Clear Sky Science · en
Breaking the cell wall for efficient DNA delivery to diatoms
Algae as Tiny Green Factories
Diatoms—microscopic algae that drift in the oceans—quietly make about one‑fifth of the planet’s organic carbon, helping feed marine food webs and lock away carbon dioxide. Scientists would like to turn these hardy, fast‑growing organisms into tiny green factories for making fuels, foods, and specialty chemicals. But there has been a major roadblock: it is surprisingly hard to get new DNA into diatom cells reliably. This study tackles that practical problem head‑on, describing new ways to slip genetic instructions and genome‑editing tools past the diatom’s tough outer wall.
Softening the Shell of Ocean Microbes
The work centers on the model diatom Phaeodactylum tricornutum, a favorite in labs because its genome is well mapped and it already has some basic genetic tools. The authors reasoned that the cell wall, the first physical barrier any DNA must cross, was a key bottleneck. By treating cells with an enzyme called alcalase, they partially or fully removed this wall, creating fragile “spheroplasts” and “protoplasts” that are far easier to penetrate. When they then used electroporation—brief electric pulses that open temporary pores—the number of successful transformants jumped dramatically, by roughly two orders of magnitude compared with earlier methods. Even tiny amounts of DNA, as little as one nanogram, were enough to recover engineered cells. 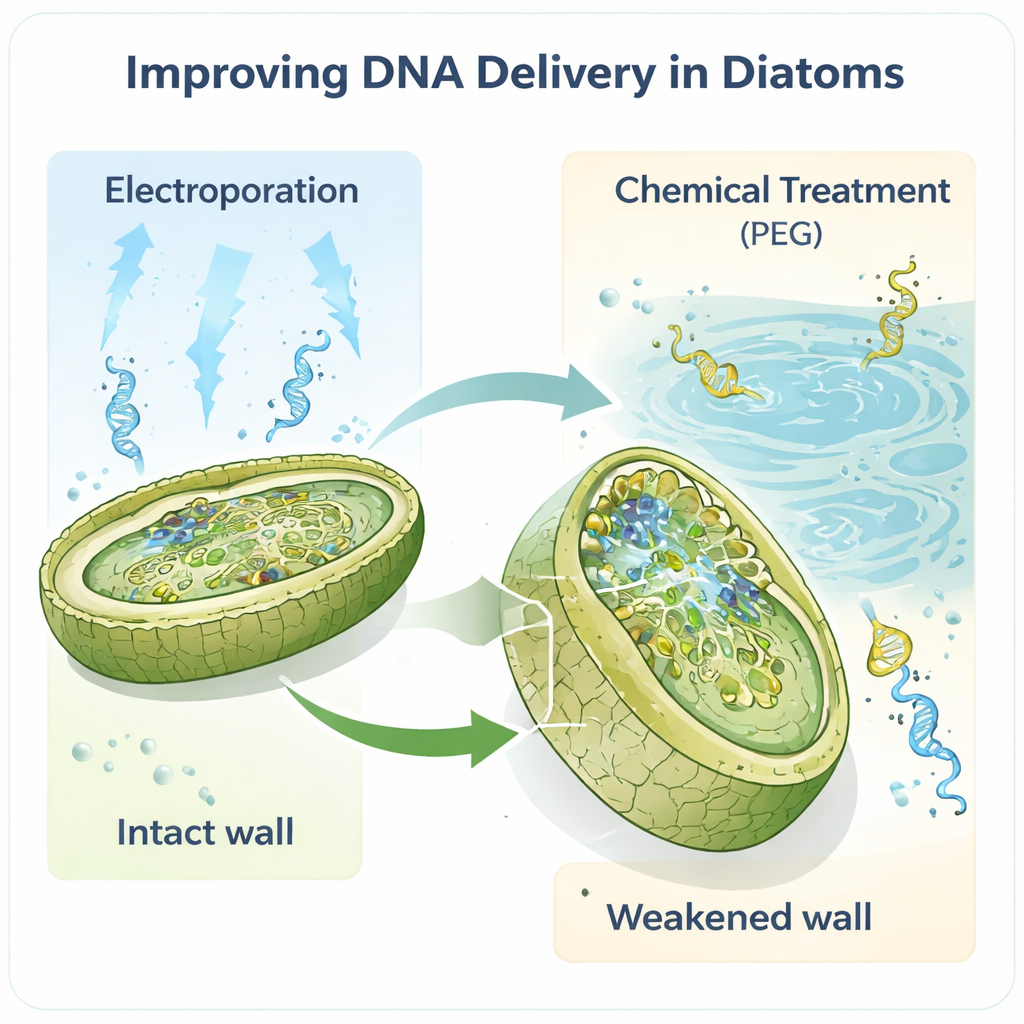
Fast DNA Delivery Without Detours Through Bacteria
Traditional ways of modifying diatoms often rely on bacterial “couriers” to ferry large circular DNA molecules, called episomes, into algae. While effective, that approach is slow, technically demanding, and can destabilize delicate DNA constructs. The new protocols show that both electroporation and a refined polyethylene glycol (PEG) chemical method can move episomes directly into diatoms, bypassing the bacterial step. Remarkably, episomes up to 55.6 thousand base pairs long were delivered and recovered intact. The same strategies also worked in a second species, Thalassiosira pseudonana, which has a more heavily mineralized wall, suggesting that this is a broadly useful toolkit rather than a one‑species trick.
