Clear Sky Science · en
Continuously tunable multistability in DNA replication networks
From Simple Switches to Many Possible States
Biological systems, from dividing cells to developing embryos, rarely operate like simple light switches that are just on or off. Instead, they often hold many long‑lasting states, like a dimmer switch that can pause at countless positions. This paper reports a DNA‑based chemical network that behaves more like that dimmer: it can stably maintain not just a few distinct states, but essentially any composition along a continuous spectrum. That ability could become a powerful foundation for future smart materials, molecular memories, and biochemical computers that process analog, not just digital, information.
Why Multiple Stable States Matter
Many cellular decisions—whether a stem cell matures, a cell self‑destructs, or a gene network remembers a past signal—depend on “multistability,” the ability to rest in more than one stable pattern of activity under the same conditions. Traditional synthetic versions of such systems have been built from gene circuits, enzymes, or DNA strands that turn one another on or off. These designs typically yield a handful of discrete outcomes, like classic bistable switches with two long‑lived states. Scaling that idea up is difficult: an N‑state system can require on the order of N×(N−1) carefully tuned inhibitory links, and the system still hops only among a small set of fixed points rather than a smooth continuum of possibilities.
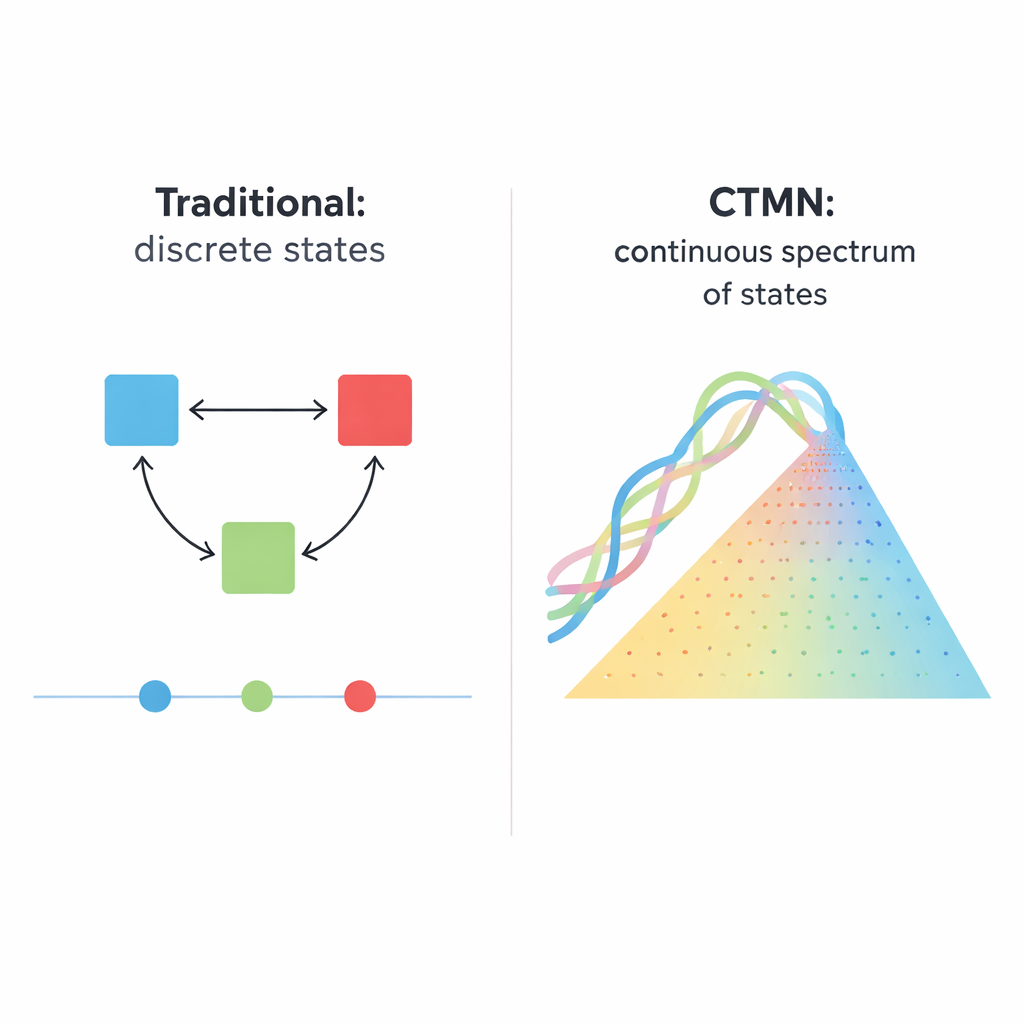
A New Kind of DNA Network: Continuously Tunable
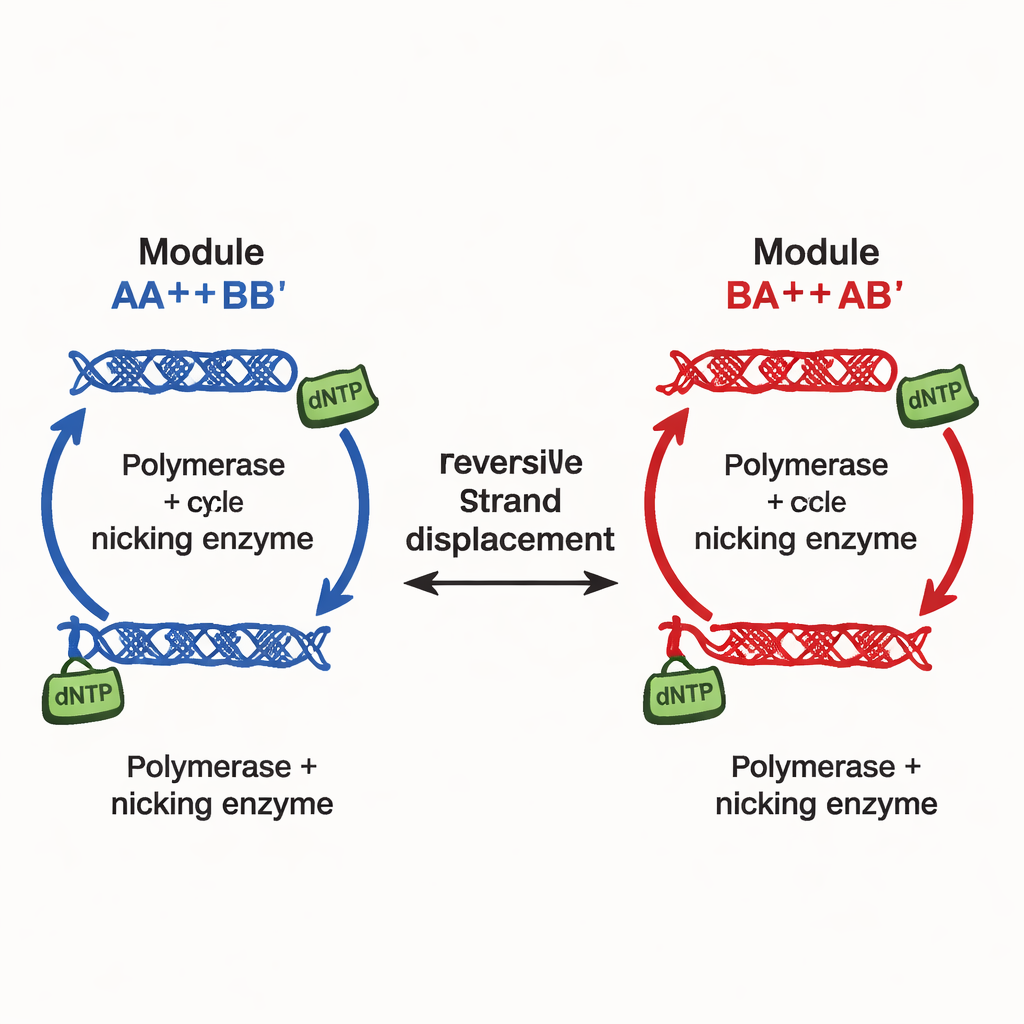
The authors introduce a different design principle, implemented in what they call continuously tunable multistability networks (CTMNs). Instead of relying on many separate inhibitory signals, they use direct competition between DNA replication modules joined by reversible strand‑exchange reactions. Each module is made from designer DNA duplexes that can both participate in strand displacement and serve as templates for DNA polymerase, aided by a nicking enzyme that cuts and regenerates key segments. The chemical fuel—nucleotide building blocks called dNTPs—drives ongoing polymerization and nicking, keeping the system far from equilibrium. Under these conditions, the competing modules effectively “freeze” into whatever relative proportions they started with, yielding not just two or three outcomes but a whole continuous line (or, for more modules, a plane) of long‑lived states.
