Clear Sky Science · en
Af-CUT&Tag: a sensitive and antibody-free chromatin profiling method using genetically encoded tags and high-affinity binders fused to Tn5
Peeking Inside Our DNA Control Room
Every cell in your body constantly decides which genes to turn on or off, shaping everything from liver repair to cancer risk. Scientists study these decisions by mapping where key proteins sit on our DNA, but the best tools for doing this have long depended on fragile, expensive antibodies that do not always work well. This paper introduces a new, antibody-free way to chart these DNA–protein interactions, called Af-CUT&Tag, that is more sensitive, more reliable, and powerful enough to work even on tiny samples and individual cells.
Why Antibodies Hold Us Back
Classical chromatin mapping methods need antibodies—large Y-shaped proteins that recognize a specific target protein—to guide an enzyme to the right spots on DNA. When antibodies work perfectly, they allow researchers to see where transcription factors and other regulators are bound across the genome. But in practice, antibodies can be hard to obtain, inconsistent in quality, and blocked by common chemical tweaks on proteins, such as phosphorylation or acetylation. These issues can blur the resulting maps, limit the proteins that can be studied, and make it difficult to compare results between labs or experiments.
Tiny Tags and Smart Binders Replace Antibodies
Af-CUT&Tag sidesteps antibodies entirely by giving the protein of interest a tiny genetic name tag. Using CRISPR genome editing, the researchers fuse short peptide tags, such as HiBiT or ALFA, onto the natural protein inside cells or tissues. They then use engineered partner proteins—high-affinity binders that recognize these tags—directly fused to the Tn5 enzyme, which cuts DNA and attaches sequencing adapters. When the binder–Tn5 fusion attaches to the tag on the protein of interest, it snips nearby DNA and marks those sites for sequencing. Because the tags are very small and the binder–Tn5 fusion is much smaller than an antibody, this system enters cells and nuclei easily, binds with exceptional precision, and is not thrown off by chemical modifications on the target protein. 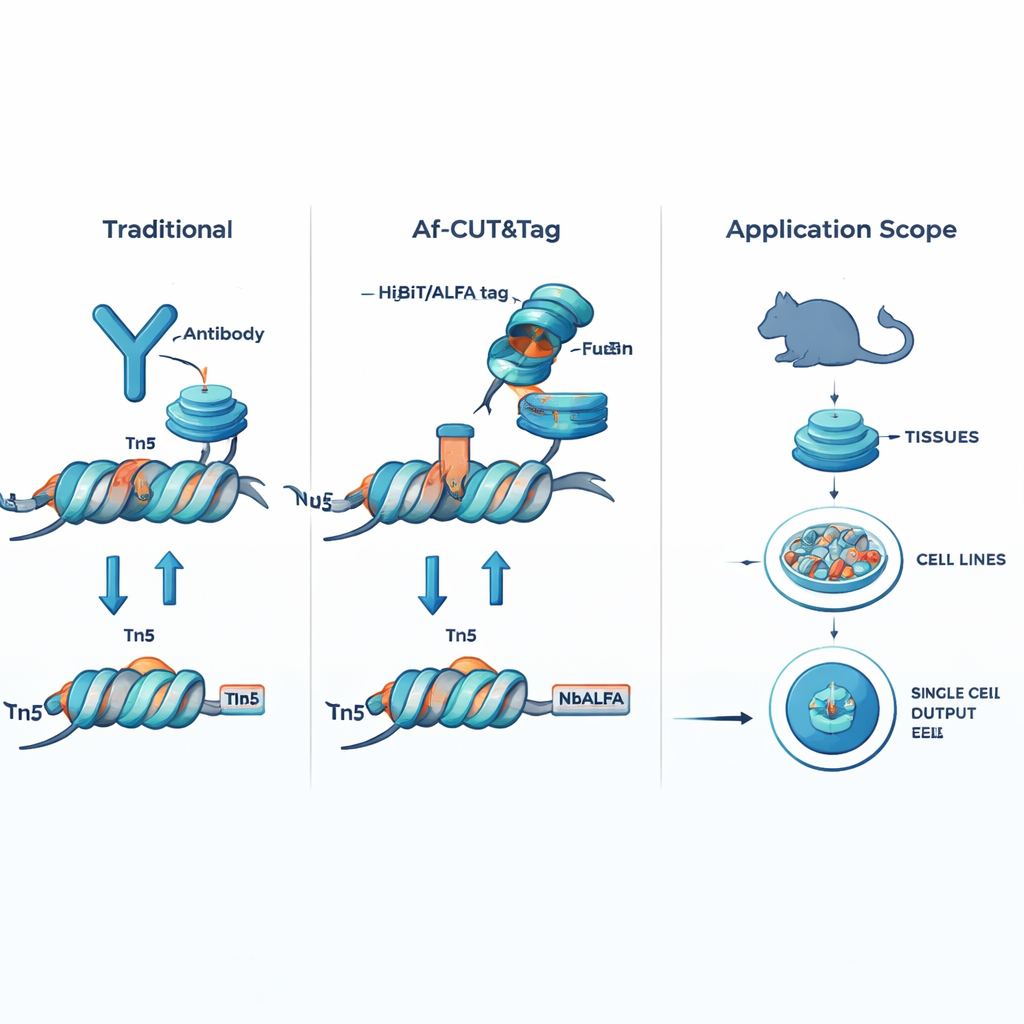
