Clear Sky Science · en
Inferring chromatin architecture at a single locus through probabilistic in situ DNA localization
How DNA’s 3D Shape Controls When Genes Turn On
Our DNA is often pictured as a straight ladder, but inside cells it folds into intricate loops and coils. These shapes matter: they help decide which genes turn on, when, and where. This study introduces a new way to look at the tiny 3D arrangements of DNA around a single gene inside developing fruit fly embryos, revealing how subtle shifts in DNA folding can change patterns of gene activity that shape the body plan.
Watching Gene Control in a Developing Embryo
As an embryo develops, thousands of genes must switch on and off at just the right moments. Many of these decisions are made by short stretches of DNA called enhancers, which can lie tens of thousands of letters away from the genes they control. To work, enhancers must physically approach their target gene in 3D space, looping the DNA so distant sites touch. But these loops are tiny, dynamic, and hard to see. The authors focused on a single gene, brinker (brk), in the fruit fly Drosophila. This gene helps pattern the early embryo, turning on in a stripe along the side of the egg. Three nearby DNA control elements—two enhancers (E1 and E2) and a promoter-proximal element (PPE) next to the gene—cooperate to produce this precise pattern.
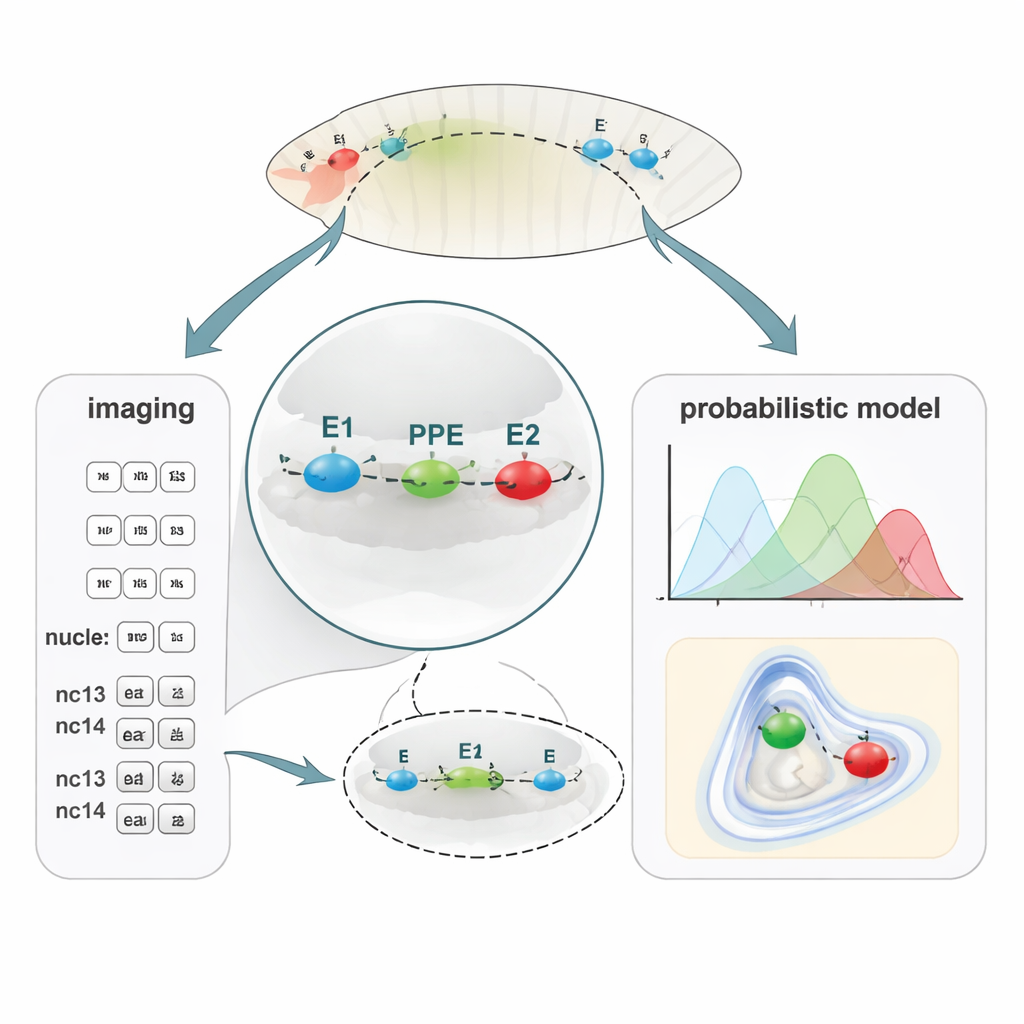
A New Way to Map Tiny DNA Distances
To link DNA folding with gene activity, the team developed PLOTTED (Probabilistic Localization of Oligopaint Tagged Target Element Distances). First, they used a DNA-labeling method called Oligopaint FISH to attach three different fluorescent dyes to the E1, PPE, and E2 regions in fixed fly embryos. Using a super-resolution confocal microscope, they measured 3D distances between these three colored spots in tens of thousands of nuclei across embryos at successive nuclear cycles, from just before gene activity begins (pre-nc13) through later