Clear Sky Science · en
Improved in vivo gene knockout with high specificity using multiplexed Cas12a sgRNAs
Editing Genes More Reliably
Gene editing promises new ways to study biology, treat disease, and even control insect pests, but in real animals it often works less cleanly than headlines suggest. Many cells escape editing or are only partly altered, which can blur experimental results and limit real-world use. This paper describes a new CRISPR-based method in fruit flies that makes gene knockouts more complete and more precise, offering a blueprint for more reliable genome editing in complex organisms.
Why Standard CRISPR Often Falls Short
Traditional CRISPR tools like Cas9 cut DNA at a chosen spot using a single or a small number of guide RNAs. In living animals, this approach faces several obstacles. Some guides simply work poorly; some target sites are hard for the enzyme to reach; and the cell’s repair machinery often “fixes” the break with tiny changes that leave the gene still working. The result is a mosaic: neighboring cells in the same tissue can carry different mutations, or none at all. That patchiness makes it difficult to see what happens when a gene is truly turned off, and it poses a challenge for applications such as gene therapy or gene drives that must act efficiently in most cells.
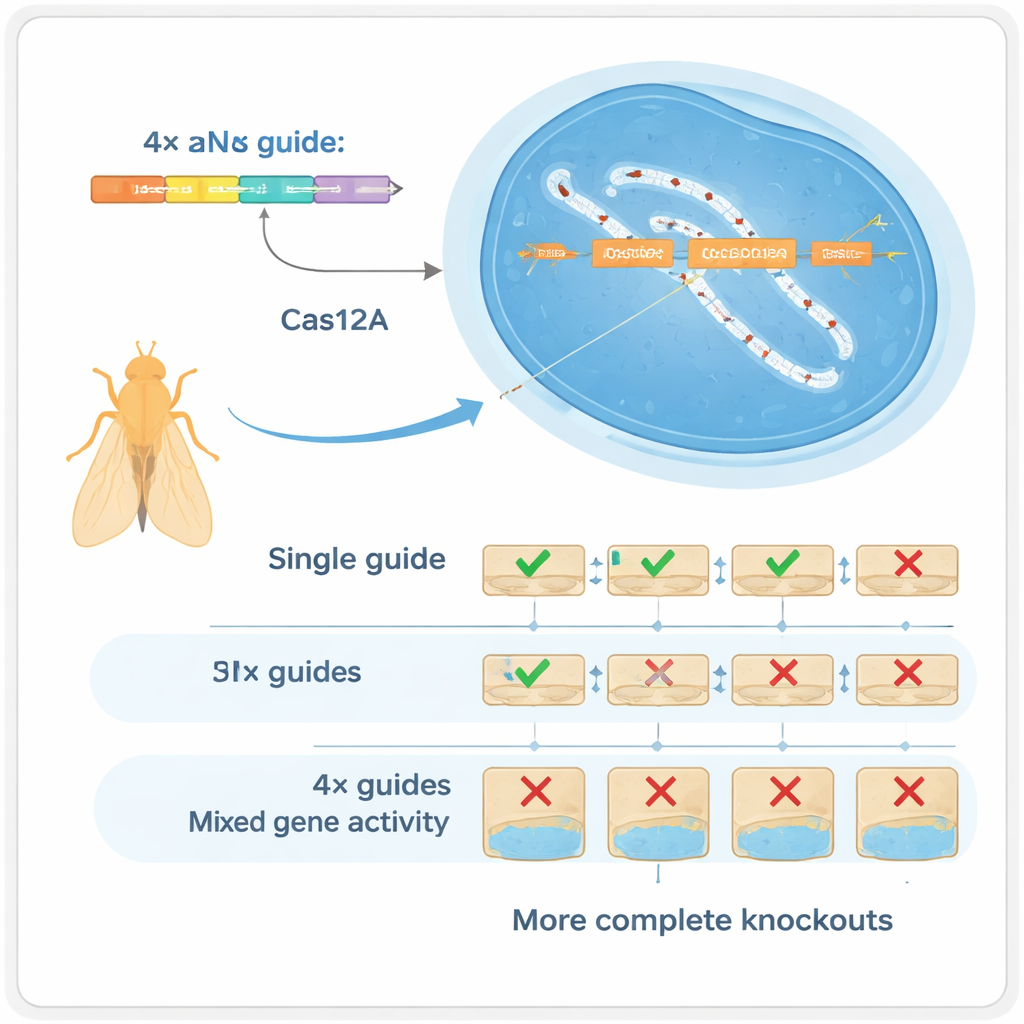
Four Guides Are Better Than One
The authors turned to another CRISPR enzyme, Cas12a, which can process compact arrays of guides much more easily than Cas9. They built a Drosophila toolkit in which each gene is targeted by an array of four guide RNAs, all encoded on a single small DNA fragment that can be mass-produced. In carefully controlled tests, they showed that using four guides per gene dramatically reshapes the kinds of DNA changes produced: instead of mostly tiny insertions or deletions at a single site, the system frequently creates larger deletions between cut sites that almost always destroy gene function. This “multiplexing” works in two ways: if one guide fails, others can still do the job (redundancy), and when several cut at once they can remove bigger chunks of the gene (synergy).
