Clear Sky Science · en
PTRAMP, CSS and Ripr form a conserved complex required for merozoite invasion of Plasmodium species into erythrocytes
Why this matters for fighting malaria
Malaria parasites must slip inside our red blood cells to cause disease. This study uncovers a tiny set of proteins that form a shared “boarding bridge” used by several major malaria species, including those that infect humans. By revealing how this bridge is built and how antibodies can sometimes block it, the work points to new ways to design vaccines that might protect against more than one type of malaria at once. 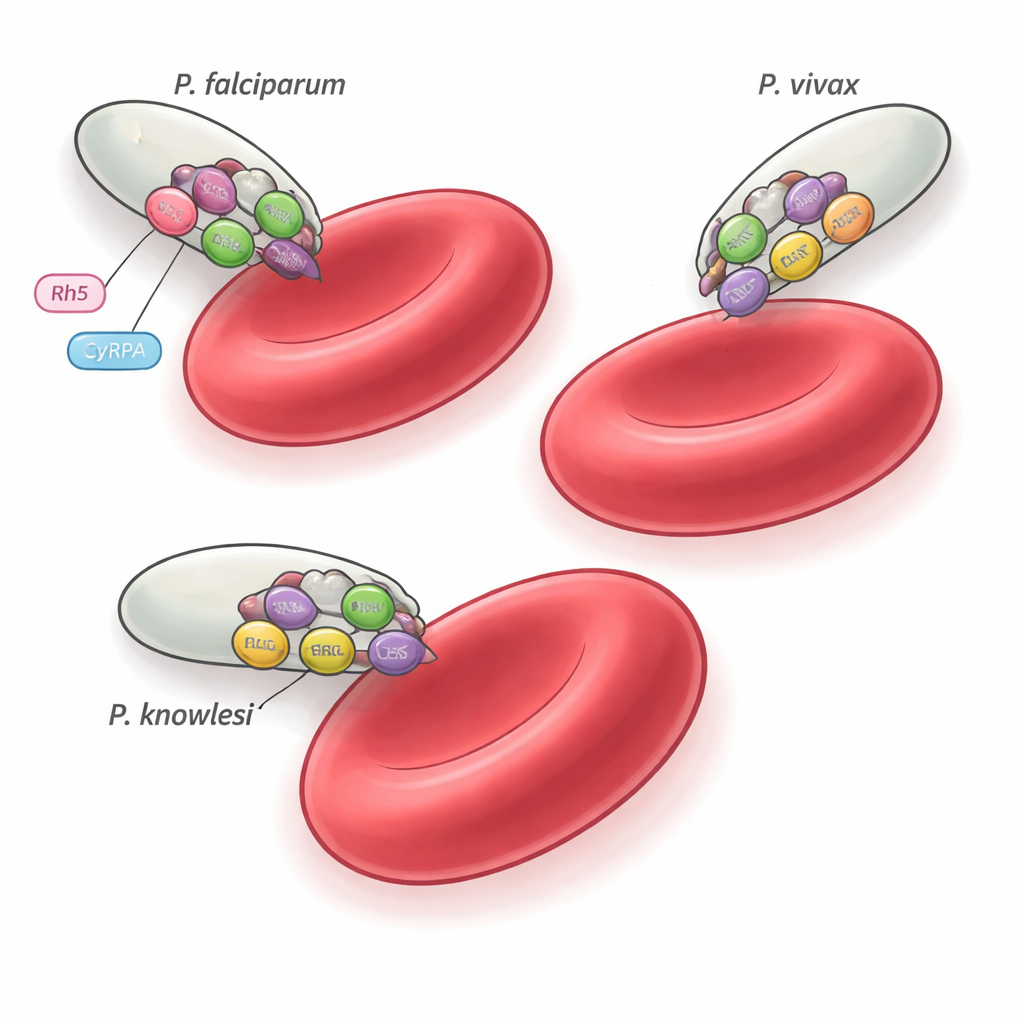
A common toolkit across many malaria parasites
There are more than 200 Plasmodium species, but only a handful, such as P. falciparum, P. vivax, and P. knowlesi, infect humans. These species sit on different branches of the parasite family tree and favor different types of red blood cells, yet all must invade erythrocytes with exquisite precision. Earlier work in P. falciparum identified a five-part assembly, the PCRCR complex, that is essential for invasion and locks onto a human red blood cell receptor called basigin. One piece of that complex, a protein named Rh5, is unique to P. falciparum and its close relatives, raising a puzzle: what do other malaria species use instead? The current study focuses on three companion proteins—PTRAMP, CSS, and Ripr—that are conserved across the genus and tests whether they form a universal invasion machine.
