Clear Sky Science · en
Coiled-coil heterodimer-mediated split base editing systems enable flexible and robust nucleotide substitutions
Fixing DNA with Fewer Side Effects
Many diseases are caused by tiny typos in our DNA—single-letter mistakes that can have life‑threatening consequences. Scientists have powerful tools, called base editors, that can correct these typos without cutting the DNA, but these tools are bulky and hard to deliver safely into the body. This paper introduces a new way to split base editors into smaller, smarter pieces that still work as well as—or even better than—the original, opening new paths toward treating genetic diseases in the liver, muscles, and beyond.
Why Size Is a Problem for Gene Fixing Tools
Base editors combine parts of the CRISPR system with enzymes that can swap one DNA letter for another, such as changing an A to a G or a C to a T. These precise swaps are promising for treating conditions like high cholesterol or muscular dystrophy. However, a leading delivery vehicle for gene therapy, adeno‑associated virus (AAV), can only carry about 4.7 thousand DNA letters, while standard base editors are larger than that. Earlier work tried splitting these editors using special protein “scissors” called inteins, or by shrinking them with miniature enzymes, but these approaches often reduced efficiency, added design complexity, or limited where in the genome the tools could act.
Using Molecular Velcro to Rebuild the Editor
To solve this, the researchers designed a “split” base editor held together by tiny protein zippers known as coiled‑coil heterodimers—like a piece of molecular Velcro. They separated the base editor into two parts: one carrying the DNA‑targeting Cas9 nickase and another carrying the editing enzyme. Short coiled‑coil peptides on each half seek each other out and snap together when both halves reach the same stretch of DNA. This creates coiled‑coil base editors (CC‑BEs), including types that edit C (CC‑CBE), A (CC‑ABE), and more complex variants that can make different kinds of letter changes. 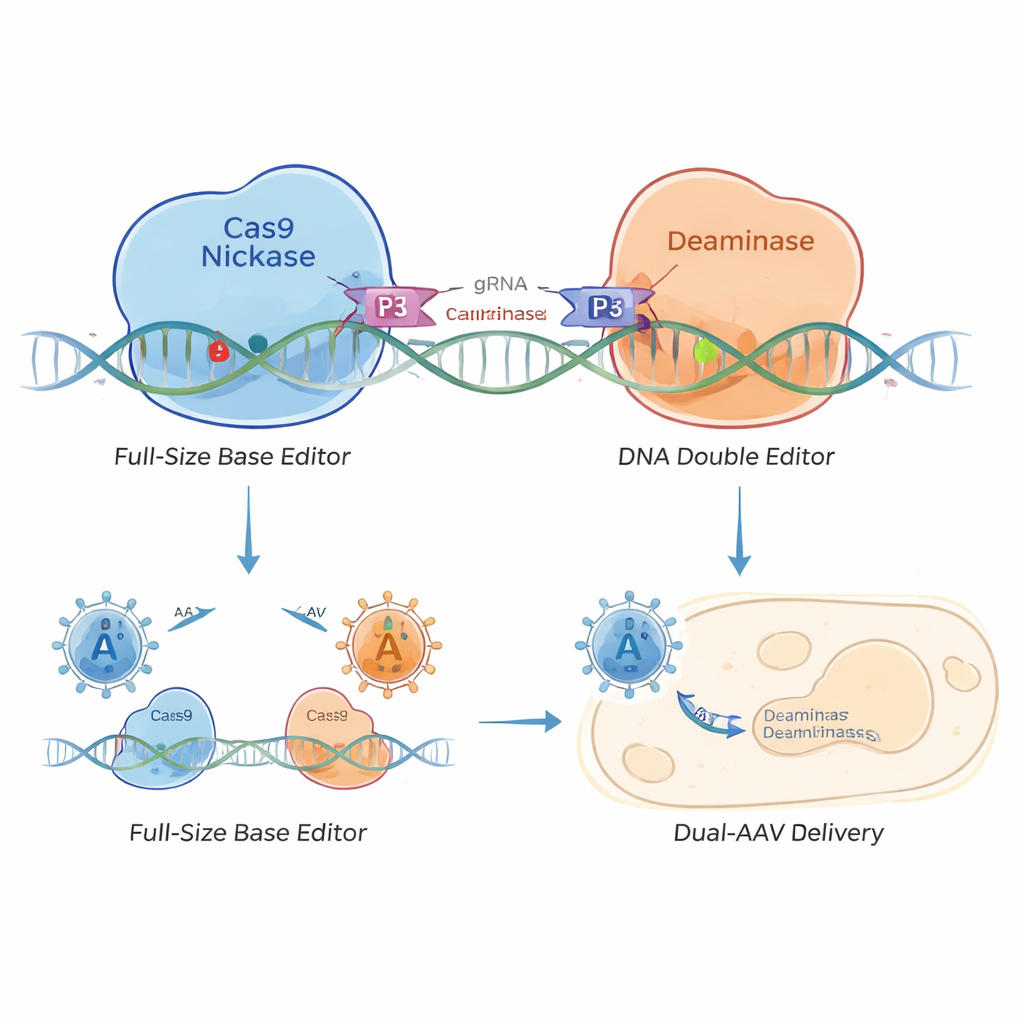
Flexible, Powerful, and Precise Editing
The team tested CC‑BEs in several cell types, including human cells and primary pig cells, and with different forms of Cas9 that recognize a wider range of DNA sequences. The CC‑CBE versions not only worked reliably at many sites but also showed a broader “editing window,” meaning they could change target letters across a wider stretch of DNA, giving scientists more flexibility in choosing guide RNAs. The CC‑ABE versions reached similar efficiency to the best existing adenine editors while often producing fewer off‑target by‑products. The researchers also adapted the coiled‑coil strategy to newer editor designs—such as compact TadCBE and precise ABE9—and again found that splitting the tools with coiled‑coil pairs preserved or improved performance.