How the Molecular Machinery Locks in Memory
In their simplest two‑module network, CTMN‑1, the team shows experimentally and through detailed kinetic modeling that the system can settle into a family of stable mixtures spanning almost the full range from “mostly module A” to “mostly module B.” Polymerization converts active duplexes into extended forms that are temporarily removed from strand‑exchange competition; nicking then restores them, closing an autoregulatory loop. When both modules share this loop, they suppress mutual interconversion so strongly that the system remains locked near its starting composition for many hours, provided fuel is available. Linear stability analysis of the underlying equations reveals an entire manifold—effectively a continuum—of steady states, in contrast to the single fixed point seen when the enzymes or fuel are removed.
Programmable Responses and Molecular Memory
Because the network’s state is stored in DNA compositions, it can be steered and read out in programmable ways. The authors design short RNA “triggers” that temporarily bind specific DNA strands, biasing the competition and shifting the network to a new composition. Even after the RNA is degraded by an enzyme, the new state persists, functioning as a form of molecular memory. Repeated or stronger stimuli drive the system smoothly along the continuum of states rather than flipping it between simple on/off positions. Moreover, the DNA replication products themselves can be customized to act as signals for downstream processes: in demonstrations, different states of CTMN‑1 selectively activate one of two DNA‑based catalysts or bias RNA transcription toward one of two messenger strands, without disrupting the underlying multistability.
Scaling Up to Complex Molecular “Dimmers”
To show that this strategy is scalable, the researchers build a three‑module network, CTMN‑2, by adding more designed DNA duplexes that share a common exchange domain but encode different replication templates. Now the possible steady states fill out a two‑dimensional region, like points across a triangular phase diagram, instead of along a simple line. Again, experiments confirm that many distinct compositions are stable as long as fuel and enzymes are present, and that all collapse to a single equilibrium point when the energy supply is removed. Because adding new modules requires only new DNA constructs that plug into the same displacement framework, the overall wiring does not balloon in complexity as in traditional designs.
What This Means for Future Molecular Technologies
To a non‑specialist, the core message is that the authors have turned a DNA replication network into a highly flexible, analog memory device. Rather than choosing among a few pre‑set options, their system can reliably hold one of dozens of experimentally distinguishable states, each defined by how much of each DNA module is present, and can convert transient chemical pulses into long‑lasting shifts along this continuum. Because the output DNA strands are programmable and decoupled from the stabilizing mechanism, the same framework could, in principle, regulate a wide range of downstream biochemical reactions or materials properties with fine, graded control. This work thus points toward molecular devices that compute, remember, and adapt more like neural or mechanical analog systems than like simple digital switches.
Citation: Zhong, R., Fu, Y., Jiang, S. et al. Continuously tunable multistability in DNA replication networks. Nat Commun 17, 1782 (2026). https://doi.org/10.1038/s41467-026-68488-z
Keywords: DNA multistability, molecular memory, synthetic biochemical networks, DNA strand displacement, analog molecular computing