Sharper Maps from Fewer Cells
The team first showed that their binder–Tn5 fusions still grab the tags very tightly and cut DNA efficiently. They then compared Af-CUT&Tag against state-of-the-art antibody-based methods for mapping RNA polymerase II, the enzyme that reads protein-coding genes, and the CTCF protein, a major organizer of genome structure. Across human cell lines, Af-CUT&Tag produced cleaner signal at gene starting points, fewer off-target cuts in open but irrelevant DNA regions, and higher-quality sequencing libraries. Remarkably, it generated robust maps from as few as about 500 cells, and the same strategy could be adapted to many different binders targeting distinct DNA features. The method was also extended to single cells by barcoding each cell’s fragments, allowing the construction of detailed, cell-by-cell chromatin maps.
Following Liver Repair in Real Time
To demonstrate what this technology can reveal in living animals, the authors applied Af-CUT&Tag to mouse liver regeneration. They used viruses and CRISPR to add HiBiT tags to two key “Hippo pathway” regulators, YAP1 and TAZ, which help control organ size and repair. After surgically removing part of the liver, they isolated liver cell nuclei before surgery and 24 hours afterward, then used Af-CUT&Tag to see where YAP1 and TAZ were bound on the genome. The maps showed that, early after injury, these proteins reduce their presence at genes that drive fat processing and synthesis, while liver cells temporarily accumulate fat droplets. At the same time, YAP1/TAZ increase binding at genes involved in clearing heme, a potentially toxic component of blood, and at Mir122, a gene that produces a microRNA heavily enriched in liver.