From Cell Dishes to Living Mice
Crucially, the authors showed that these split editors work inside living animals using dual‑AAV delivery, where each viral particle carries half of the editor. In mice, a CC‑ABE targeting the Pcsk9 gene in the liver achieved A‑to‑G edits at rates up to about 70%, sharply lowering Pcsk9 protein and blood LDL cholesterol without signs of liver damage. 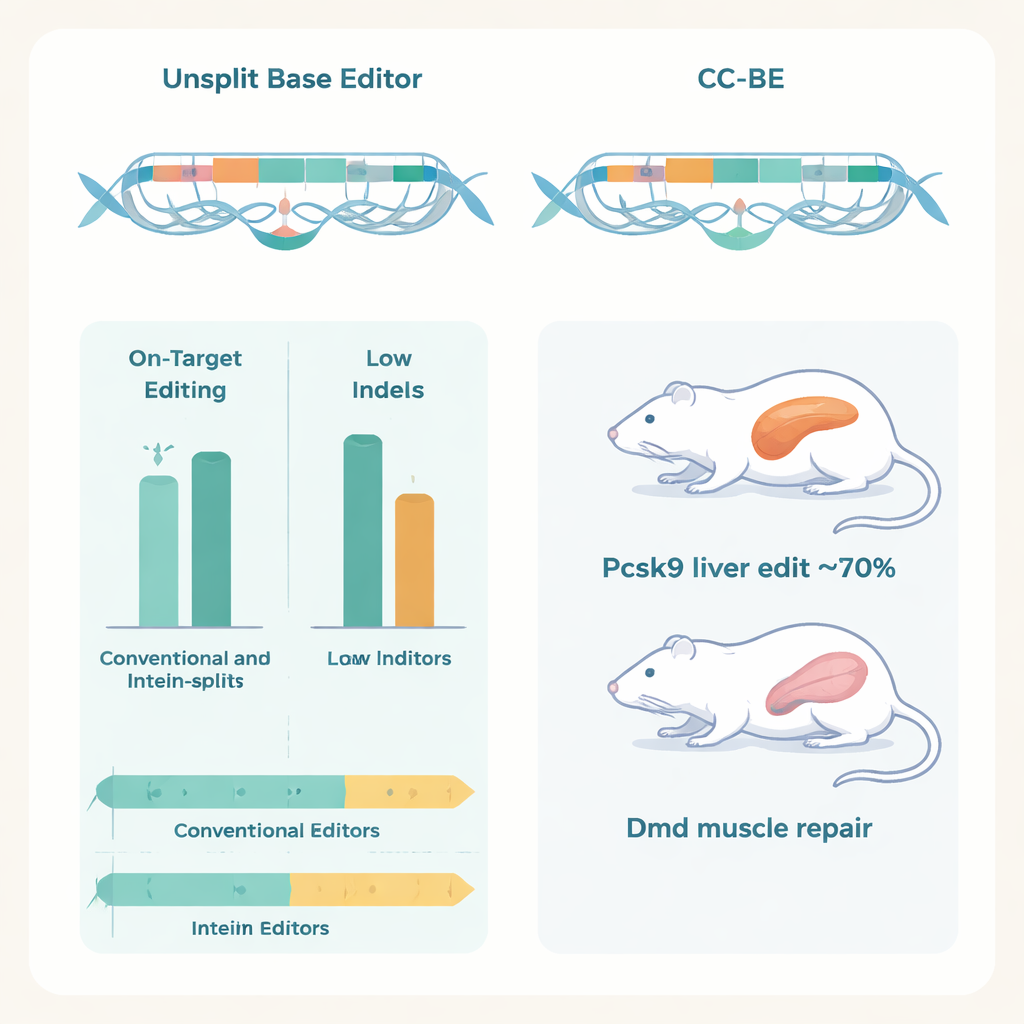
A New Path Toward Practical DNA Repair
In essence, this study presents a simple but powerful engineering trick: use small coiled‑coil “hooks” to split large base editors into AAV‑sized modules that reunite only where they are needed. For non‑experts, the takeaway is that scientists can now package highly capable DNA‑repair tools into clinically proven viral carriers without sacrificing accuracy or strength. This coiled‑coil approach could make it easier to design treatments for a wide range of single‑letter genetic diseases in organs like the liver, heart, and muscle, bringing finely tuned gene correction a step closer to real‑world therapies.
Citation: Mu, S., Li, Q., Chen, M. et al. Coiled-coil heterodimer-mediated split base editing systems enable flexible and robust nucleotide substitutions. Nat Commun 17, 1765 (2026). https://doi.org/10.1038/s41467-026-68469-2
Keywords: base editing, gene therapy, CRISPR, AAV delivery, genetic disease