Clear Sky Science · en
Identification of thermotolerant non-canonical PAMs for robust one-pot CRISPR-Cas12a detection
Turning Up the Heat on DNA Tests
Fast, accurate tests for infections and genetic changes are vital for medicine, but today’s DNA tests often need complex lab gear and can miss rare or subtle targets. This study shows how simply running a CRISPR-based test a bit hotter unlocks many more useful DNA “handles,” making one-tube diagnostics faster, more sensitive, and able to tell apart even single-letter differences in genetic code. 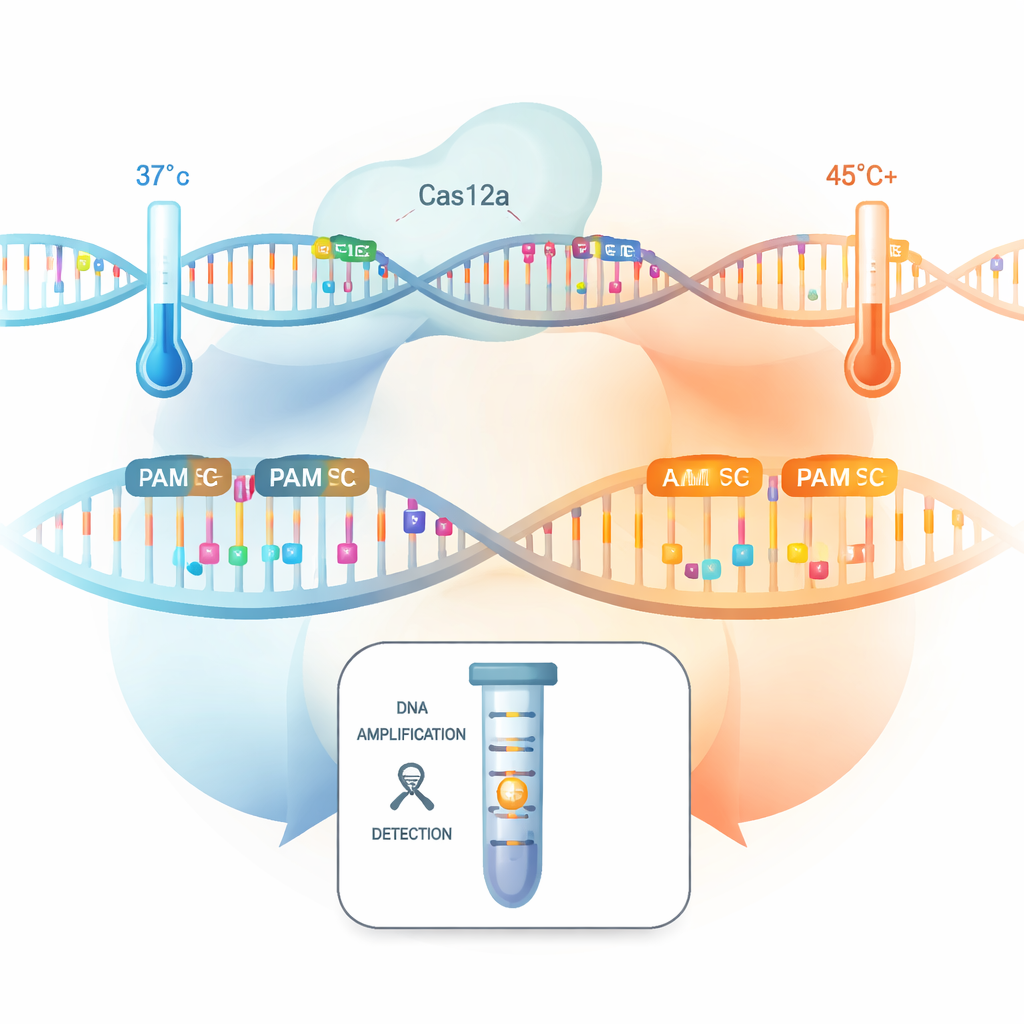
Why CRISPR Needs a Zip Code
CRISPR tools like Cas12a work by homing in on specific stretches of DNA, guided by a short RNA that acts like a search string. But they can only latch on and start working if a nearby four-letter tag, called a PAM, has the right sequence—traditionally a narrow set starting with “TTT.” That is like needing a house number before you can deliver a letter: if the right number is missing, the message never arrives. This PAM requirement limits where CRISPR can look, a serious drawback when doctors need to detect particular mutations, drug-resistance markers, or viral variants that do not sit next to these classic tags.
