Clear Sky Science · en
A dual context-aware basecaller for nanopore direct RNA sequencing
Why decoding RNA letters matters
Every cell in your body is constantly reading and rewriting messages written in RNA, the working copy of our genes. New “nanopore” machines can read individual RNA molecules directly, promising to reveal how genes are switched on, how RNAs are spliced, and how chemical marks on RNA influence health and disease. But there is a catch: these devices actually measure tiny electrical currents, which then must be translated—“basecalled”—into the familiar A, C, G and U letters. If that translation is wrong, the biological story we infer can be badly distorted. This paper introduces Coral, a new artificial‑intelligence system that makes this translation much more accurate.
Reading electricity instead of letters
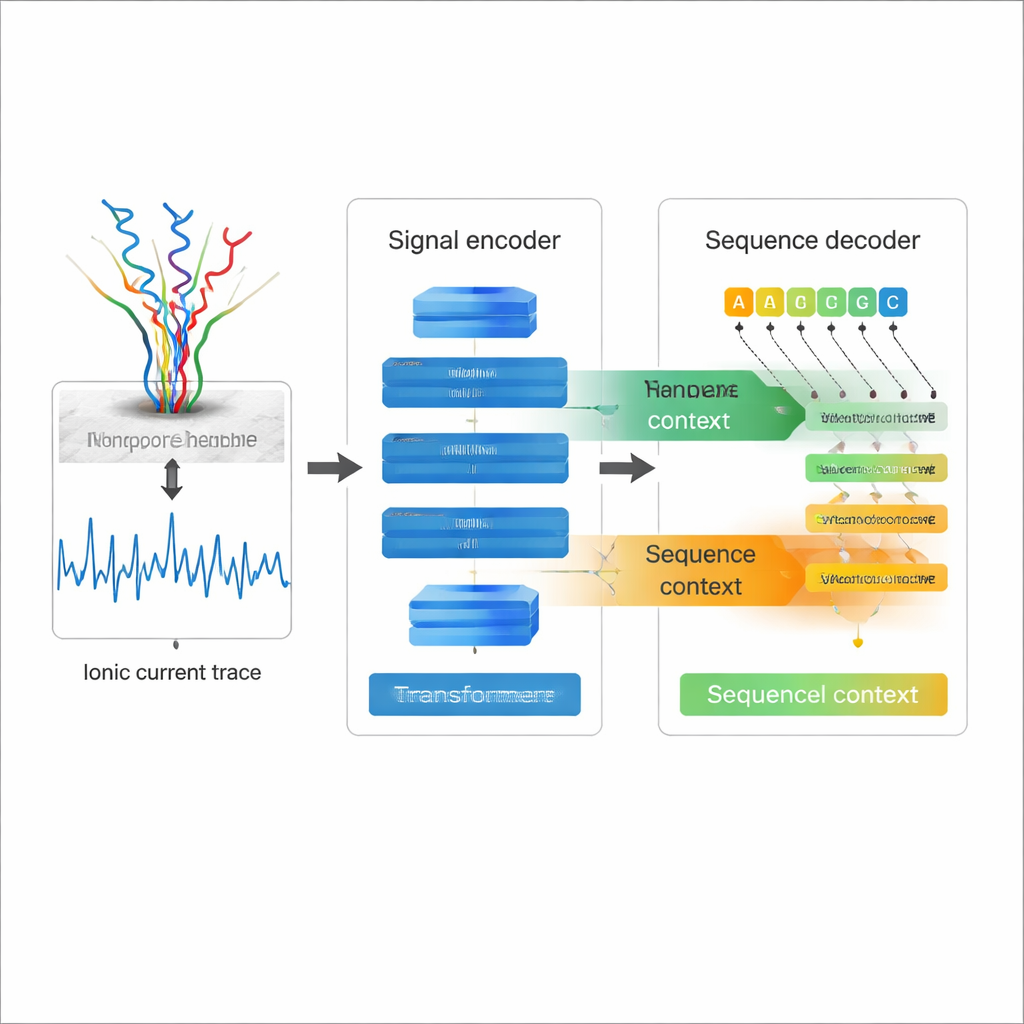
Nanopore direct RNA sequencing works by threading a single RNA strand through a molecular hole— a nanopore—while measuring how the electric current changes as each nucleotide passes. Those wiggly current traces contain the information about the RNA sequence and its chemical modifications. Traditional RNA sequencing instead converts RNA into DNA and amplifies it, steps that can introduce bias and erase many natural chemical marks. Direct RNA sequencing avoids those problems, but the price has been a relatively high error rate when turning current traces into sequences, especially for challenging features like repeated bases and complex RNA shapes. Better basecalling is essential if scientists want to trust the fine details of these long RNA reads.
