Clear Sky Science · en
HIF sustain a transcriptional regulatory circuit of EPAS1 expression in renal clear cell carcinoma
Why kidney cancer needs a closer look
Clear cell renal cell carcinoma is the most common form of kidney cancer, and many patients now receive drugs that block a protein called HIF‑2α. These medicines can shrink tumors, but they do not work equally well for everyone. This study asks a simple but important question: what makes some kidney tumors produce so much HIF‑2α, and can understanding this "on switch" help us predict who will benefit most from treatment and how to design new therapies?
The oxygen sensor that goes rogue
Healthy kidney cells use a safety protein called VHL to keep oxygen-sensing factors, known as HIFs, under tight control. When oxygen is plentiful, VHL marks HIF proteins for destruction, preventing them from turning on genes that promote blood vessel growth and cell division. In clear cell kidney cancer, VHL is usually lost or damaged. As a result, one HIF family member, HIF‑2α (encoded by the EPAS1 gene), escapes destruction and accumulates. Earlier work showed that high HIF‑2α activity is linked to faster tumor growth and poorer outcomes, and that blocking HIF‑2α can slow or stop tumors in some patients. However, the molecular steps that boost EPAS1 gene activity in kidney tumors were not well understood.
A hidden DNA switch for a cancer gene
Using tumor samples from patients, single-cell data, and cancer cell models, the authors show that HIF‑2α messenger RNA is markedly higher in clear cell kidney tumors than in normal kidney tissue or other kidney tumor types. They then scan the EPAS1 region of the genome for chemical and structural marks that signal active regulatory DNA. This reveals a powerful enhancer—a kind of long-distance gene switch—located about 70,000 DNA letters upstream of EPAS1 that is specifically active in clear cell tumors. In tumor cells, the chromatin at this enhancer is open, decorated with activation marks, and physically loops to contact the EPAS1 promoter, while in normal kidney tubule cells this region is largely silent. Activity at this enhancer correlates with higher EPAS1 levels in large cancer datasets. 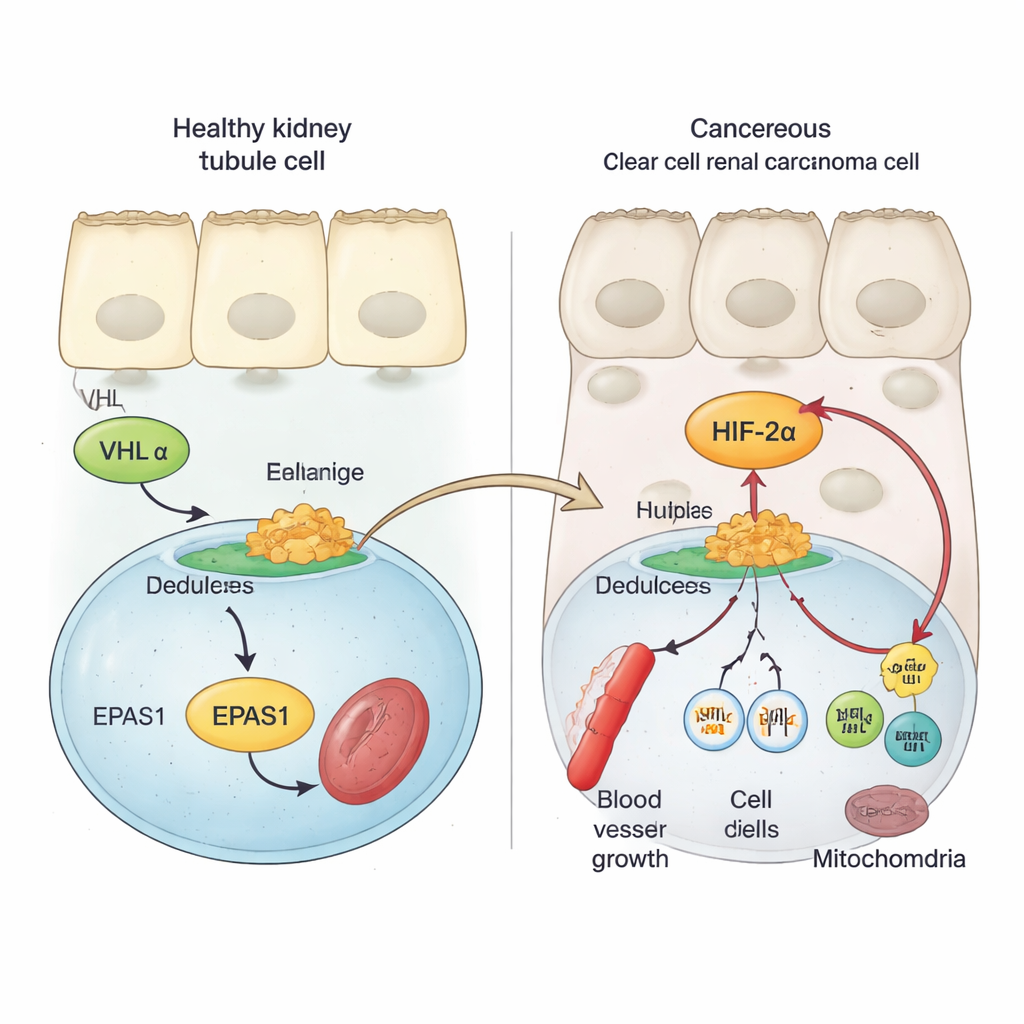
