Clear Sky Science · en
Robust and interpretable prediction of gene markers and cell types from spatial transcriptomics data
Turning Routine Tissue Slides into Molecular Maps
When a biopsy is taken, doctors usually see only what the microscope reveals: the shapes and patterns of cells in pink-and-purple stains. Yet beneath those colors lies a hidden world of genes switching on and off, influencing how a cancer grows and responds to treatment. This study introduces STimage, a new artificial intelligence (AI) system that aims to read that molecular script directly from standard pathology images, potentially offering faster, cheaper insights without extra laboratory tests.
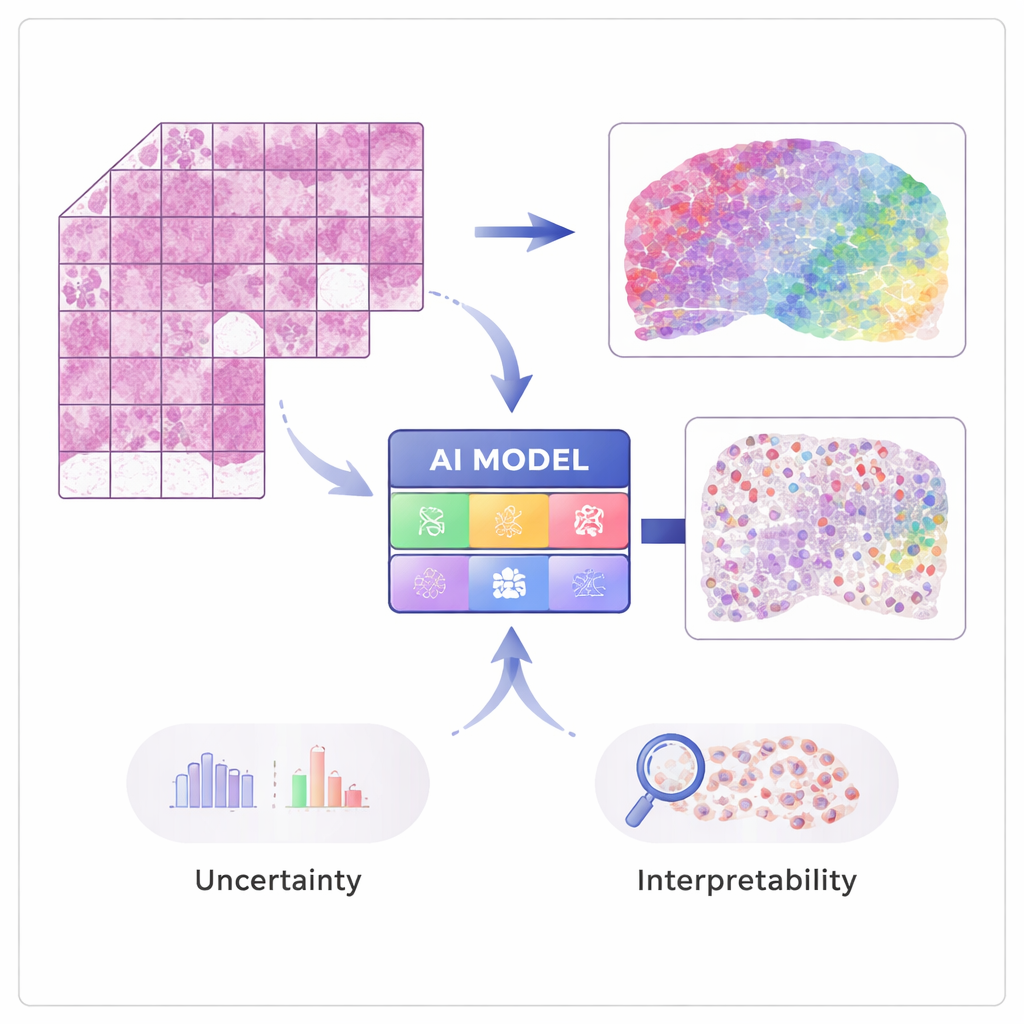
From Pictures to Gene Activity
Modern techniques in “spatial transcriptomics” can measure the activity of tens of thousands of genes while preserving where each signal came from in the tissue. These methods are powerful but expensive and not yet routine in hospitals. STimage is trained on a modest number of such spatial datasets, where each tissue image is matched with detailed gene measurements at many tiny spots. The AI learns to associate local visual patterns in the hematoxylin and eosin (H&E) slide—such as how dense or irregular the nuclei are—with the underlying gene activity, so that later it can predict gene expression and cell types from ordinary images alone.
Building a More Trustworthy AI Pathologist
A key goal of the work is not just accuracy, but reliability and explainability. Rather than outputting a single number for each gene, STimage predicts a full probability distribution, describing a likely range of gene activity at each location. It also separates two kinds of uncertainty: one driven by noisy or variable data, and another that reflects the model’s own lack of knowledge. By training many slightly different versions of the model and averaging them—an “ensemble” approach—the researchers both improve performance and gain a clearer sense of where the system is confident and where it is not, which is crucial for clinical decision-making.