A smarter translator that uses two kinds of context
Most existing nanopore basecallers treat the electrical signal as the main source of information and decode each position almost independently, which limits how well they can use the structure of the RNA sequence itself. Coral takes a different approach. It uses a Transformer-based encoder–decoder architecture, similar in spirit to modern language models. First, an encoder network built from convolutions and self‑attention layers digests the raw current signal into a compact description of how the signal changes over time. Then a decoder predicts each new RNA base one step at a time, simultaneously looking backward at the bases it has already written and sideways at the encoded signal. Two kinds of attention—within the growing RNA sequence and between sequence and signal—allow Coral to weigh both electrical and sequence context when deciding which letter comes next.
Sharper sequences and fewer missed molecules
The authors tested Coral against several leading basecallers, including Oxford Nanopore’s commercial tools, on RNA from humans and other organisms and on multiple nanopore chemistries. Across six species and older RNA sequencing kits, Coral achieved a typical median read accuracy around 97%, clearly higher than competing methods. With the latest RNA kit, its accuracy exceeded 99%. Coral produced fewer mismatches, insertions and deletions, and yielded longer, better‑aligned reads with fewer sequences that could not be mapped at all. It was especially good at handling short runs of repeated bases—very common in real data—which are a frequent source of errors for other tools. By more reliably capturing longer stretches of correct sequence, Coral also excelled at predicting short sequence patterns (k‑mers) and remained robust even when earlier decoding steps contained small mistakes.
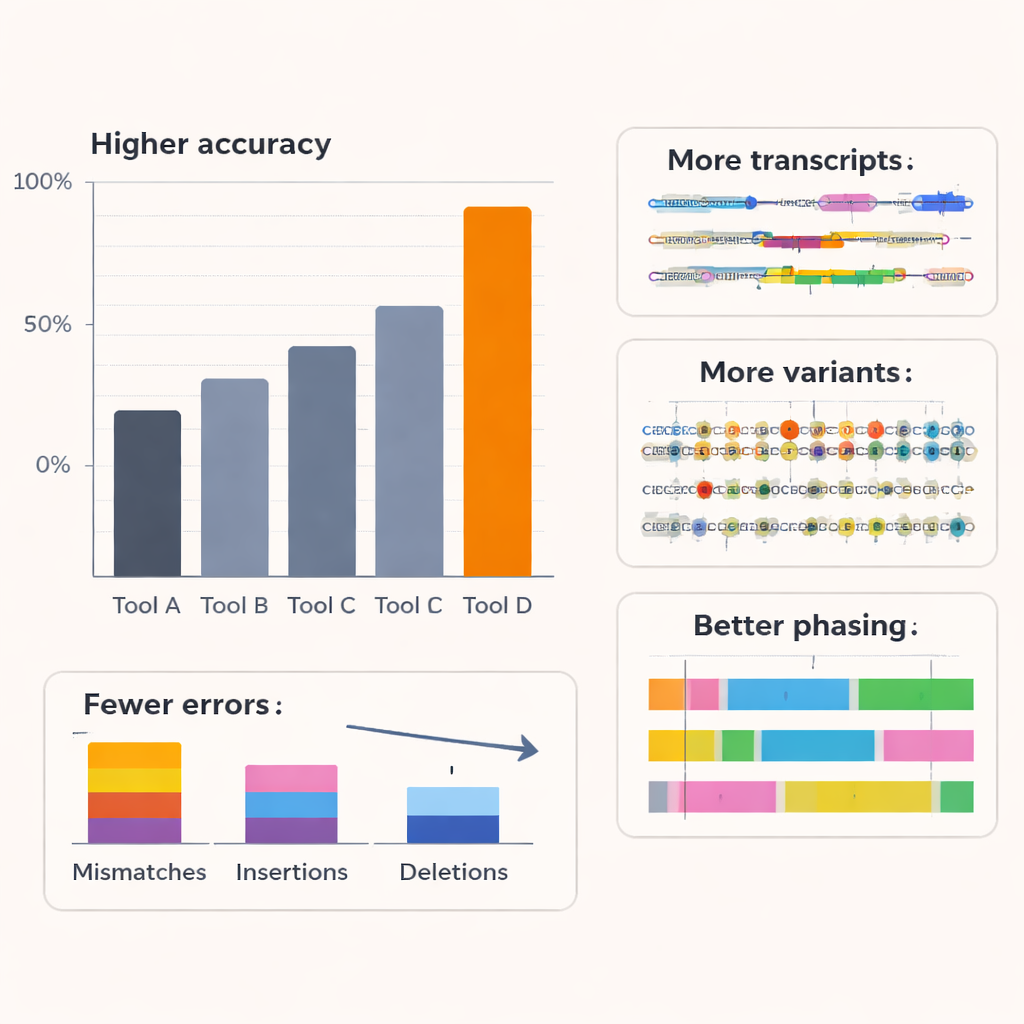
Seeing more of the transcriptome’s hidden detail
Improved basecalling is valuable only if it leads to better biology. To test this, the team examined how Coral’s output affected downstream analyses in human cell lines. Using a specialized tool to reconstruct full RNA isoforms—the different splice versions of each gene—they found that Coral’s reads exposed more known transcript structures and many additional, low‑abundance isoforms that other basecallers missed. Many Coral‑specific transcripts were supported by independent short‑read data, indicating they are