Discovering New “Doorways” at Higher Temperatures
The researchers systematically tested all 256 possible four-letter PAM combinations to see how well they powered Cas12a’s “collateral” cutting activity—its ability to slice many nearby DNA pieces once it finds its target, which is the basis for CRISPR diagnostic signals. At normal body temperature (37 °C), only a handful of non-standard PAMs performed as well as the classic ones. But when they raised the reaction temperature to about 45 °C, something striking happened: 82 different non-canonical PAMs suddenly produced strong signal-cutting activity, rivaling the standard tags. The team believes extra warmth loosens the local DNA structure and makes the Cas12a complex more flexible, lowering the barrier to recognizing these previously “weak” PAMs.
Strong Signaling, Gentle on the Template
Cas12a has two cutting modes. In “cis” mode, it snips the specific DNA it was told to find; in “trans” mode, once activated, it rapidly chops any nearby signal-carrying strands, which lights up the test. The researchers found that at higher temperatures, non-canonical PAMs gave very strong trans-cutting—great for a bright, fast readout—but kept cis-cutting of the main DNA relatively weak. That combination is ideal for one-tube tests, where DNA must first be amplified without being destroyed. They also showed that at these warmer settings Cas12a becomes much pickier about mismatches between the guide RNA and the target: even a single wrong letter can shut down the signal, especially when using non-canonical PAMs. 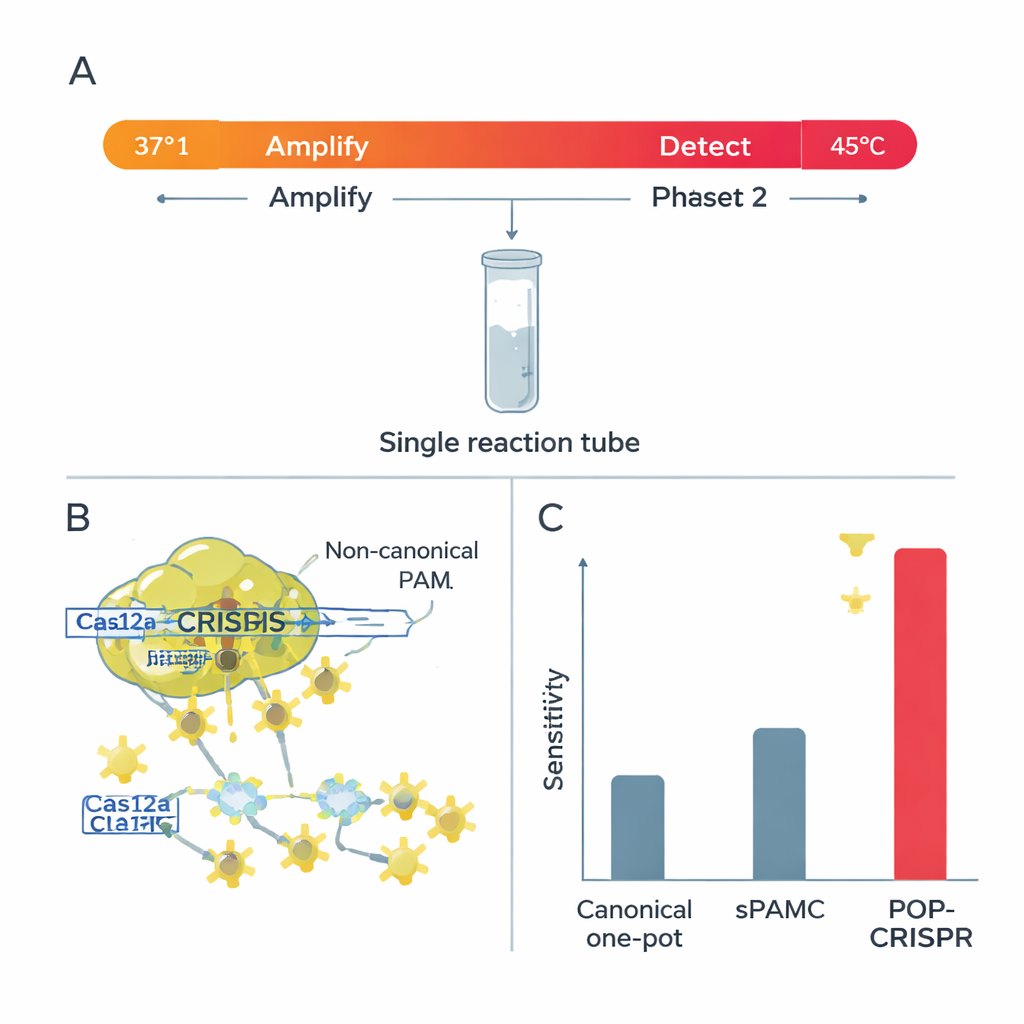
A Smarter One-Pot Test: POP-CRISPR
Building on these insights, the team created POP-CRISPR, a “poikilothermal” (variable-temperature) one-pot assay. The reaction starts around 37 °C, where an isothermal amplification method quietly multiplies any target DNA or RNA while Cas12a is relatively inactive. After about 10 minutes, the mix is heated to around 45 °C, a sweet spot where Cas12a—now guided to non-canonical PAM sites—turns on powerful trans-cutting and generates a strong fluorescent signal. This simple two-step temperature program boosted detection sensitivity by roughly tenfold compared with earlier one-pot CRISPR methods that also used weaker PAMs, and clearly outperformed conventional canonical-PAM one-pot formats.
