Clear Sky Science · en
ClpC1-targeting peptide natural products differentially dysregulate the proteome of Mycobacterium tuberculosis
Why breaking bacterial cleanup crews matters
Tuberculosis remains one of the world’s deadliest infectious diseases, and drug-resistant strains of Mycobacterium tuberculosis are making it harder to treat. Most existing antibiotics hit the bacterium’s cell wall or its ability to make energy and copy genetic material. This study explores a very different weak spot: the cell’s internal “cleanup crew” that maintains protein health. By understanding how new natural-product–based molecules disrupt this cleanup system, scientists hope to design future drugs that can kill TB bacteria in new and more durable ways.
The cell’s protein housekeeping hub
Like all living cells, the TB bacterium constantly builds, folds, repairs, and recycles its proteins. A central player in this protein housekeeping network is a large machine called the ClpC1–ClpP1P2 complex. One part (ClpC1) acts as a chaperone, recognizing and pulling in damaged or unneeded proteins, while the other part (ClpP1P2) chops them into pieces for recycling. This system is essential for the bacterium’s survival, especially under stress such as heat or antibiotic attack. Several natural peptide compounds—ecumicin, ilamycins (rufomycins), and cyclomarins—are already known to bind to ClpC1 and show strong TB-killing activity, suggesting this complex could be a powerful drug target. 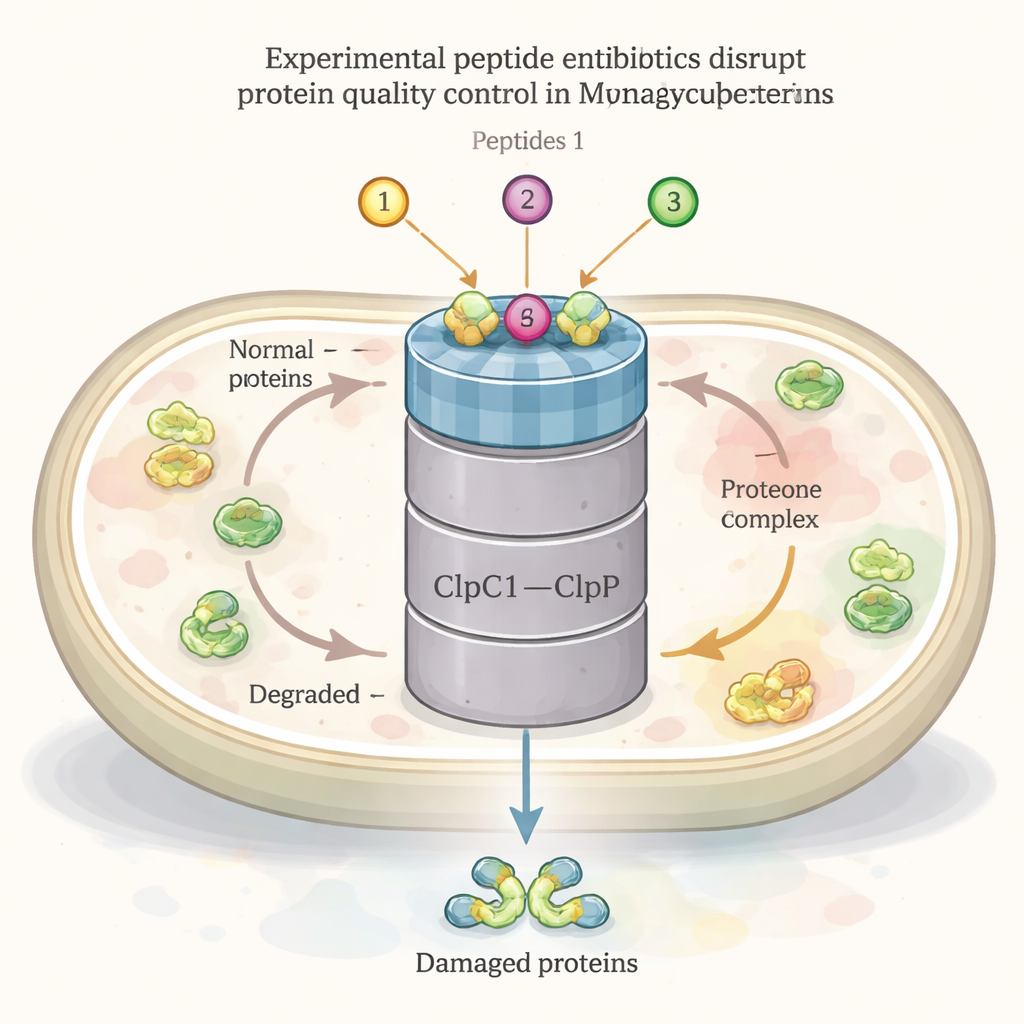
Three related drugs, three different impacts
The researchers examined one representative from each of the three peptide families: an ecumicin analogue (Ecu*), ilamycin E (IlaE), and a cyclomarin derivative (desoxycyclomarin, dCym). Although all three latch onto the same region of ClpC1, they do not behave the same way inside the cell. Using quantitative proteomics—an approach that measures thousands of proteins at once—they found that Ecu* caused the broadest disruption, significantly altering levels of nearly 17% of all bacterial proteins. IlaE changed about 12%, and dCym about 7%. Only 72 proteins were consistently affected by all three compounds, and most of those decreased in abundance. This shows that each molecule pushes the ClpC1 machine in a different direction, reshaping the cell’s protein landscape in distinct ways rather than simply “turning the machine off.”