stages (nc13 and nc14). They then fed all these distances into a custom computational pipeline that filters out noisy measurements and builds probability maps showing where each element is most likely to sit relative to the others. Instead of a single static “loop,” PLOTTED produces a landscape of likely chromatin shapes at each developmental stage.
When DNA Compacts, the Gene Wakes Up
In normal embryos, the researchers found that as the embryo reaches nuclear cycle 13, both enhancers move closer to the PPE: the local DNA neighborhood around the brk gene becomes more compact. After this point, distances between the three elements remain relatively stable. Importantly, this timing matches the onset of brk expression, suggesting that the tightening of the DNA configuration helps enable the gene to turn on. PLOTTED also revealed that this compact arrangement appears more frequently in regions of the embryo where brk is active, and looser configurations dominate where the gene is repressed, reinforcing the link between 3D architecture and transcription.
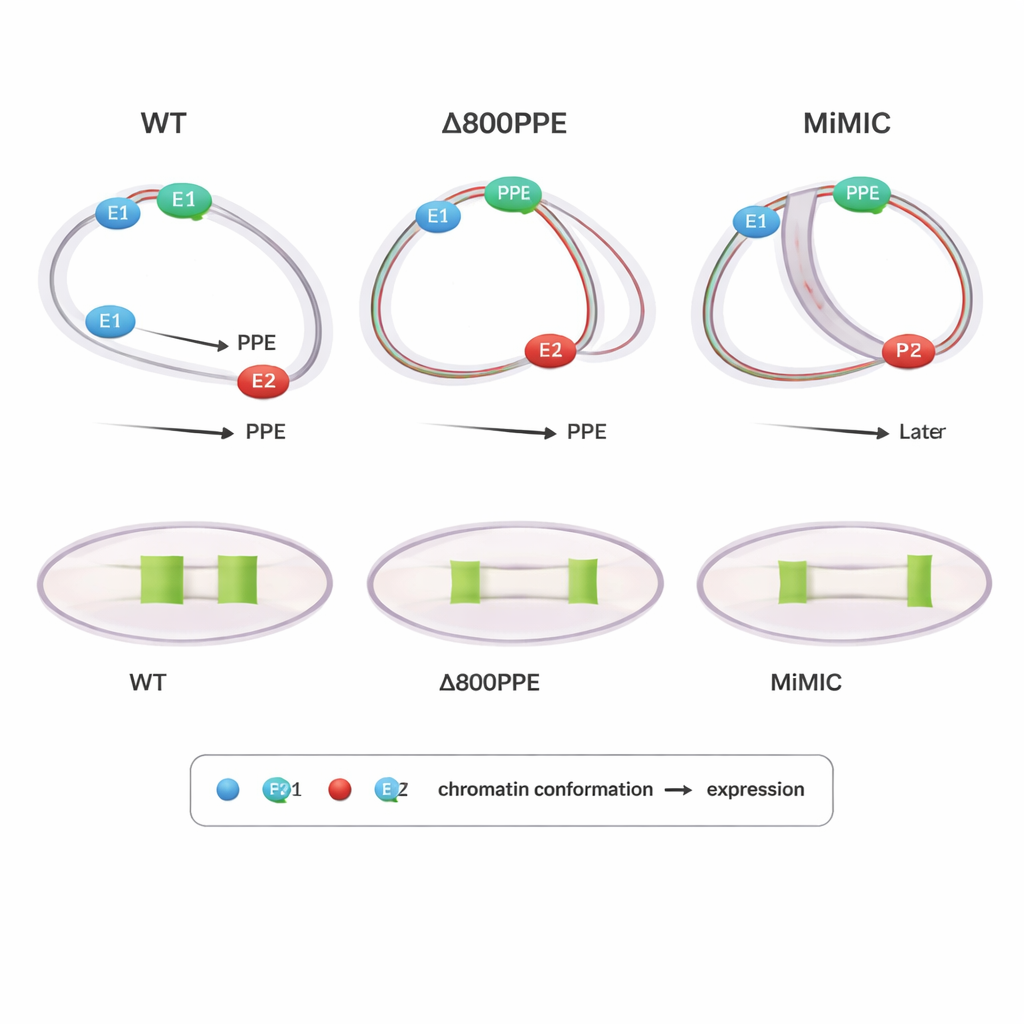
Mutations Reveal How Timing and Position Matter
To probe cause and effect, the authors examined flies carrying engineered changes at the brk locus. In one mutant, 800 base pairs of the PPE were deleted, weakening this central element; in another, a 7.3 kilobase DNA cassette (MiMIC) was inserted between E1 and the PPE, effectively pushing them farther apart and adding an extra promoter. Both mutants showed delayed or reduced brk expression. PLOTTED showed why: in the PPE-deletion line, the compaction of distances between the PPE and both enhancers occurred later than normal, and at late stages the PPE stayed too close to E1, preventing E2 from driving the broad expression pattern seen in wild type. In the MiMIC line, the PPE associated too early and strongly with E2 and only later drew closer to E1, again disturbing the normal handoff between enhancers. These results suggest that not just whether elements come together, but when and to which partner they are closest, is crucial for correct gene output.
