Clear Sky Science · en
A common DNA deletion altering the 3’UTR of mdr1 is associated with reduced mefloquine susceptibility in P. vivax parasites from Cambodian patients
Why this malaria study matters
For people living in or traveling to malaria-endemic regions, effective drugs are a lifeline. But malaria parasites are constantly evolving ways to survive treatment. This study focuses on Plasmodium vivax, the main cause of malaria outside Africa, and uncovers a subtle genetic change that may already be helping the parasite tolerate one important drug, mefloquine. Understanding this shift now could help public health officials stay a step ahead of emerging drug resistance.
A stubborn parasite and its shrinking treatment options
Malaria in humans is caused by several related parasites, but P. vivax is especially difficult to eliminate. It can hide in the liver for months before reappearing in the blood, and often circulates at levels too low for standard tests to spot. For decades, the frontline drug chloroquine worked well against P. vivax, but signs of resistance have spread. As a result, the World Health Organization now recommends using artemisinin-based combination therapies (ACTs), which pair a fast-acting drug with a longer-lasting partner such as mefloquine. Because laboratory culture of P. vivax is challenging, scientists rely heavily on patient samples and genetic clues to track how the parasite is responding to these medicines.
A tiny missing piece of DNA in a key drug-resistance gene
The researchers sequenced the genomes of 206 P. vivax samples taken from malaria patients in Cambodia between 2021 and 2023. They searched for stretches of DNA that were either deleted or duplicated. Among many rearrangements, one stood out: more than 80% of the parasites carried a small DNA deletion located just after the end of the coding region of a gene called mdr1 (multidrug resistance 1). This gene encodes a transporter protein that can pump various compounds, including drugs, across a membrane inside the parasite. Unlike previously reported mutations, this 837-base-pair deletion did not change the protein sequence itself. Instead, it lay in the so‑called 3' untranslated region (3'UTR), a segment of the gene that is not translated into protein but can strongly influence how much protein is ultimately made.
Independent deletions and altered messages from the same gene
To see whether similar changes appeared elsewhere, the team reanalyzed nearly 830 P. vivax genomes from 25 countries. They discovered another, shorter deletion overlapping the same downstream region of mdr1 in parasites from South America. Detailed comparisons and confirmatory PCR tests showed that the Southeast Asian and South American deletions were distinct, suggesting they arose independently—an indication that natural selection may favor alterations in this regulatory region. Using RNA sequencing on 95 Cambodian infections, the researchers showed that parasites with the deletion produced an alternative version of the mdr1 messenger RNA, with a different 3'UTR. After accounting for transcript length, parasites carrying the deletion had nearly twice as much mdr1 mRNA as those without it, implying that the missing DNA or the new 3'UTR sequence helps stabilize or boost expression of the gene.
