Clear Sky Science · en
Structures of nucleotide-bound human telomerase at several steps of its telomeric DNA repeat addition cycle
Why the ends of our chromosomes matter
Every time a cell divides, the protective caps at the ends of our chromosomes—called telomeres—get a little shorter. If they become too short, cells stop dividing or malfunction, contributing to aging and disease. A molecular machine called telomerase can rebuild these caps and is unusually active in most cancers. This study reveals, in unprecedented detail, how human telomerase grips and extends telomeres step by step, offering clues to how we might one day fine‑tune this enzyme in age‑related disorders and cancer.
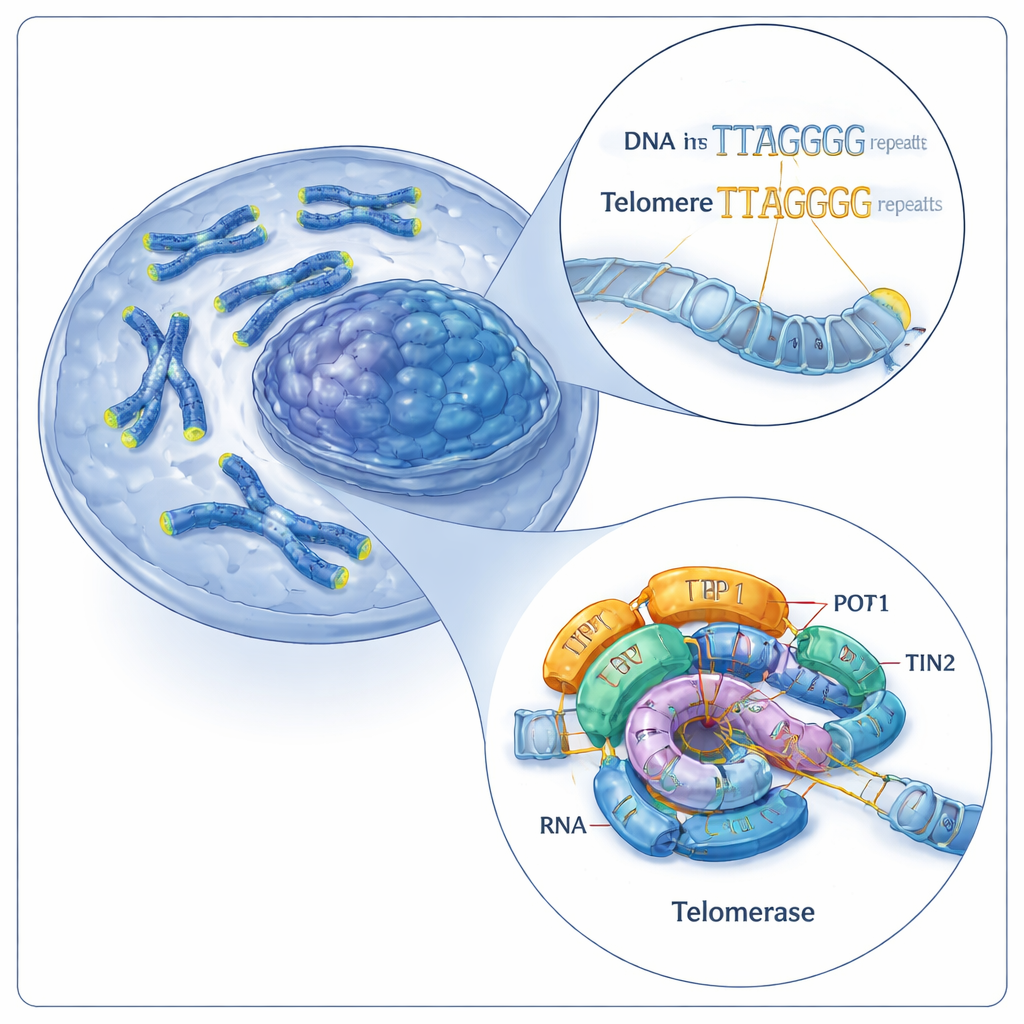
The cell’s chromosome cap repair crew
Telomeres are made of short, repeated DNA sequences (in humans, a six‑letter pattern: TTAGGG). Telomerase lengthens telomeres by adding more of these repeats onto chromosome ends. It does this using two main parts: a protein called telomerase reverse transcriptase (TERT) and a built‑in RNA molecule (hTR) that acts as a template. Together, these components form a catalytic core, which is further helped by accessory proteins known as shelterin, including a trio called TPP1–POT1–TIN2. Previous work had visualized human telomerase in just one working state, leaving a major question open: how does this machine repeatedly add new repeats without falling off the DNA?
Freezing telomerase in action
The researchers rebuilt human telomerase in human cells and then assembled it with carefully designed pieces of telomeric DNA and non‑reactive nucleotide analogs—molecules that look like the letters of DNA but cannot be fully incorporated. Using high‑resolution cryo‑electron microscopy, they captured snapshots of telomerase at three key stages of its repeat‑addition cycle: the start (initiation), the middle of copying (elongation), and just before finishing a repeat (pre‑termination). Each snapshot showed the catalytic core of telomerase embracing a short hybrid of DNA and RNA, with an incoming nucleotide poised in the active site, ready to be added to the growing DNA strand.
A tiny zipper that controls a big process
Across all three stages, a surprising theme emerged: the DNA–RNA hybrid inside telomerase remains essentially the same length—only four base pairs long, temporarily extended to five when a new nucleotide enters. A specific amino acid in TERT, nicknamed a “zipper head,” consistently marks the end of this short hybrid and prevents it from growing longer. As new DNA letters are added, one base pair at the far end melts away, so the hybrid never extends beyond this tight four‑to‑five‑pair window. This compact design likely makes it easier for the strands to separate and shift position, allowing the enzyme to move along and start the next repeat without fully letting go of the DNA. Subtle differences in which letters of DNA and RNA occupy this short hybrid explain why some telomeric end sequences bind more tightly to telomerase than others.
