Clear Sky Science · en
Ultrathin liquid cells for microsecond time-resolved cryo-EM
Watching Proteins at Work in Real Time
Many of the machines that keep our cells alive are made of proteins, and we often know what these proteins look like when frozen in place. But what we really want to see is how they move while they do their jobs. This study introduces a new way to watch these motions on incredibly short timescales—millionths of a second—without sacrificing the razor-sharp detail that modern cryo‑electron microscopy (cryo‑EM) can provide.
A New Window Into the Microscopic World
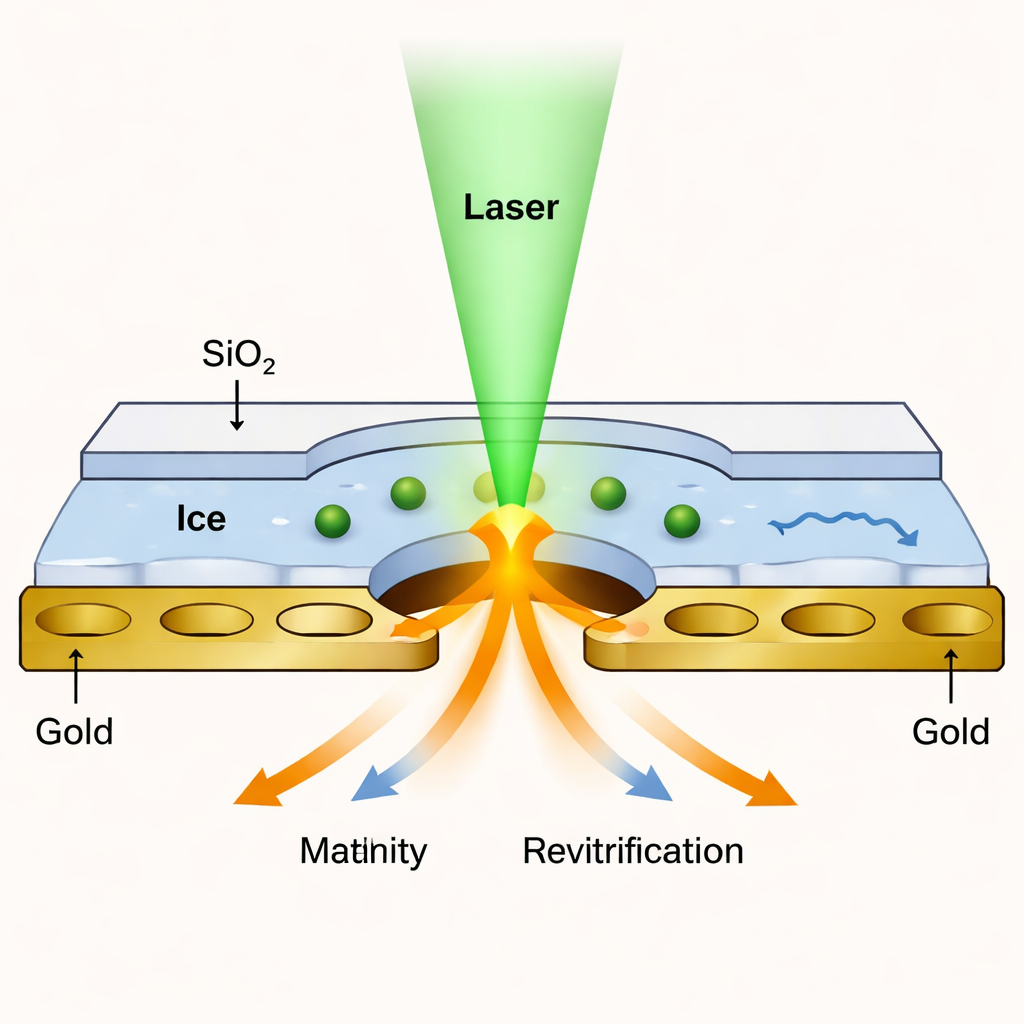
Cryo‑EM has revolutionized structural biology by imaging flash‑frozen proteins at near‑atomic resolution. However, traditional methods show only still pictures. To capture motion, researchers developed “microsecond time‑resolved cryo‑EM,” where a laser briefly melts a frozen sample so proteins can move, and then the sample is rapidly re‑frozen to trap them in new positions. The problem has been that the tiny liquid film created by the laser tends to break up after a few tens of microseconds, limiting how long the proteins can be observed. The new work solves this bottleneck by enclosing the sample in an ultrathin liquid cell, keeping it stable long enough to watch slower, more complex motions.
Building an Ultrathin Liquid Cell
The team created a kind of nanoscopic sandwich: the protein solution is frozen on a standard perforated gold grid, then both sides are coated with layers of silicon dioxide only about 1.4 nanometers thick—just a few atoms. These glass‑like layers act as transparent lids that prevent the liquid from evaporating when the laser melts the ice. Short laser pulses heat the sealed sample to a controlled temperature and then allow it to re‑freeze within microseconds. Because the membranes are so thin, they still let enough electrons through for the microscope to produce images with nearly the same resolution as in conventional cryo‑EM, down to about 1.7–1.8 angstroms for a test protein called apoferritin.