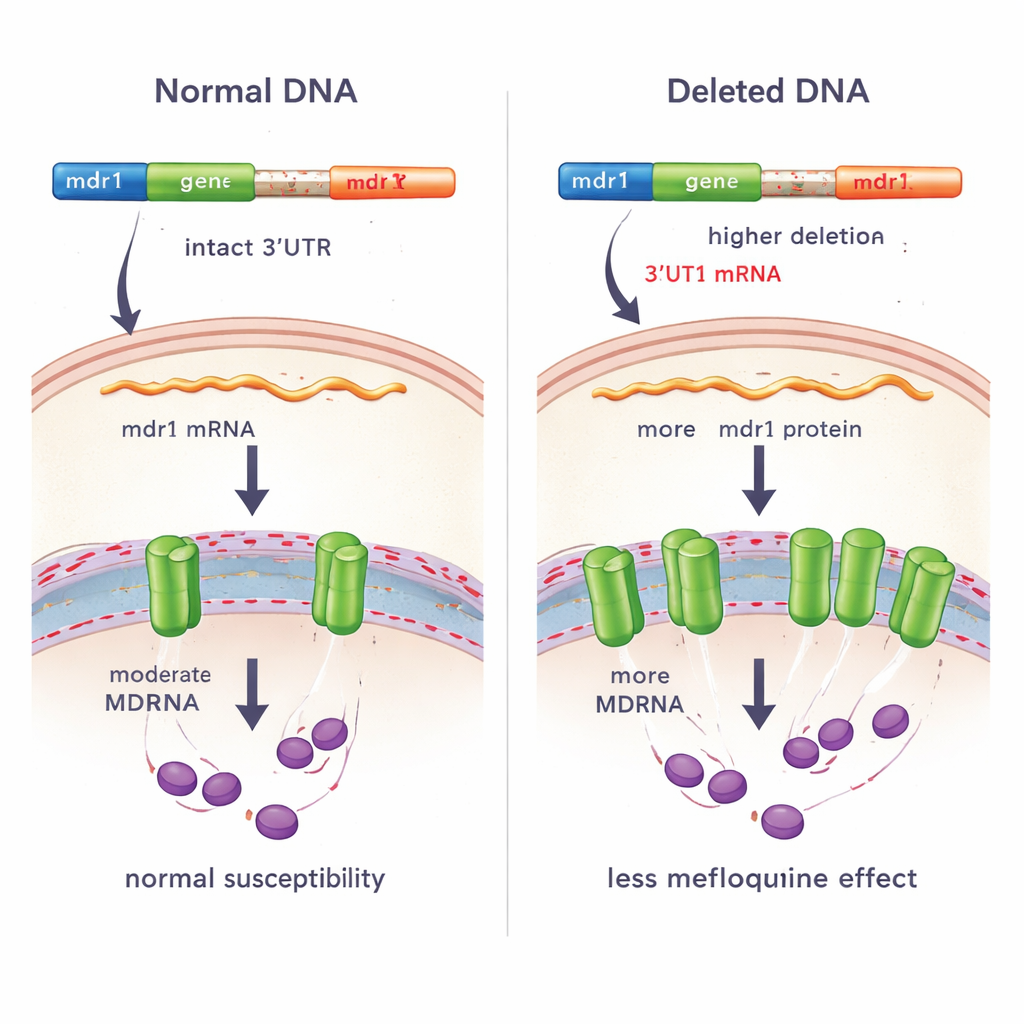
Linking the deletion to drug response in patients and lab tests
Because mdr1 has been implicated in resistance to several antimalarial drugs, the team investigated whether the deletion was tied to how parasites responded to treatment. They first compared how quickly parasites were cleared from patients' blood after artesunate treatment. Among 167 infections, there was no significant difference in clearance rates between parasites with and without the deletion, suggesting that this genetic change does not directly affect sensitivity to artesunate itself. However, in a smaller set of 14 infections where the researchers measured parasite growth in the presence of mefloquine outside the body, parasites carrying the deletion tended to have higher IC50 values—meaning they required more drug to inhibit growth—than parasites without the deletion. This pattern is consistent with reduced susceptibility to mefloquine, although the sample size was limited.
A worrying rise over time and what it could mean
To understand how widespread the deletion has become, the authors screened 592 archived P. vivax samples collected across Cambodia from 2014 to 2024. Before mefloquine was introduced as part of ACTs for vivax malaria around 2016–2017, roughly 30% of parasites carried the deletion. After the switch, the frequency climbed above 60% and then stabilized, a trend that was statistically strong and remained even when focusing on eastern Cambodia alone. The deletion was already present at low levels before mefloquine was widely used against P. vivax, possibly due to indirect exposure when the drug was used to treat P. falciparum or because the change also affects responses to other medicines. Together with recent reports of slightly slower parasite clearance after artemisinin treatment, the results raise concern that P. vivax in Cambodia may be moving toward decreased effectiveness of the current artesunate–mefloquine combination.
What this means for malaria control
For non-specialists, the core message is that even when a parasite’s main drug-resistance gene looks intact, small changes in the nearby “control wiring” of its DNA can alter how strongly the gene is turned on. In this case, a common deletion in a non-coding region of mdr1 appears to boost the gene’s activity and is associated with slightly weaker effects of mefloquine on P. vivax. While there is not yet clear evidence of treatment failure in Cambodia, the rising frequency of this deletion suggests that drug pressure may be quietly favoring parasites with this trait. Recognizing and monitoring such subtle genetic warning signs early can help health authorities adjust drug policies, design better surveillance tools, and protect the dwindling arsenal of effective antimalarial treatments.
Citation: Ko, K., Tebben, K., Andrianinarivomanana, T. et al. A common DNA deletion altering the 3’UTR of mdr1 is associated with reduced mefloquine susceptibility in P. vivax parasites from Cambodian patients. Nat Commun 17, 1748 (2026). https://doi.org/10.1038/s41467-026-68456-7
Keywords: Plasmodium vivax, malaria drug resistance, mefloquine, mdr1 gene, Cambodia