From Bench to Bedside in Minutes
The POP-CRISPR system was tested on real clinical samples. It sensitively detected human papillomavirus (HPV-16) in vaginal swabs and Mycoplasma pneumoniae in respiratory swabs, including very low-level “gray zone” cases that challenge standard methods. Crucially, POP-CRISPR could distinguish drug-resistant Mycoplasma strains carrying a single-letter change in a ribosomal gene, even though no classic PAM sat near that mutation—a task that stymies many existing Cas12a tests. To make the approach field-friendly, the authors paired a rapid two-minute heat-and-Chelex lysis step (avoiding full DNA extraction) with a palm-sized fluorescence reader controlled by a smartphone. Together, this workflow can turn a raw swab into a digital yes/no result in about 20 minutes.
What This Means for Everyday Testing
By raising the reaction temperature and exploiting a much broader set of PAM “doorways,” this work turns Cas12a into a more flexible and precise sensor. POP-CRISPR delivers faster, more sensitive, and more specific nucleic acid detection in a single closed tube, while also expanding the range of genetic sites that can be targeted—including single-base mutations linked to drug resistance or cancer. For patients, that could translate into quicker on-site diagnosis and better-tailored treatments; for researchers and test developers, it offers a powerful new way to design CRISPR diagnostics that are both more robust and more discriminating.
Citation: Tian, T., Zhang, T., Zhang, W. et al. Identification of thermotolerant non-canonical PAMs for robust one-pot CRISPR-Cas12a detection. Nat Commun 17, 1771 (2026). https://doi.org/10.1038/s41467-026-68476-3
Keywords: CRISPR diagnostics, Cas12a, PAM sites, point-of-care testing, nucleic acid detection