Testing Across Cancers, Technologies, and Hospitals
The team evaluated STimage on diverse datasets from breast, skin, and kidney cancers, as well as an immune-related liver disease. It learned to predict important cancer and immune markers, often matching the true spatial patterns seen in independent experiments. The model held up when challenged with data from different laboratories, sample preparation methods, and even different underlying technologies, including single-cell-resolution platforms and older, lower-resolution systems. In head-to-head comparisons with several existing AI tools, STimage and its ensemble variants usually came out on top, particularly when judging how well the predicted patterns matched the real distribution of gene activity across the tissue.
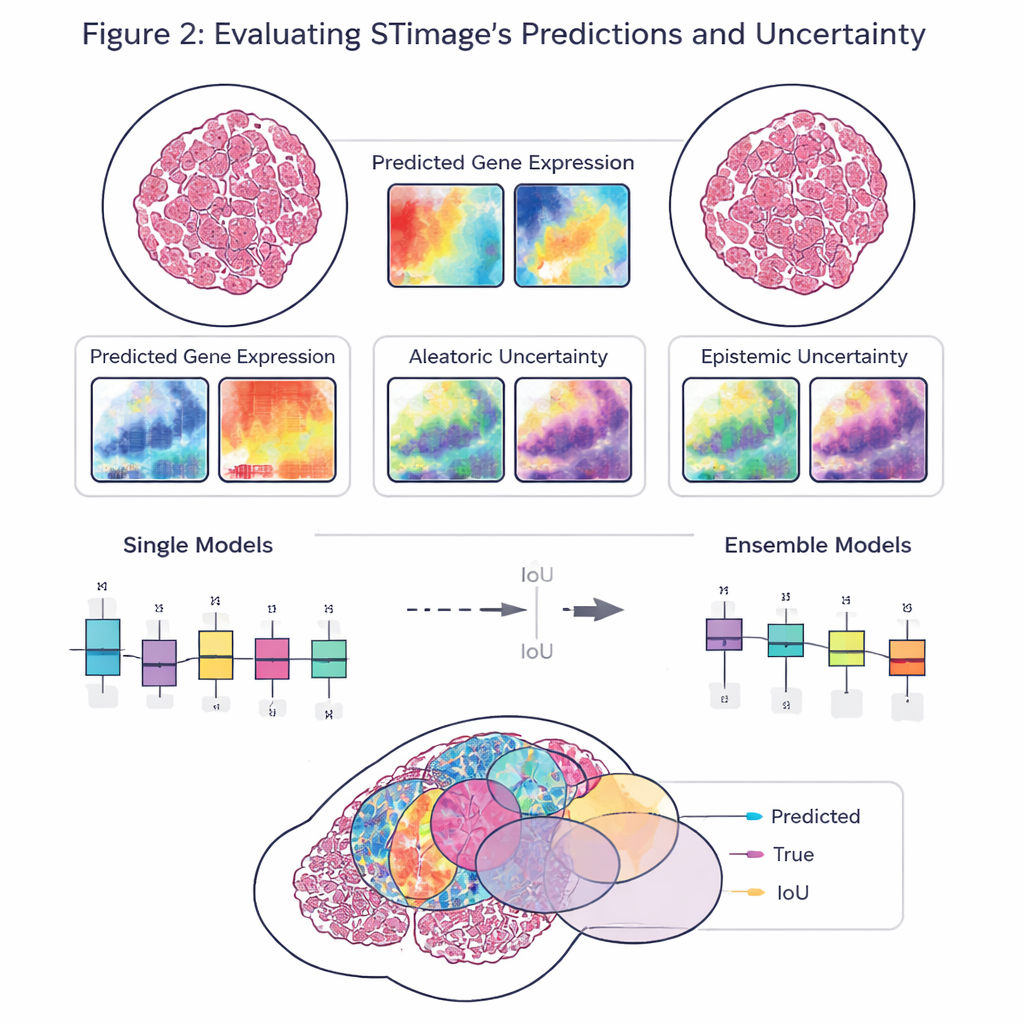
Seeing Inside Tumors: Cells, Survival, and Drug Response
STimage goes beyond gene prediction to infer which cell types occupy each region, using high-resolution datasets where each cell’s identity is known. The model could distinguish cancer cells from immune and supporting cells and map their arrangement across a slide. The authors then applied STimage to large collections of routine cancer images from The Cancer Genome Atlas. Even without spatial measurements, the AI’s predicted gene profiles were closely aligned with real bulk gene data. These predictions were strong enough to group patients into higher- and lower-risk categories and to help distinguish those more likely to respond completely to certain breast cancer therapies.
Why This Matters for Future Patients
For patients and clinicians, the promise of STimage is a kind of “molecular overlay” on the familiar pathology slide. Instead of ordering multiple expensive tests, a single scanned image could one day reveal where aggressive gene programs are active, how immune cells are distributed, and which markers point to better or worse outcomes or different drug responses. While the method is still being refined and correlation with true measurements is not perfect, its ability to capture spatial patterns, estimate its own uncertainty, and highlight which cells drive its predictions makes it a practical step toward more informative, transparent digital pathology.
Citation: Tan, X., Mulay, O., Xie, J. et al. Robust and interpretable prediction of gene markers and cell types from spatial transcriptomics data. Nat Commun 17, 1781 (2026). https://doi.org/10.1038/s41467-026-68487-0
Keywords: digital pathology, spatial transcriptomics, cancer biomarkers, deep learning, tumor microenvironment