural products