Clear Sky Science · en
Identification of antimicrobial peptides from ancient gut microbiomes
Digging Up Ancient Clues to Fight Modern Germs
As antibiotic resistance rises around the world, doctors are running out of ways to treat dangerous infections. This study takes an unusual approach to the problem: it looks not to high-tech chemistry labs, but to the fossilized remains of human feces—ancient gut "time capsules"—to uncover natural germ-fighting molecules that may still work against today’s bacteria.
Ancient Microbes as Hidden Medicine Cabinets
Long before modern medicine, the human gut was already a battleground where helpful microbes and invading germs competed for space and food. Many friendly gut bacteria make antimicrobial peptides—short protein fragments that punch holes in bacterial cells or otherwise disable them. Modern research has mostly searched today’s microbiomes for these germ-killing peptides. But our current gut communities have evolved alongside antibiotic use and modern lifestyles, giving harmful bacteria many chances to develop resistance. In contrast, ancient gut communities, preserved in dried feces called coprolites, formed in a world without prescription antibiotics. That makes them a promising source of "forgotten" defenses that modern pathogens may not yet have learned to evade.
A Lightweight AI Tool to Read Fossilized DNA
To explore this ancient pharmacy, the researchers built a new computer tool called AMPLiT (AMP Lightweight Identification Tool). Instead of needing large supercomputers, AMPLiT runs efficiently on an ordinary laptop while still maintaining high accuracy. It scans huge DNA datasets from microbiome samples and flags short sequences that are likely to encode antimicrobial peptides. The team fine-tuned AMPLiT’s design so it could handle millions of fragments from damaged, centuries-old DNA in a matter of hours, cutting training time by about 80% compared with earlier methods, yet keeping near state-of-the-art performance in spotting likely germ-killing candidates.
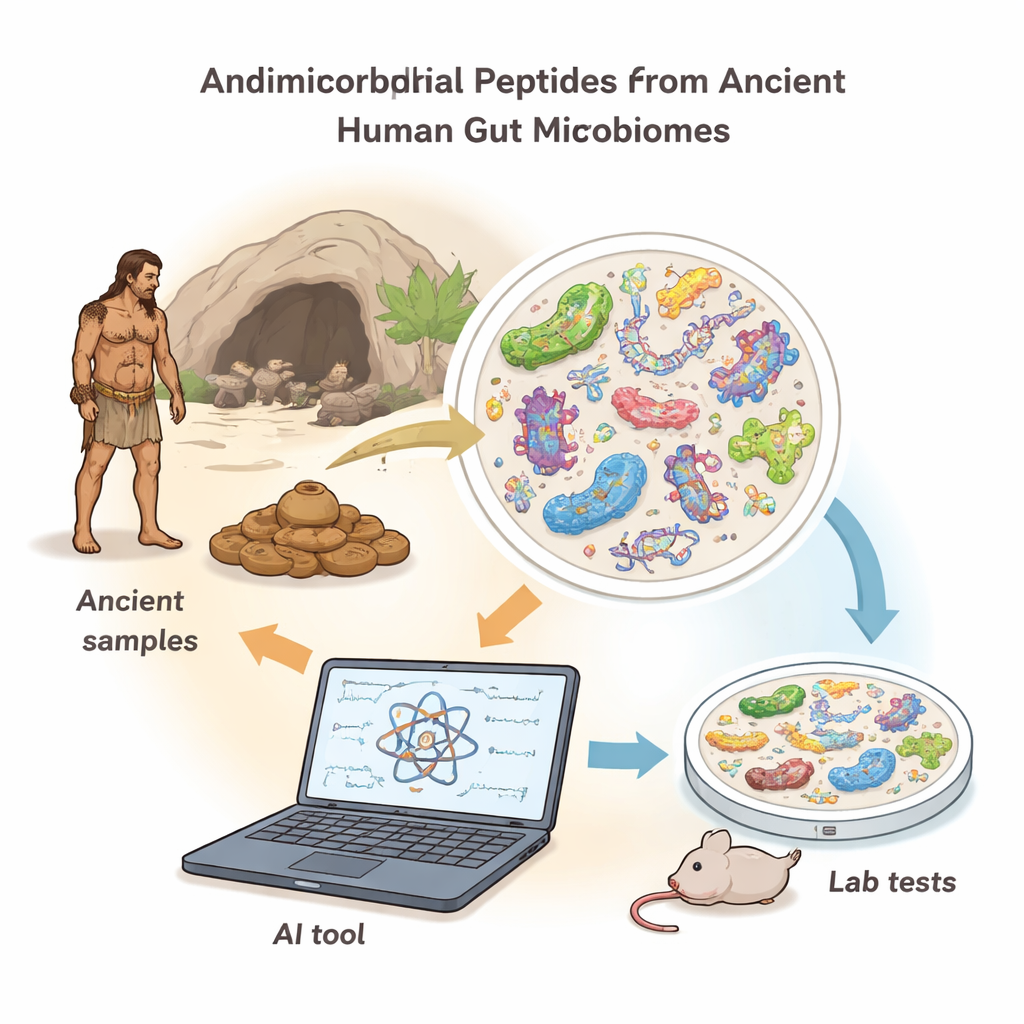
Resurrecting Germ Killers from an Ancient Gut Resident
Using AMPLiT, the scientists examined gut DNA from seven ancient humans who lived 1,000–2,000 years ago in North America. After removing environmental contamination and focusing on short, practical peptide lengths, the tool predicted hundreds of thousands of possible antimicrobial sequences. A stricter series of filters—presence in multiple individuals, chemical properties, and low predicted toxicity—narrowed this down to 41 high-confidence candidates, 40 of which could be chemically made in the lab. When tested against both "Gram-positive" and "Gram-negative" bacteria (two major groups that include many common pathogens), 36 of the 40 peptides slowed or stopped bacterial growth at relatively low doses, an unusually high success rate for this kind of discovery work.
The Surprising Star: A Vanishing Gut Ally
Roughly two-thirds of the most active peptides came from a single gut bacterium: Segatella copri, a close relative of a microbe formerly grouped under Prevotella copri. This species was abundant in ancient guts and is still common in people with more traditional, less industrialized diets, but it is now rare in many urban, Western-style populations. By tracing where the peptide genes sit in the Segatella genome, the team found that most are actually fragments of larger, everyday "housekeeping" genes that the microbe appears to have repurposed as weapons—an efficient evolutionary trick. Many of these ancient peptides look quite different from those in modern databases, suggesting they represent genuinely new chemical designs, not just minor variations on known antibiotics.
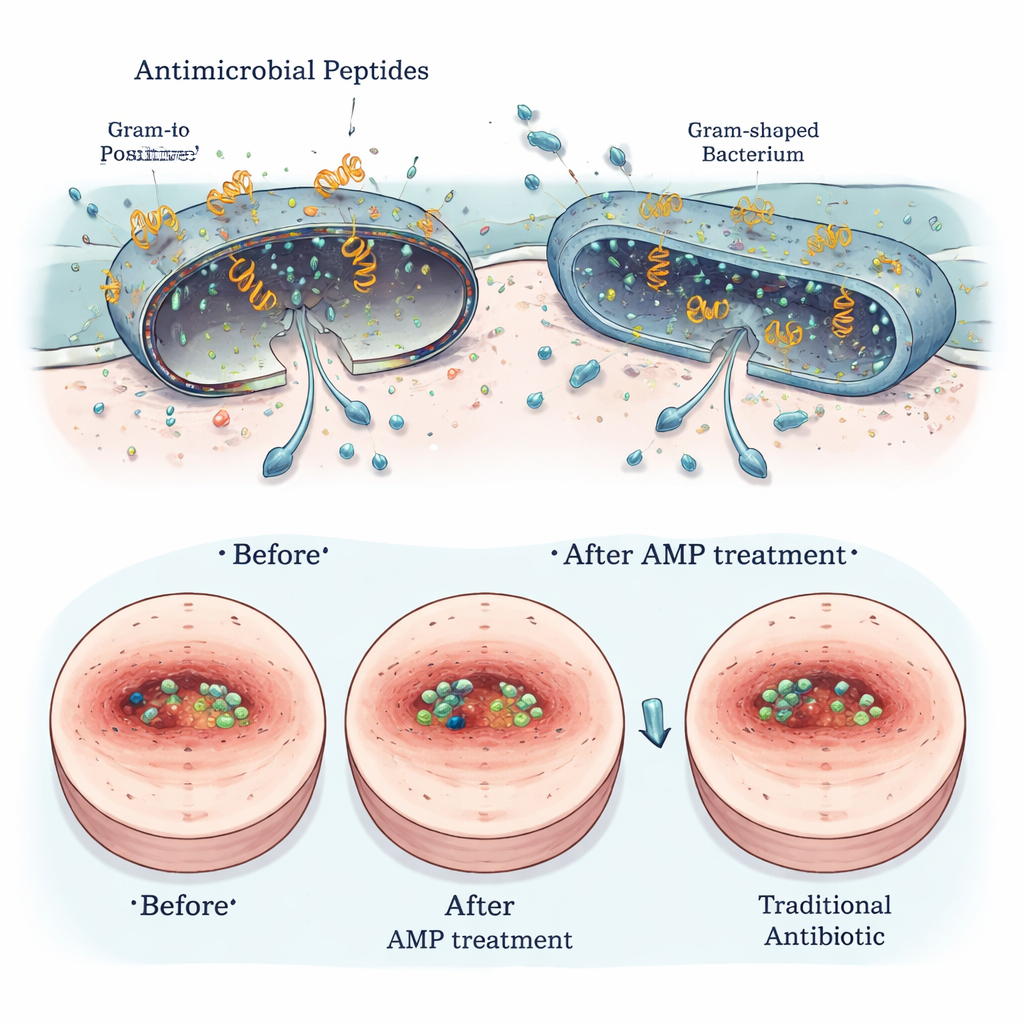
Safe for Hosts, Tough on Bugs—and Promising in Wounds
Several of the most promising Segatella peptides were tested for safety and real-world usefulness. In lab dishes, they caused little to no damage to red blood cells and only mild effects, if any, on human gut-like cells. High-resolution microscopy showed that the peptides physically disrupted the outer membranes of harmful bacteria, while sparing mammalian cells. In infected wound models in rodents, selected peptides applied to the skin reduced bacterial burden, sped up wound closure, and lessened signs of inflammation, with performance comparable to established antibiotics such as vancomycin and polymyxin B, especially against Gram-positive bacteria like Staphylococcus aureus.
What This Means for Future Medicines
To a non-specialist, the message is straightforward: our ancestors’ gut microbes may hold blueprints for new antibiotics that still work against today’s hard-to-treat infections. This study shows that with smart, efficient AI tools like AMPLiT, scientists can mine ancient DNA for antimicrobial peptides that are both potent and relatively gentle on human cells. While many steps remain before any of these molecules become drugs, the work suggests that reviving "lost" microbial partners such as Segatella copri—or at least borrowing their molecular weapons—could help replenish our shrinking arsenal against resistant bacteria.
Citation: Chen, S., Yuan, Y., Wang, Y. et al. Identification of antimicrobial peptides from ancient gut microbiomes. Nat Commun 17, 1788 (2026). https://doi.org/10.1038/s41467-026-68495-0
Keywords: antimicrobial peptides, ancient microbiome, Segatella copri, antibiotic resistance, metagenomic mining