Clogging the shredder versus cutting the power
To put these effects in context, the team compared the peptides with bortezomib, a cancer drug that broadly blocks protein-degrading enzymes. In TB bacteria, bortezomib caused general buildup of many proteins, as expected when the shredders are switched off. By contrast, the peptides produced more selective patterns: some known ClpC1 targets accumulated, others were depleted, and global inhibition of all protein breakdown was not observed. Detailed lab tests with purified components showed that each peptide altered how specific substrates were handled. For example, all three interfered with destruction of one regulatory protein (PanD), but only Ecu* blocked breakdown of a model disordered protein, and only dCym impaired degradation of the stress-protective protein Hsp20. These differences hint that the overall shape and flexibility of each target protein influence how it is affected when ClpC1 is “mis-tuned” by a given compound. 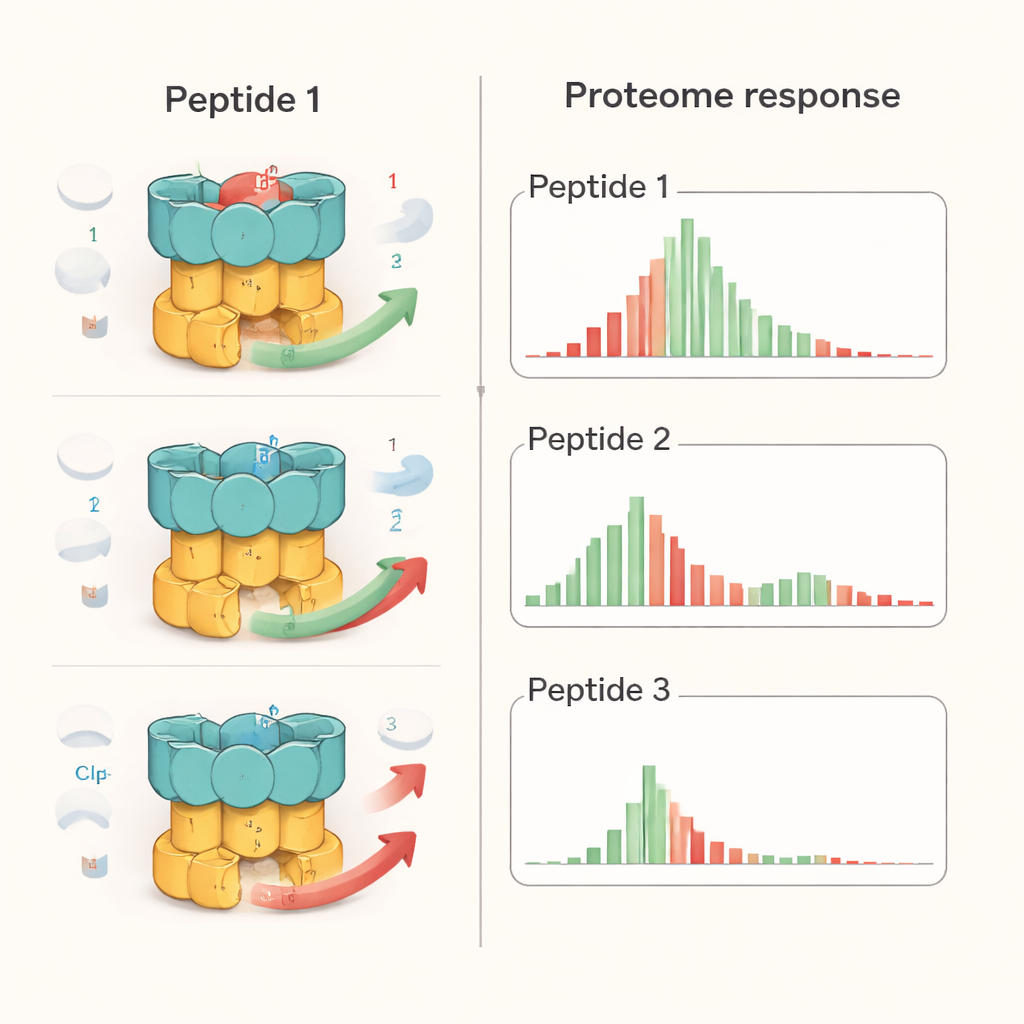
Stress responders and bacterial self-rescue
The TB bacterium has backup systems to cope when protein damage overwhelms the main cleanup machinery. One such system involves another chaperone, ClpC2, which can bind cyclomarin-like molecules and reduce their impact on ClpC1. This study found that although all three peptides can bind ClpC2 in the test tube, only the cyclomarin derivative (dCym) actually triggered higher ClpC2 levels in live bacteria and became more potent when ClpC2 was experimentally reduced. In contrast, Ecu* and IlaE remained effective even when ClpC2 was knocked down, suggesting they largely bypass this self-rescue route. The team also discovered a strong, compound-specific link to Hsp20, a small protein that helps prevent harmful clumps: Ecu* (and to a lesser extent IlaE) caused dramatic increases in Hsp20 levels and Ecu* directly bound Hsp20, revealing a new connection between these drug candidates and the cell’s stress-response network.