Building a three-part invasion bridge
Using sequence searches across many parasite genomes, the authors show that PTRAMP, CSS, and Ripr are present in all major Plasmodium lineages, unlike Rh5, which is restricted to one subgroup. Structure prediction with AlphaFold, combined with detailed biochemical measurements, reveals that PTRAMP and CSS join to form a stable pair, held together by a specific disulfide bond between two conserved cysteines. This two-protein platform then clamps onto the tail end of Ripr, creating a three-part PTRAMP–CSS–Ripr (PCR) complex. Experiments with proteins from P. falciparum, P. vivax, and P. knowlesi show that this complex forms in all three species with high affinity, and that only a small region at the C‑terminal end of Ripr is needed for tight binding. 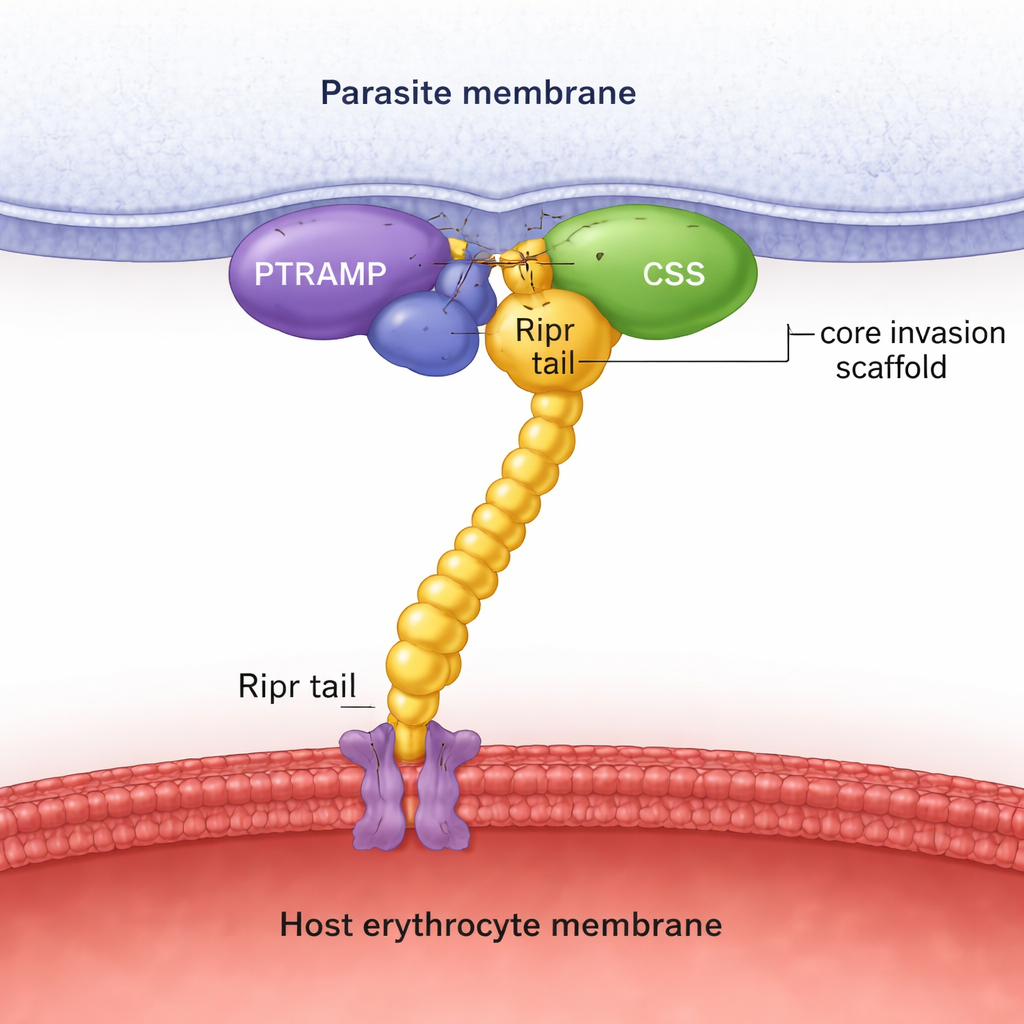
Seeing the structure in atomic detail
To move beyond predictions, the team solved crystal structures and collected cryo–electron microscopy images. The crystal structure of the P. vivax PTRAMP–CSS pair shows exactly how a short stretch of PTRAMP threads across CSS and forms the key disulfide linkage. Another structure captures a potent antibody bound to two of Ripr’s growth factor–like domains, mapping an inhibitory epitope on the Ripr tail. Cryo‑EM analysis of the P. knowlesi PCR complex confirms the overall shape predicted by AlphaFold: PTRAMP and CSS sit near the parasite membrane and grip Ripr, whose elongated body extends toward the host cell. Together, these structural snapshots support the idea that the PCR trio forms a rigid scaffold that physically spans the gap between parasite and red blood cell during invasion.
Antibodies that recognize more than one species
Because these proteins are conserved, the authors asked whether human infections naturally generate antibodies that cross‑react between species. Plasma from patients infected with P. falciparum, P. vivax, or P. knowlesi showed strong antibody responses to CSS and Ripr from multiple species, suggesting that people develop antibodies against shared regions of the PCR complex. The team then produced monoclonal antibodies and nanobodies targeting P. vivax PTRAMP, CSS, and