Clear Sky Science · en
Robust characterization and interpretation of rare pathogenic cell populations from spatial omics using GARDEN
Why the hunt for rare cells matters
Modern biology can measure which genes are switched on in thousands of tiny locations across a tissue, giving scientists a detailed "map" of what cells are doing and where they sit. Hidden in these maps are rare cells that can drive cancer spread, brain degeneration, or tissue repair, but they are easy to miss because they are so few and often scattered. This paper introduces GARDEN, a new computational approach designed to reliably spot and interpret these rare, disease‑linked cells in complex tissue maps.
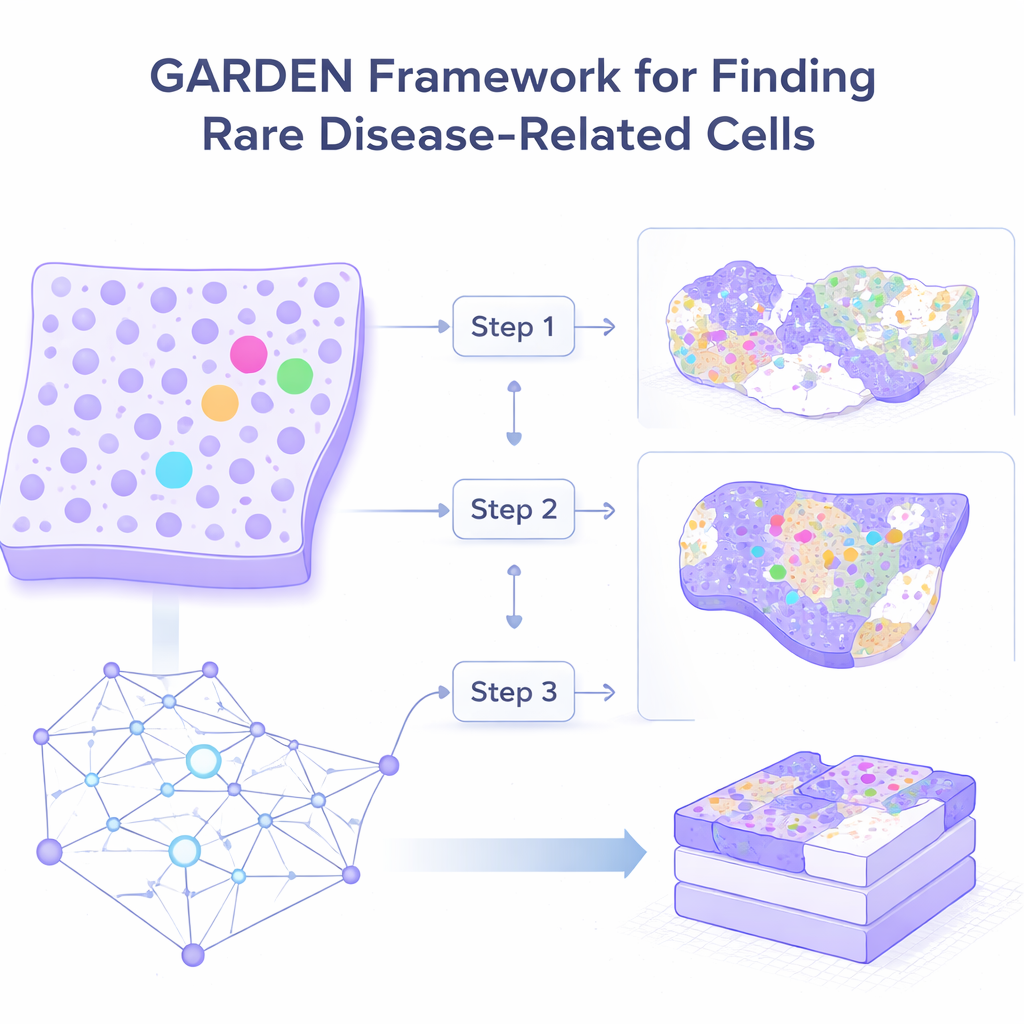
Seeing tissues as living neighborhoods
Traditional methods for analyzing spatial gene maps tend to focus on the most common cell types, smoothing over subtle local differences. GARDEN takes a different view: it treats each tiny tissue location as a point in a neighborhood network, where connections represent physical closeness and shared activity. Using a type of deep learning that can adjust its focus on the fly, GARDEN learns which neighboring cells deserve more attention and which can be safely ignored. This “dynamic attention” lets the method preserve small, sharp differences instead of blending them away.
Separating real biological change from noise
Because real tissues are messy, GARDEN must distinguish true rare cell patterns from random fluctuations. To do this, the system creates artificial “what‑if” versions of the tissue by shuffling gene activity while keeping positions the same. It then trains itself to tell apart genuine microenvironments from these fakes, sharpening its sense of what real local structure looks like. At the same time, it learns to reconstruct the original gene activity from its internal representation, which forces it to keep biologically meaningful detail. This combination makes GARDEN particularly good at finding small pockets of unusual cells.
Rebuilding tissues in 3D
Most spatial datasets are thin slices through a much thicker organ. A rare cell type might appear in only a few slices, making it easy to misinterpret or overlook. GARDEN includes a registration module that aligns many consecutive slices into a coherent three‑dimensional shape. By matching patterns of gene activity and spatial layout across slices, it can reconstruct structures such as layered brain regions and then trace how rare cells spread through depth. In mouse brain data, for example, GARDEN consistently aligned slices into a realistic 3D hemisphere and revealed how scarce cell types line specific layers and pathways.
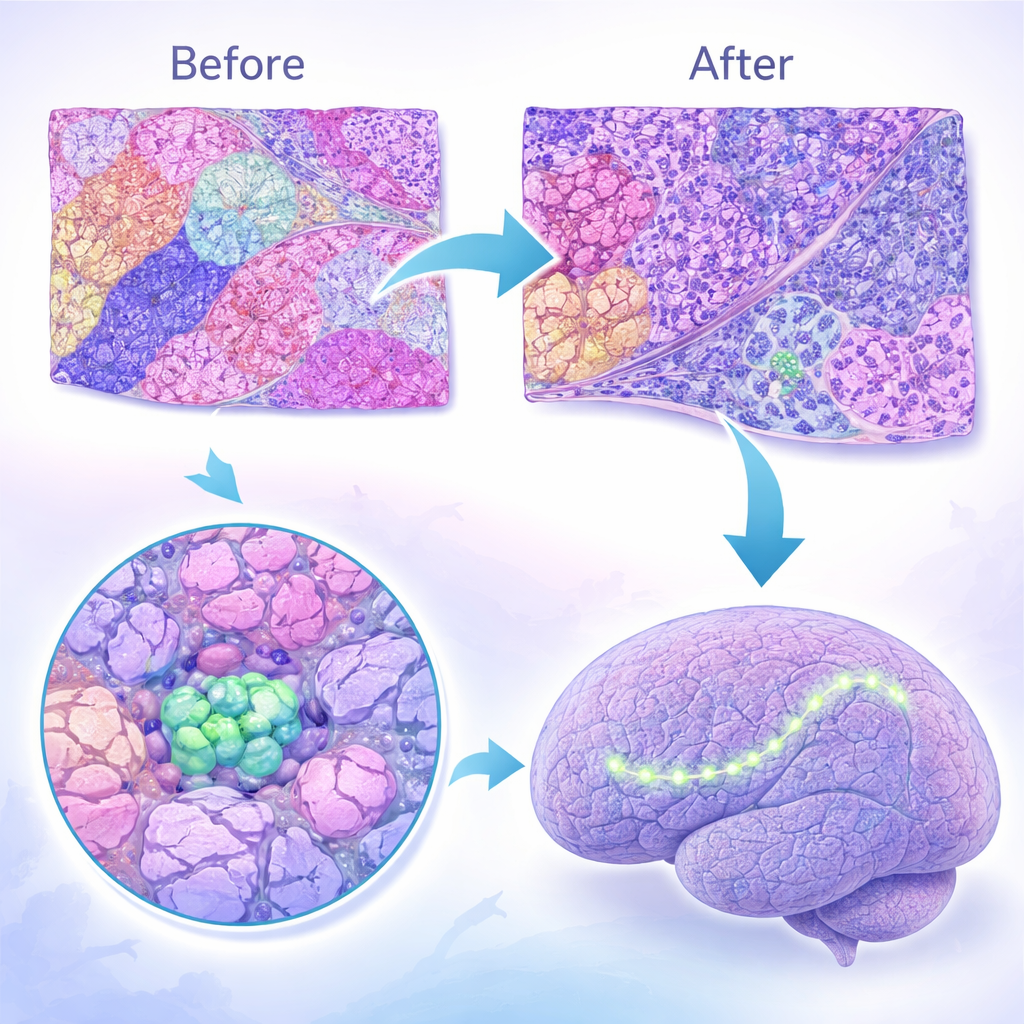
Discoveries in cancer and Alzheimer’s disease
Applied to a wide range of real tissues, GARDEN repeatedly uncovered small but important structures that other tools missed. In the mouse olfactory bulb and across different animal brains, it identified tiny zones rich in immature nerve cells or specialized medulla regions, supported by gene markers and known anatomy. In high‑resolution colorectal cancer samples, GARDEN split invasive tumor areas into distinct subgroups tied to different genes that promote growth, invasion, or blood‑vessel changes, suggesting multiple aggressive behaviors within what looked like a single mass. In nasopharyngeal carcinoma, it detected very small, scattered immune hubs called tertiary lymphoid structures, and showed that in some patients these hubs were in a highly active, antibody‑producing state. In mouse models of Alzheimer’s disease, GARDEN pinned down rare microglia and astrocyte populations clustered around amyloid‑β plaques, along with gene programs linked to inflammation and synapse loss.
From gene activity to possible treatments
Beyond locating rare cells, GARDEN helps explain what makes them special. By probing which genes are most important for its decisions, and grouping these genes into functional modules, the method highlights immune signals, regulatory factors, and cell‑to‑cell communication pathways that define each rare niche. In breast cancer, for instance, GARDEN revealed inflammatory macrophages sitting next to early‑stage ductal carcinoma in situ and traced communication routes and gene pairs associated with potential metastasis. In spatial chromatin (DNA accessibility) maps, it cleanly separated fine brain layers and followed how key developmental genes gradually switch on. Altogether, GARDEN offers a powerful way to turn complex spatial omics data into concrete biological stories, pointing researchers toward rare cell types and circuits that may become early diagnostic markers or targets for more personalized therapies.
Citation: Zhang, X., Yu, Z., Hao, G. et al. Robust characterization and interpretation of rare pathogenic cell populations from spatial omics using GARDEN. Nat Commun 17, 1792 (2026). https://doi.org/10.1038/s41467-026-68500-6
Keywords: spatial omics, rare cell populations, GARDEN, cancer microenvironment, Alzheimer’s disease