Clear Sky Science · en
A stochastic mechanism drives fast substrate translocation in the AAA+ machine ClpB
How Cellular Machines Keep Proteins Moving
Inside every cell, tiny molecular machines constantly tug, pull, and reshape proteins to keep them from clumping and to repair damage. One such machine, called ClpB, helps rescue proteins that have become tangled—an essential job for cell survival under stress. This study asks a simple but deep question: how does ClpB turn the chemical fuel ATP into the mechanical work of pulling a protein chain through its central pore, and does it do so in neat, clockwork steps or in a more random, diffusive way?
A Protein Rescue Ring at Work
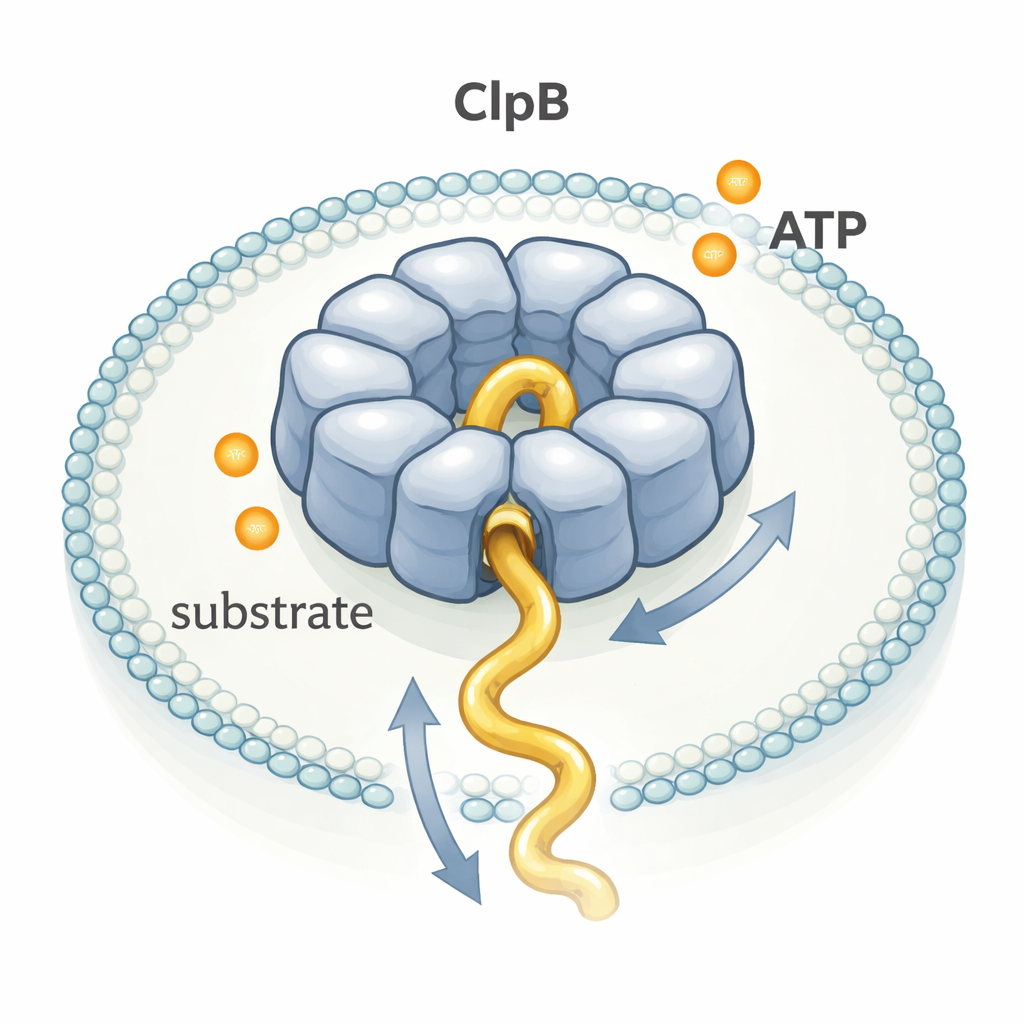
ClpB belongs to a large family of enzymes called AAA+ machines that assemble into ring-shaped structures with a central tunnel. Many have been pictured in exquisite detail using cryo–electron microscopy, leading to a popular "hand-over-hand" model: each ATP molecule burned would drive a small, precise step, like a team of people pulling a rope in sequence. Yet other experiments hinted that ClpB might move proteins far faster than its slow rate of ATP consumption would seem to allow. To resolve this puzzle, the authors watched single ClpB molecules in real time as they moved a flexible test protein, κ-casein, through their pore.
Watching One Molecule at a Time
The researchers trapped individual ClpB rings and single κ-casein molecules together inside tiny lipid bubbles stuck to a glass surface. They attached fluorescent dyes to specific points on ClpB and on the protein chain, and then used single-molecule FRET, a technique that reports distance changes between dyes on the nanometer scale. When κ-casein slipped into the ClpB pore, the dyes came close and FRET shot up as a brief spike; when the chain left, the signal dropped again. By measuring the duration and height of thousands of such spikes, they could infer how quickly and how far segments of the protein threaded through the pore.
Fast, Weakly Fueled, and Nearly Temperature-Free
Surprisingly, the translocation events were extremely fast: the labeled segment of κ-casein typically traversed several nanometers of the pore in about 1–2 milliseconds—roughly a thousand times faster than the average time between ATP hydrolysis events in ClpB. Most events were this short, and they followed a broad, power-law distribution rather than a single characteristic timescale. Changing the temperature from 10 to 32 °C barely slowed these motions, implying that the energy barrier for movement is tiny compared with that of classic "power-stroke" motors like kinesin. Likewise, lowering the ATP concentration strongly reduced how often events occurred, but had almost no effect on how long each event lasted. ATP, in other words, controlled when ClpB engaged and how frequently it acted, more than the speed of individual threading bursts.
