Clear Sky Science · en
DNA methylation and lncRNA control asynchronous DNA replication at specific imprinted gene domains
Why the cell’s DNA copying schedule matters
Every time a cell divides, it must copy its entire DNA, but not all regions are duplicated at the same moment. Some stretches are copied early, others late, following a precise daily “timetable” for the genome. This paper explores why, in a few special gene clusters that remember which parent they came from, the mother’s and father’s copies are copied at different times. Understanding this unusual timing sheds light on how chemical tags on DNA and long non-coding RNAs (lncRNAs) help organize the genome and may influence development and disease.
Parental memories written on DNA
In mammals, a small number of “imprinted” gene domains behave differently depending on whether they are inherited from the mother or the father. This parent-of-origin effect is controlled by DNA methylation—tiny chemical marks added to specific regions known as differentially methylated regions (DMRs)—and by changes in chromosome folding. The authors studied mouse embryonic stem cells, which resemble very early embryonic cells. They compared cells with only maternal genomes, only paternal genomes, and normal hybrids carrying one of each. By tracking when pieces of DNA were copied during the cell cycle, they mapped replication timing across the genome and zoomed in on known imprinted regions.
Two imprinted regions break the rules
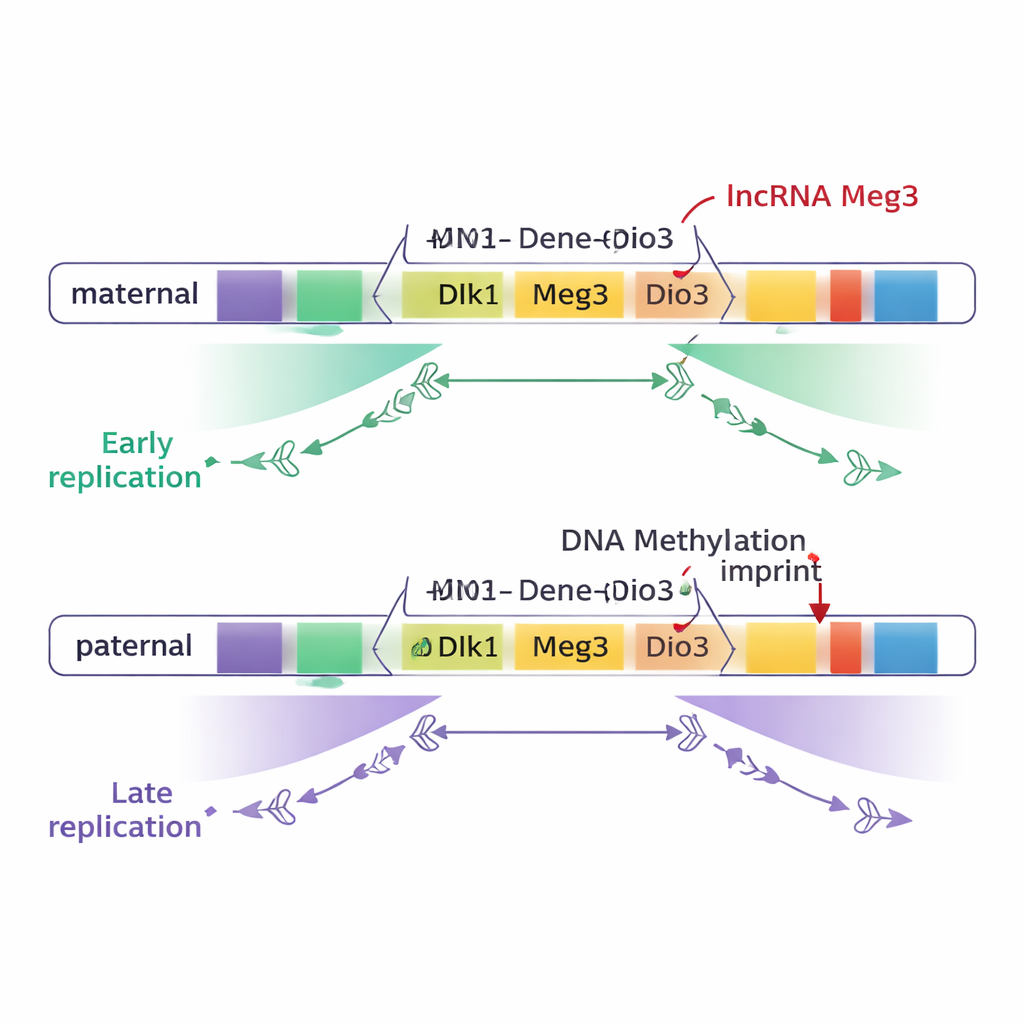
Most of the genome, including most imprinted domains, turned out to be copied at the same time on maternal and paternal chromosomes. However, two large imprinted regions stood out: the Dlk1–Dio3 domain on chromosome 12 and the Snrpn region on chromosome 7. In these zones, a wide stretch of DNA—about three-quarters of a million base pairs at Dlk1–Dio3—was copied early on one parental chromosome and late on the other. Crucially, this difference followed parental origin, not strain background: the maternal copy of key genes such as Dlk1 and the lncRNA gene Meg3 consistently replicated early, while the paternal copies replicated late.
DNA methylation sets the timing switch
To test what causes this timing difference, the researchers engineered stem cells in which the normal methylation pattern at DMRs was either erased or forced onto both parental chromosomes. When both parental copies of the Dlk1–Dio3 DMRs were methylated, the entire region replicated late on both chromosomes. When methylation was largely removed from both copies, the same region replicated early on both. Similar experiments at Snrpn produced a loss of timing differences as well. These results show that parent-specific DNA methylation is essential to create the early-versus-late replication contrast between maternal and paternal chromosomes, at least in these two domains.