miR-122 as a Guardian of Regeneration
miR-122 is a tiny RNA molecule that helps fine-tune the activity of many other genes. Using Af-CUT&Tag, the researchers saw stronger YAP1/TAZ binding near the Mir122 region, along with more open chromatin and enhanced chemical marks linked to gene activation. Experiments confirmed that miR-122 levels rise during the early regeneration window. When the team specifically removed miR-122 in mouse liver cells, animals showed more fat buildup, stronger inflammatory responses, and fewer liver cells entering the cell cycle after surgery. Together, these results suggest that YAP1/TAZ help the liver strike a careful balance during repair: briefly stockpiling fat for energy, limiting damage from heme and inflammation, and using miR-122 to restrain excessive stress while promoting tissue regrowth. 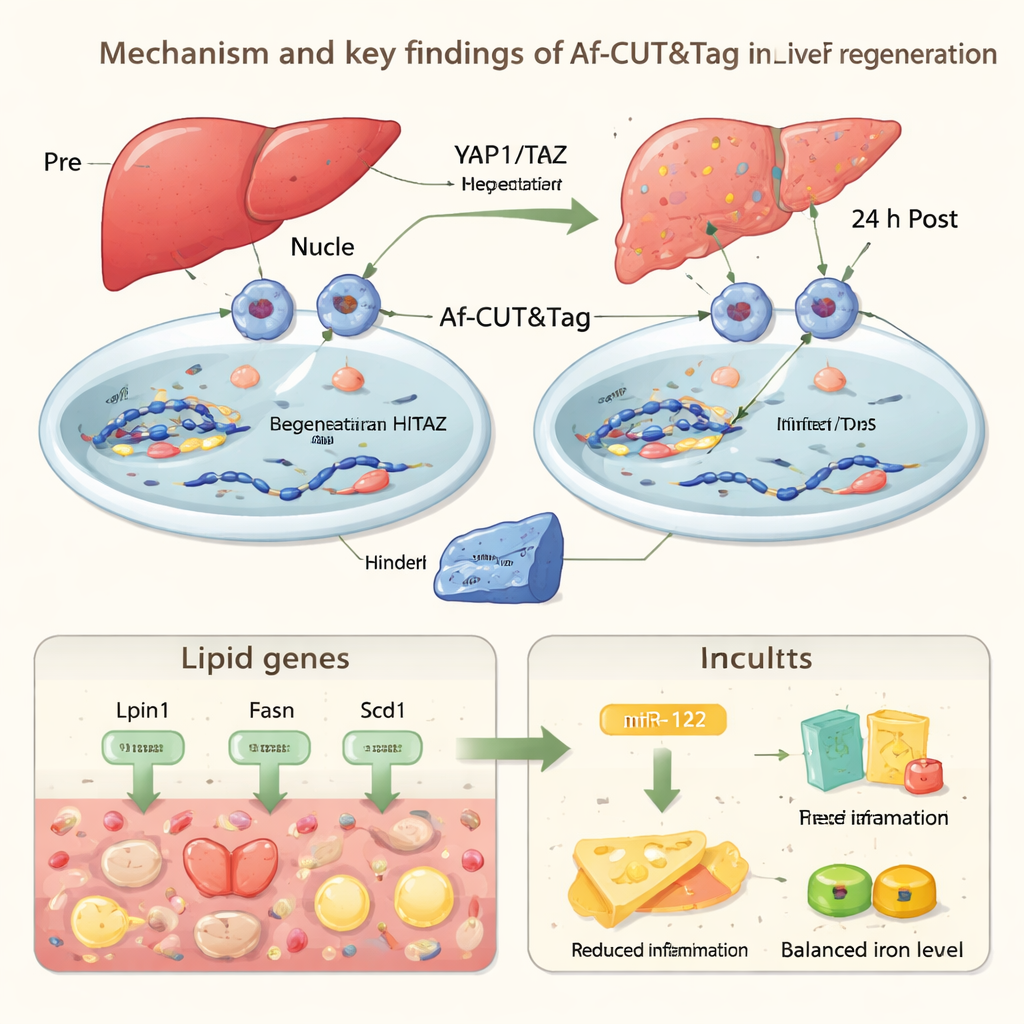
A Versatile New Lens on Gene Control
Af-CUT&Tag offers a modular, antibody-free way to see where key proteins sit on DNA in cell lines, tissues, and even individual cells. By replacing unpredictable antibodies with small genetic tags and engineered binders, it provides sharper, more reproducible maps from far fewer cells. Applied to liver regeneration, it revealed how YAP1 and TAZ coordinate metabolism, iron handling, and the microRNA miR-122 to support early repair. As genome editing and designer binders continue to improve, this strategy could become a standard tool for dissecting how gene regulation goes right in development and regeneration—and wrong in diseases such as cancer and liver failure.
Citation: Wang, X., Deng, X., Qiu, L. et al. Af-CUT&Tag: a sensitive and antibody-free chromatin profiling method using genetically encoded tags and high-affinity binders fused to Tn5. Nat Commun 17, 1746 (2026). https://doi.org/10.1038/s41467-026-68454-9
Keywords: chromatin profiling, epigenetics, liver regeneration, YAP1 TAZ, microRNA-122