Ripr, and tested whether these could block parasite growth in culture. Some antibodies, especially one named 5B3 that binds Ripr’s tail, were able to inhibit invasion by P. knowlesi and, at higher concentrations, by P. falciparum. Surprisingly, the same antibodies did not stop P. vivax or the closely related monkey parasite P. cynomolgi, even though they could bind the proteins from those species.
A shared scaffold with species-specific add‑ons
The functional tests show that the PCR complex itself does not grab onto red blood cells; instead, it likely serves as a conserved structural scaffold onto which each species bolt their own receptor‑binding partners. In P. falciparum, for example, the PCR trio partners with CyRPA and Rh5 to engage basigin on human erythrocytes. In P. vivax and P. knowlesi, the equivalent receptor‑binding proteins and host targets remain unknown, but the same PTRAMP–CSS–Ripr core appears to organize the invasion machinery. The fact that some antibodies can cross‑react yet only inhibit certain species underscores how subtle differences in complex assembly and timing can alter vulnerability to immune attack.
What this means for future malaria vaccines
For a non‑specialist, the take‑home message is that malaria parasites of very different species share a common “plug” they use to enter red blood cells—made of the PTRAMP, CSS, and Ripr proteins—even though they connect that plug to different receptors on the cell surface. This conserved plug is now structurally mapped and, in some cases, can be hit by antibodies that work across species. While more work is needed to identify the full set of partner proteins and to improve inhibitory antibodies, the discovery of a universal invasion scaffold opens the door to vaccine designs that might protect against multiple kinds of malaria rather than just one.
Citation: Seager, B.A., Lim, P.S., Xiao, X. et al. PTRAMP, CSS and Ripr form a conserved complex required for merozoite invasion of Plasmodium species into erythrocytes. Nat Commun 17, 1780 (2026). https://doi.org/10.1038/s41467-026-68486-1
Keywords: malaria invasion, Plasmodium proteins, merozoite entry, cross-species vaccine, erythrocyte infection