A long RNA fine-tunes early replication
DNA methylation alone, however, was not the whole story. The Dlk1–Dio3 region also produces a long non-coding RNA called Meg3 as part of a large RNA “polycistron.” On the maternal chromosome, unmethylated DMRs allow Meg3 to be expressed; on the paternal chromosome, methylation keeps it silent. By creating precise deletions that turned off Meg3 while leaving DNA methylation intact, the authors showed that loss of Meg3 RNA caused specific parts of the maternal domain to shift from early to later replication. In other words, Meg3 RNA helps promote early copying across nearby stretches of the maternal chromosome, adding a second layer of control on top of DNA methylation.
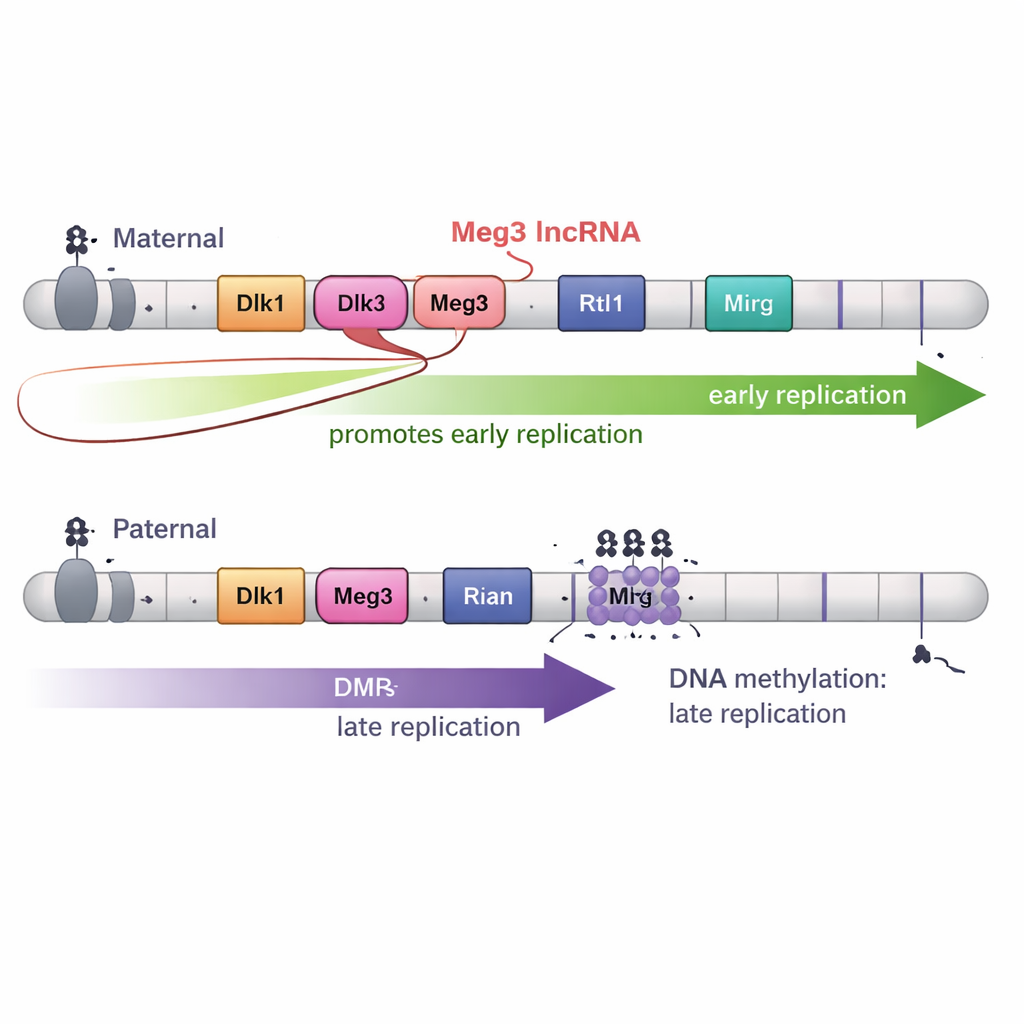
3D folding and development add further twists
Because genome folding is often linked to replication timing, the team also mapped 3D chromosome architecture in high detail. Surprisingly, in the Dlk1–Dio3 region, the borders of 3D domains did not match the boundaries of early and late replication. Even when methylation patterns were altered and replication timing changed, the basic folding units—topologically associating domains—shifted in different ways. Finally, when the stem cells were differentiated into neural progenitor cells, the striking timing differences between maternal and paternal copies largely disappeared, even though imprinting marks and many aspects of 3D structure remained. This suggests that developmental cues can override the special early-versus-late pattern seen in stem cells.
What this means in simple terms
This study reveals that, in a few special gene neighborhoods that remember which parent they came from, the cell’s DNA copying schedule is controlled by a combination of chemical marks on DNA and long non-coding RNAs. DNA methylation at imprinted control regions sets a basic early-or-late timing for each parental chromosome, and the Meg3 lncRNA further nudges nearby maternal DNA to be copied earlier. These effects act largely independently of how the chromosome folds in 3D. During development, as cells specialize, this unusual timing fades, showing that it is a feature of the pluripotent stem cell state. Together, the work explains how parental epigenetic “memories” and non-coding RNAs can locally override the genome’s usual replication timetable.
Citation: Imaizumi, Y., Charon, F., Surcis, C. et al. DNA methylation and lncRNA control asynchronous DNA replication at specific imprinted gene domains. Nat Commun 17, 1844 (2026). https://doi.org/10.1038/s41467-026-68558-2
Keywords: genomic imprinting, DNA methylation, replication timing, long non-coding RNA, embryonic stem cells