High Efficiency Without Extra Collateral Damage
Creating more DNA breaks raises obvious safety questions. Could multiple cuts in one region accidentally wipe out nearby genes? Could guides misfire elsewhere in the genome more often? To address this, the researchers measured cell death, surveyed effects on neighboring genes, and invented a clever assay to visualize chromosome repair events called loss of heterozygosity across large stretches of DNA. They found that clustering four cuts within a single gene was well tolerated: it did not increase cell death compared to conventional Cas9 approaches and rarely disturbed neighboring genes unless a guide landed extremely close to a regulatory element. Large-scale screens using more than 2,000 guides across one-third of the fly genome showed that over 99% of guide arrays were active at their intended target, while reproducible off-target activity was below 1%, indicating very high specificity even in a multiplexed setup.
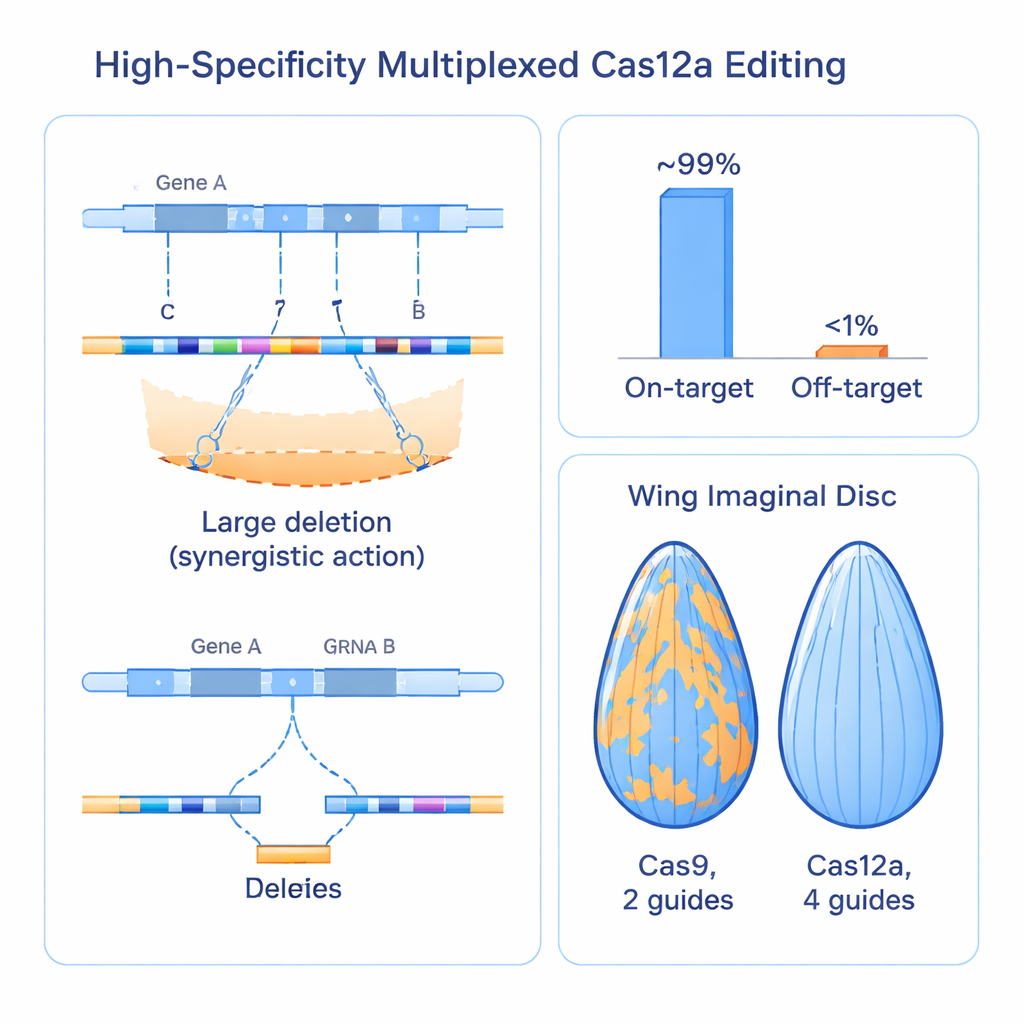
Beating Established Cas9 Systems in Real Tissues
To see whether these molecular improvements translate into clearer biology, the team compared their Cas12a–four-guide system directly against widely used Cas9-based resources targeting more than 100 genes in the fly. In tissues such as the eye, intestine, and developing wing, the Cas12a approach produced stronger and more uniform loss-of-function effects than Cas9, which often left obvious patches of unedited, normal tissue. When they used wing size as a quantitative readout, the new system consistently generated larger and more reproducible growth defects for known regulators, revealing that some genes previously scored as weak or nonessential were in fact missed because older tools did not inactivate them thoroughly enough. The heightened power of the method even uncovered a previously unknown, essential role for a gene called trade embargo in wing development and survival.
What This Means for Future Gene Editing
In simple terms, this work shows how to turn CRISPR from a sometimes-messy scalpel into a more decisive switch for turning genes off in living animals. By combining Cas12a with four guides per gene, the authors achieve near-complete knockouts with very low unintended effects, all in a format that is practical to scale across hundreds of genes. Although developed in fruit flies, the underlying principles—using multiple guides for redundancy and synergy, and carefully checking for chromosomal side effects—are broadly applicable. This strategy could improve basic research, sharpen genetic screens, and inform safer designs for future medical and ecological applications of gene editing.
Citation: Port, F., Buhmann, M.A., Zhou, J. et al. Improved in vivo gene knockout with high specificity using multiplexed Cas12a sgRNAs. Nat Commun 17, 877 (2026). https://doi.org/10.1038/s41467-026-68434-z
Keywords: CRISPR, Cas12a, gene knockout, Drosophila, genome editing specificity