DNA Folding Varies Across the Embryo
Because PLOTTED preserves spatial information inside intact embryos, the team could also ask whether DNA architecture differs in various body regions. Comparing lateral zones where brk is active to ventral zones where it is repressed, they found that the three elements sit closer together in active regions and spread farther apart in repressed domains. Along the head-to-tail axis, they observed that the E1–PPE distances change differently in the front versus the back of the embryo, hinting that regional cues tune chromatin architecture to fine-tune gene expression patterns. These findings support a view in which the 3D arrangement of regulatory DNA is context-dependent, tracking both time and position in the developing organism.
Why This Matters Beyond Fruit Flies
In simple terms, this study shows that how DNA folds around a single gene is closely tied to when and where that gene turns on. The new PLOTTED method provides a practical way to map these tiny DNA neighborhoods in intact tissues using widely available microscopes and straightforward chemistry, combined with powerful probabilistic modeling. Although demonstrated in fly embryos, the approach can be applied to many organisms and disease models. As scientists increasingly discover that misfolded chromatin underlies developmental disorders and cancers, tools like PLOTTED will help reveal how small shifts in the 3D layout of enhancers and genes can ripple out into large changes in cell fate and health.
Citation: Le, M.T., McGehee, J., Dunipace, L. et al. Inferring chromatin architecture at a single locus through probabilistic in situ DNA localization. Nat Commun 17, 1752 (2026). https://doi.org/10.1038/s41467-026-68460-x
Keywords: chromatin architecture, enhancer-promoter interactions, Drosophila embryogenesis, gene regulation, super-resolution imaging