Back-and-Forth Motion with a Forward Bias
To see whether chains always move in one direction, the team performed more complex three-color FRET experiments, dyeing both ends of the pore and the substrate. This allowed them to distinguish whether κ-casein entered from the top or bottom of the ring and whether it passed all the way through or retreated. They discovered six distinct patterns: complete forward and backward threading, longer "partial" visits where the chain sampled both ends of the pore before leaving, and fleeting encounters near just one end. Roughly three quarters of complete events went in the forward direction, but a substantial fraction went backwards, revealing that ClpB allows bidirectional diffusion of its substrate with only a modest forward bias. When ATP was replaced by a slowly hydrolyzed analog, events became rare and much slower, and the forward preference essentially disappeared.
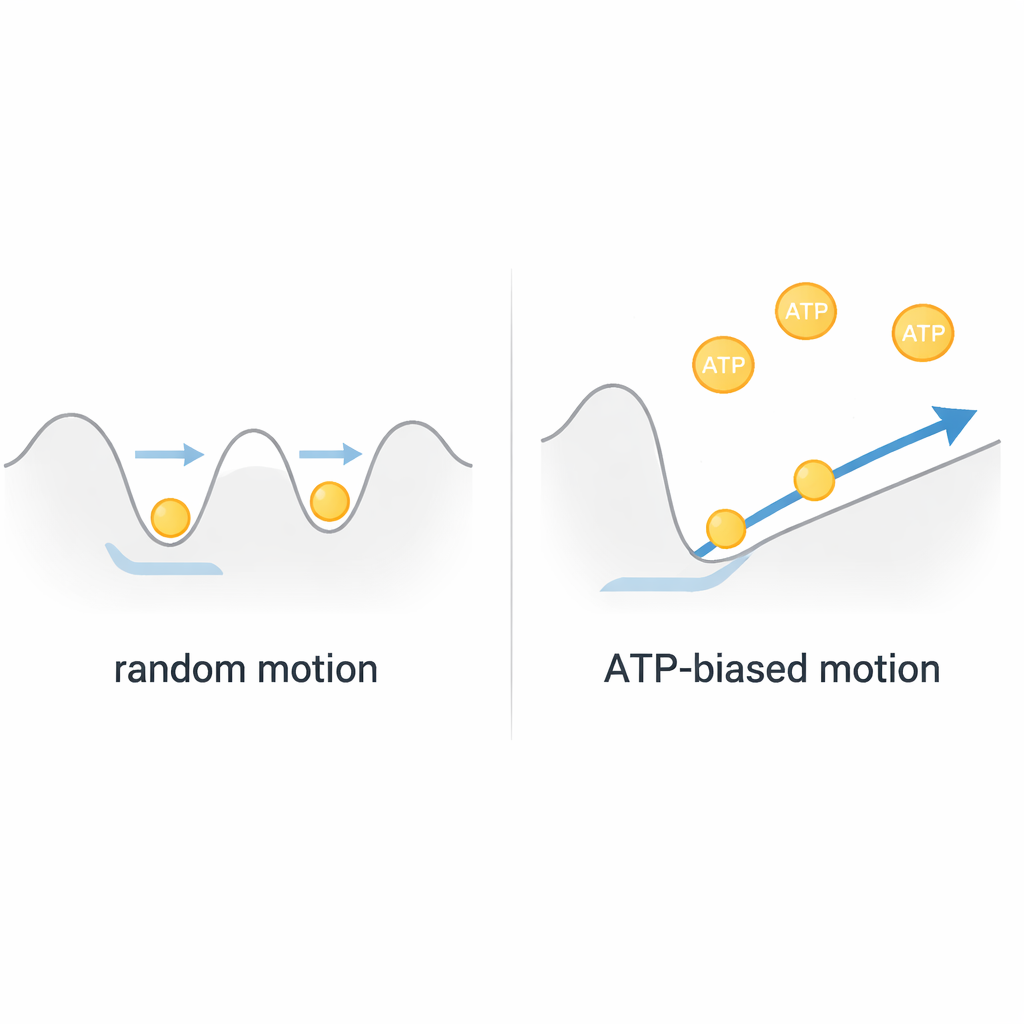
A Brownian Motor, Not a Simple Winch
Putting these observations together, the authors conclude that ClpB works more like a Brownian motor than a rigid, fuel-driven winch. The protein chain jiggles thermally within a shallow energy landscape inside the pore, moving back and forth at high speed. ATP does not directly power large, discrete pulling strokes; instead, it tunes the shape of this landscape via fast motions of internal "pore loops," subtly favoring movement in one direction. Because each complete threading event correlates with only about one or two ATP molecules consumed, ClpB achieves remarkably efficient, rapid transport by rectifying random motion rather than fighting it. This view reshapes how we think about many AAA+ machines, suggesting that biological nanomotors may often harness randomness instead of eliminating it.
Citation: Casier, R., Levy, D., Riven, I. et al. A stochastic mechanism drives fast substrate translocation in the AAA+ machine ClpB. Nat Commun 17, 1773 (2026). https://doi.org/10.1038/s41467-026-68478-1
Keywords: protein translocation, AAA+ machine, Brownian motor, molecular chaperone, single-molecule FRET