Sharper Views and Fairer Angles
One hidden challenge in cryo‑EM is that proteins tend to stick to the air–water interface in the thin layer of ice, lining up in similar orientations and making it harder to reconstruct a full 3D view. The silicon dioxide coatings in these liquid cells change the surface from water‑air to water‑solid and make it more friendly to water. As a result, proteins are less likely to cling in a single pose. When the authors tested a large cellular machine called the 50S ribosomal subunit, they found that the angular distribution of particles became almost perfectly even, essentially eliminating the long‑standing problem of “preferred orientation” while still maintaining high resolution in the final reconstructions.
Timing the Motion of a Molecular Lever
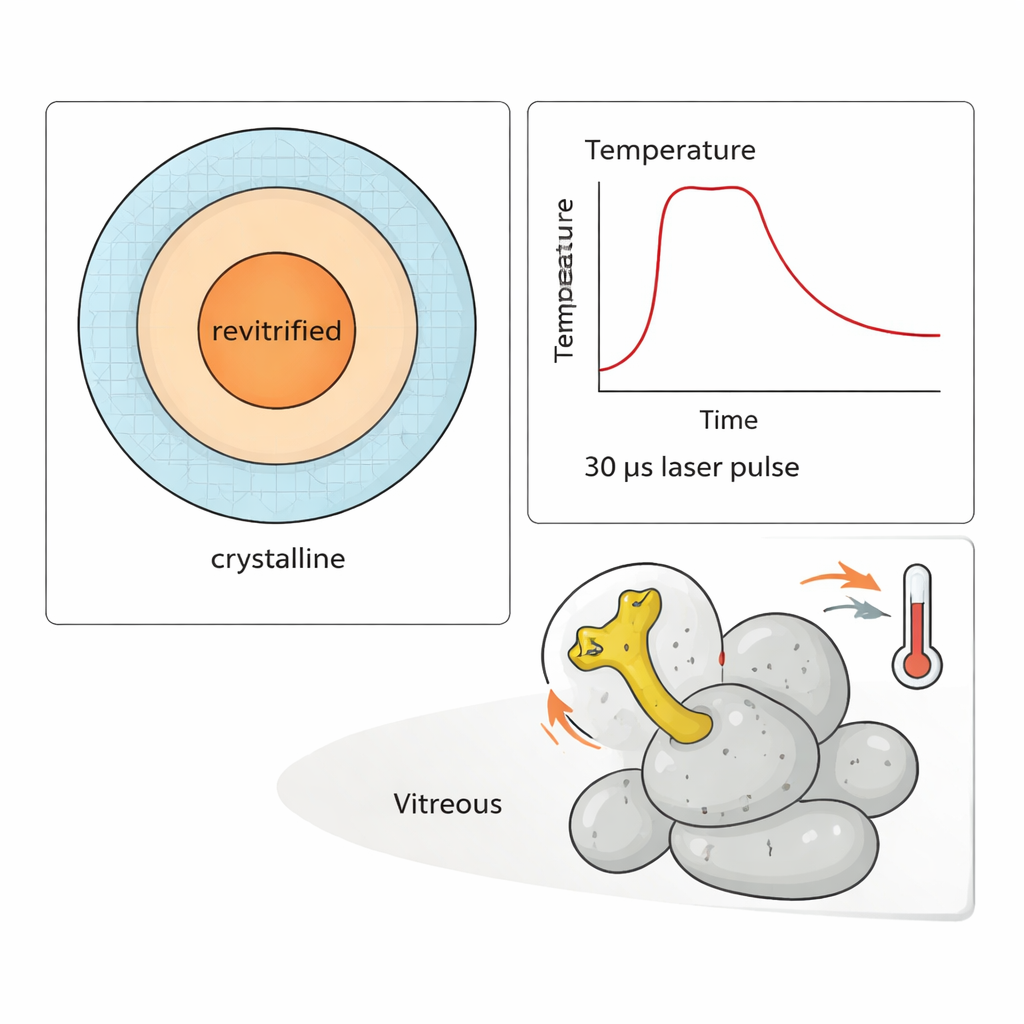
To showcase the power of their method, the researchers carried out a “temperature jump” experiment on the 50S subunit. A flexible arm on this particle, known as the L1 stalk, swings like a lever during protein synthesis. By delivering trains of 30‑microsecond laser pulses, they could heat the sample to different temperatures for up to about 300 microseconds and then refreeze it. Simulations and measurements of which regions re‑froze as glass allowed them to estimate the temperature for each observed particle. Analyzing thousands of images, they showed that the amplitude of L1 stalk motion clearly increases with temperature—but only after hundreds of microseconds have passed. At early times, the distribution of conformations still reflects the original, room‑temperature state before freezing.
Why This Matters for Future Biology
For non‑specialists, the key message is that this ultrathin liquid cell design dramatically extends how long proteins can be observed in motion without blurring the structural details. It shifts microsecond time‑resolved cryo‑EM from capturing only the very fastest events to probing slower, biologically important rearrangements, like the delayed response of the L1 stalk to a heat pulse. With further refinements, this approach could bridge the gap to the millisecond range and beyond, and it also offers new ways to prepare samples, reduce imaging artifacts, and trigger reactions directly on the grid. In practical terms, this means scientists are getting closer to making “molecular movies” that link the shapes of proteins to what they actually do inside living cells.
Citation: Curtis, W.A., Wenz, J., Krüger, C.R. et al. Ultrathin liquid cells for microsecond time-resolved cryo-EM. Nat Commun 17, 1799 (2026). https://doi.org/10.1038/s41467-026-68515-z
Keywords: time-resolved cryo-EM, protein dynamics, liquid cell electron microscopy, ribosome L1 stalk, ultrathin silicon dioxide membranes