A self-reinforcing loop driven by kidney identity factors
The study finds that HIF proteins do more than respond to low oxygen—they actually help drive their own production. When the researchers restore VHL or remove a key HIF partner protein (HIF‑1β), EPAS1 levels fall and the enhancer becomes less accessible. Direct mapping of protein–DNA contacts shows that HIF binds to two enhancer regions upstream of EPAS1, including the clear cell–specific enhancer. In kidney tumor cells, this enhancer is also occupied by PAX8 and HNF‑1β, transcription factors that help define kidney cell identity and are known to support tumor growth. Disrupting PAX8 or HNF‑1β, or mutating their binding sites within the enhancer, reduces HIF‑2α RNA and protein and weakens enhancer activity. Together, these findings reveal an auto-regulatory circuit: HIF‑2α, working with kidney lineage factors, boosts the activity of an EPAS1 enhancer, which in turn drives more HIF‑2α production.
Genetic risk and tumor behavior converge on the same circuit
The EPAS1 region has long been flagged in genome-wide studies as a hotspot for kidney cancer risk, with certain inherited variants linked to higher cancer rates and to VHL mutations in tumors. By combining genetic data with gene expression measurements, the authors show that individuals carrying the high-risk version of a key EPAS1 variant tend to have higher HIF‑2α levels in their tumors—and even in normal kidney tubule cells when HIF is experimentally stabilized. This suggests that inherited DNA differences can tune how strongly the enhancer responds, priming some kidneys to ramp up HIF‑2α more once VHL is lost. Experiments in glioblastoma cells show that the same enhancer can also be engaged in certain brain tumors, hinting that this regulatory module may be reused in other cancers where HIF‑2α is important. 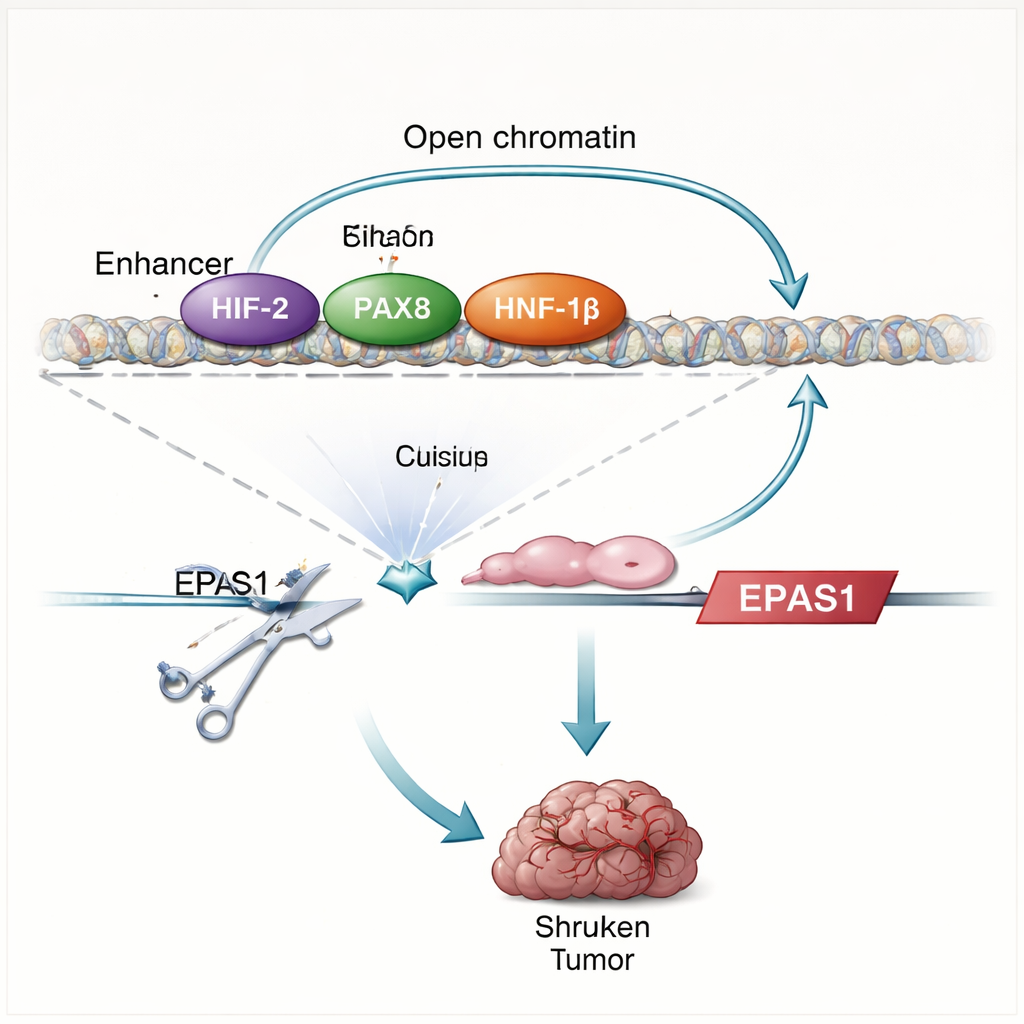