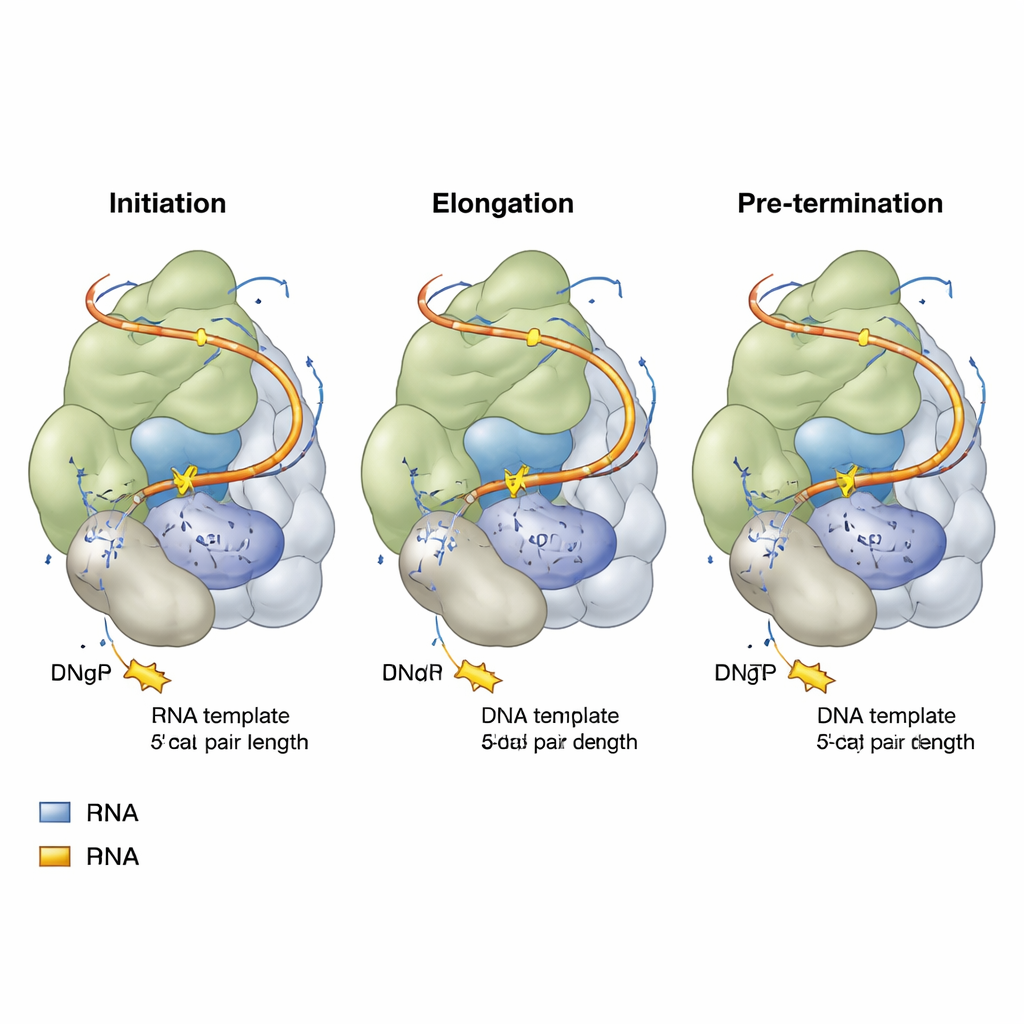
Flexible RNA linkers and moving parts
The RNA template within telomerase does not float freely; it is tethered by flexible stretches on both sides, called 5′ and 3′ template linkers. As telomerase progresses from initiation to pre‑termination, the upstream (5′) linker is pulled taut like a bungee cord, eventually reaching a fully stretched state that helps signal the end of a repeat. Meanwhile, the downstream (3′) linker bulges outward and approaches another RNA structure (the P6.1 stem‑loop) and nearby protein regions. If these linkers become too short or too long, or if P6.1 is altered, telomerase’s ability to add repeat after repeat drops sharply. Importantly, certain protein regions—the TEN domain and a telomerase‑specific insert called the TRAP wedge—act as adjustable guides, contacting both the DNA and these RNA linkers. Mutations that reduce the bulk of the TRAP wedge can actually make the enzyme more processive, suggesting this wedge normally acts as a controlled barrier that times template movement.
New insights into a therapeutic target
By combining structural snapshots with functional tests of targeted mutations, the study proposes a detailed mechanical model for how human telomerase repeatedly extends telomeres. The enzyme holds a very short DNA–RNA hybrid, uses a zipper‑like residue to enforce that length, and relies on flexible RNA linkers and mobile protein elements to advance the template and keep the DNA anchored during multiple rounds of copying. For non‑specialists, the key message is that telomerase works not as a static copying machine but as a finely tuned, spring‑loaded device whose geometry and moving parts determine how long and how efficiently it can maintain telomeres. Understanding this choreography opens the door to drugs that might either dampen telomerase in cancers or stabilize its function in diseases of premature aging.
Citation: Balch, S., Franco-Echevarría, E., Ghanim, G.E. et al. Structures of nucleotide-bound human telomerase at several steps of its telomeric DNA repeat addition cycle. Nat Commun 17, 1847 (2026). https://doi.org/10.1038/s41467-026-68560-8
Keywords: telomerase, telomeres, cryo-EM, genome stability, cancer biology