Turning stress into a therapeutic advantage
Because the ClpC1 system is especially important under harsh conditions, the researchers tested the drugs while the bacteria were subjected to mild heat stress, which increases the load of misfolded proteins. Under these conditions, both Ecu* and dCym became much more lethal, reaching toxicity similar to that of bortezomib, even though they act more selectively. This demonstrates that carefully targeting the protein quality-control machinery can be as effective as shutting down all major proteases, but with potentially more precise control. The work also showed that many of the changes caused by Ecu* in protein levels did not simply mirror changes in gene activity, underscoring that disrupting ClpC1 mainly perturbs regulation after proteins are made.
What this means for future TB treatments
For non-specialists, the key message is that TB bacteria rely on a finely balanced internal cleanup and stress-response system to survive. The three natural-product–derived peptides tested here all grab the same central machine but push it into different modes, leading to diverse and highly specific distortions in the bacterium’s protein makeup. Importantly, ecumicin-like and ilamycin-like compounds avoid a built-in bacterial rescue mechanism that can blunt cyclomarin’s effect, yet still cripple the pathogen under stress. These properties make them promising scaffolds for future tuberculosis drugs, including designer molecules that could force TB cells to selectively destroy their own vital proteins or that could be built into “protein-targeting chimeras” to precisely direct bacterial self-digestion.
Citation: Barter, I.K., Bedding, M.J., Leodolter, J. et al. ClpC1-targeting peptide natural products differentially dysregulate the proteome of Mycobacterium tuberculosis. Nat Commun 17, 1725 (2026). https://doi.org/10.1038/s41467-026-68423-2
Keywords: tuberculosis, protein quality control, ClpC1, antibiotic resistance, natural product antibiotics