real rather than artifacts. Coral also detected more artificial reference transcripts with known concentrations in a spike‑in experiment and estimated their abundance more accurately. Beyond transcript discovery, Coral improved the detection of gene‑fusion events in a breast‑cancer cell line and increased the number and reliability of genes showing allele‑specific expression, where one parental copy of a gene is more active than the other.
Clearer genetic variants and family lines
Because long RNA reads can span distant genetic variants, they are powerful tools for determining which variants travel together on the same chromosome copy—a process called haplotype phasing. Using a well‑studied human sample with a gold‑standard variant map, the authors showed that Coral’s higher‑quality reads led to more accurate detection of single‑nucleotide changes and far fewer phasing errors: switch errors and overall mismatch rates within phased blocks dropped by up to about three‑quarters compared with other methods, while substantially more variants could be phased at all. Simulation studies varying the underlying read accuracy confirmed that once basecalling approaches about 95% accuracy, performance in transcript discovery, allele‑specific expression, and phasing improves sharply and then plateaus. Coral sits in this high‑benefit zone, suggesting it captures most of the biologically relevant information present in the noisy nanopore signals.
What this means for future RNA research
For non‑specialists, the key message is that Coral acts like a far more reliable translator between the electrical language of nanopore sequencers and the genetic language of RNA. By better using context in both the signal and the growing sequence, it produces cleaner reads that uncover more transcript variants, spot rare fusion genes, and more confidently track which variants come from which parent. The software is open‑source, so researchers can adapt it to new organisms, chemistries, or even to study chemical marks on RNA itself. As nanopore technology continues to improve, tools like Coral will help turn raw current traces into trustworthy, detailed maps of the RNA world inside cells.
Citation: Xie, S., Ding, L., Yu, Y. et al. A dual context-aware basecaller for nanopore direct RNA sequencing. Nat Commun 17, 1851 (2026). https://doi.org/10.1038/s41467-026-68566-2
Keywords: nanopore RNA sequencing, basecalling, Transformer model, transcript isoforms, haplotype phasing