Breaking the loop to stop tumor growth
To test how essential this enhancer is, the team uses CRISPR genome editing to damage the HIF binding motifs within it in kidney cancer cell lines and fresh patient tumor cells. This cut reduces HIF‑2α RNA by about one-third to one-half, lowers the levels of well-known HIF‑2α target genes such as CCND1 and VEGFA, and weakens the broader hypoxia gene program. When edited cells are implanted into mice, they largely fail to form tumors, in stark contrast to control cells. The pattern of gene changes after enhancer disruption closely mirrors the effects of directly blocking HIF‑2α with a clinical drug, underscoring that this single DNA element is a major driver of the HIF‑2α pathway in these cancers.
What this means for patients and therapies
For non-specialists, the key message is that the authors have uncovered a self-amplifying DNA switch that keeps a major cancer-promoting protein, HIF‑2α, turned on in clear cell kidney tumors. This switch depends both on the loss of the VHL safety brake and on kidney-specific helper proteins, and it is influenced by inherited genetic variants. Because the enhancer is highly active in many clear cell tumors and is tightly linked to how strongly the HIF‑2α pathway runs, measuring its activity—or targeting it directly—could help identify patients who will benefit most from current HIF‑2α inhibitors and could offer new ways to shut down this pathway when tumors become drug-resistant.
Citation: Naas, S., Krüger, R., Grampp, S. et al. HIF sustain a transcriptional regulatory circuit of EPAS1 expression in renal clear cell carcinoma. Nat Commun 17, 1764 (2026). https://doi.org/10.1038/s41467-026-68576-0
Keywords: clear cell renal cell carcinoma, HIF-2α, EPAS1 enhancer, VHL mutation, kidney cancer genetics