Letting the Cell Build Its Own Genetic Circles
While probing how electroporated DNA behaved inside cells, the team stumbled on a surprising talent: diatoms can stitch DNA fragments together on their own. Linear pieces of episomes entered the cell and were repaired into circles either by imprecise “non‑homologous” joining or by more accurate, overlap‑guided “homology‑directed” repair. The authors dubbed this process “diatom in vivo assembly,” or DIVA. By designing overlapping fragments, they coaxed cells to assemble two, three, or four pieces into full episomes with high success rates, sometimes even incorporating small synthetic cassettes that add fluorescent tags or new functions. This ability turns the diatom nucleus into a miniature DNA workshop, potentially replacing labor‑intensive assembly steps usually done in yeast or E. coli.
Editing Genomes With Protein Complexes Only
Beyond adding episomes, the researchers showed they could deliver ready‑made CRISPR–Cas9 protein‑RNA complexes directly into diatom cells using their optimized electroporation method. Targeting a gene called PtAPT, which controls sensitivity to a toxic adenine look‑alike, they generated thousands of resistant mutants without introducing any extra DNA into the genome. Many of these mutants carried small insertions or deletions at the target site; some even captured bits of “carrier” DNA that had been added to buffer the electric shock. The team then co‑delivered CRISPR complexes and episomes in a single step, finding that roughly one in ten colonies carried both a genomic edit and a selectable episome—an efficient way to track otherwise invisible gene changes. 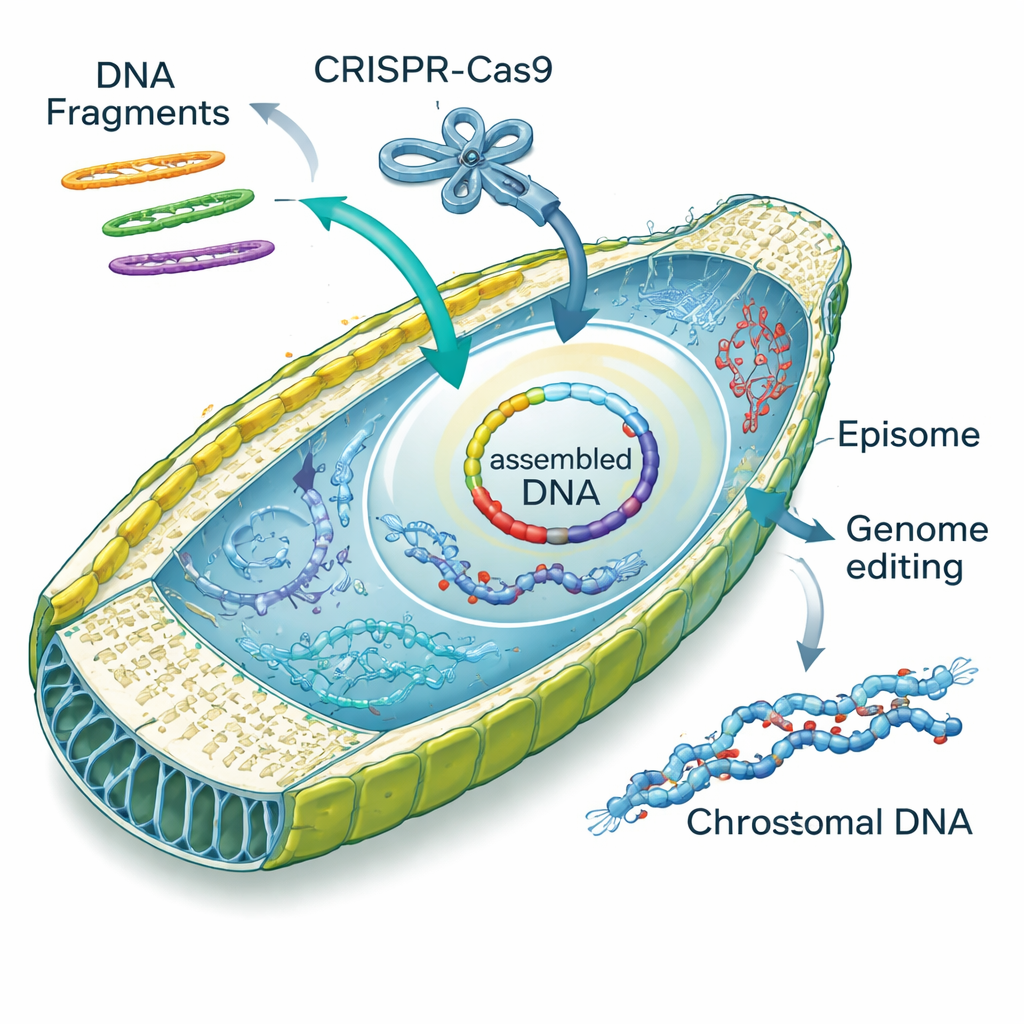
Toward Designer Diatoms for a Sustainable Future
For non‑specialists, the key message is that diatoms are moving much closer to being practical, programmable organisms. By gently breaking or softening the cell wall, the authors turned a finicky, low‑yield process into a robust pipeline that uses small amounts of DNA, works with large genetic constructs, and even lets the cell assemble and edit its own DNA. These advances shorten the path from a computer‑designed sequence to a living, tested strain. In the long run, such tools could speed efforts to build diatoms with entirely synthetic chromosomes and to harness these ocean microbes for cleaner fuels, climate‑friendly chemistry, and new biological discoveries.
Citation: Walker, E.J.L., Pampuch, M., Deng, L. et al. Breaking the cell wall for efficient DNA delivery to diatoms. Nat Commun 17, 1848 (2026). https://doi.org/10.1038/s41467-026-68562-6
Keywords: diatom biotechnology, genetic transformation, CRISPR genome editing, synthetic biology, microalgae engineering