Clear Sky Science · en
Simultaneous epigenomic profiling and regulatory activity measurement using e2MPRA
Reading the Cell’s Hidden Switches
Every cell in your body carries essentially the same DNA, yet a brain cell and a liver cell behave very differently. The secret lies in short stretches of DNA that act like dimmer switches for genes, turning them up, down, or off. This study introduces a powerful new tool, called e2MPRA, that lets scientists test thousands of these switches at once while also watching how the DNA is packaged and marked inside the cell—key steps toward understanding development, disease risk, and why some genetic variants matter while others do not.
From DNA Maps to DNA Function
Over the past decade, researchers have drawn huge maps of "candidate" regulatory switches, known as cis-regulatory elements. They use methods that show where DNA is loosely packed (open chromatin), which proteins are bound, and which histone marks decorate the surrounding DNA. These maps are impressive, but they are mostly descriptive: just because a protein binds or a mark appears does not prove that a sequence truly controls gene activity. Traditional massively parallel reporter assays (MPRAs) can test whether thousands of DNA snippets boost or silence a reporter gene, but they do not tell us which proteins bind there or what epigenetic changes accompany that activity.
A Two-in-One Assay for Activity and Epigenetic Marks
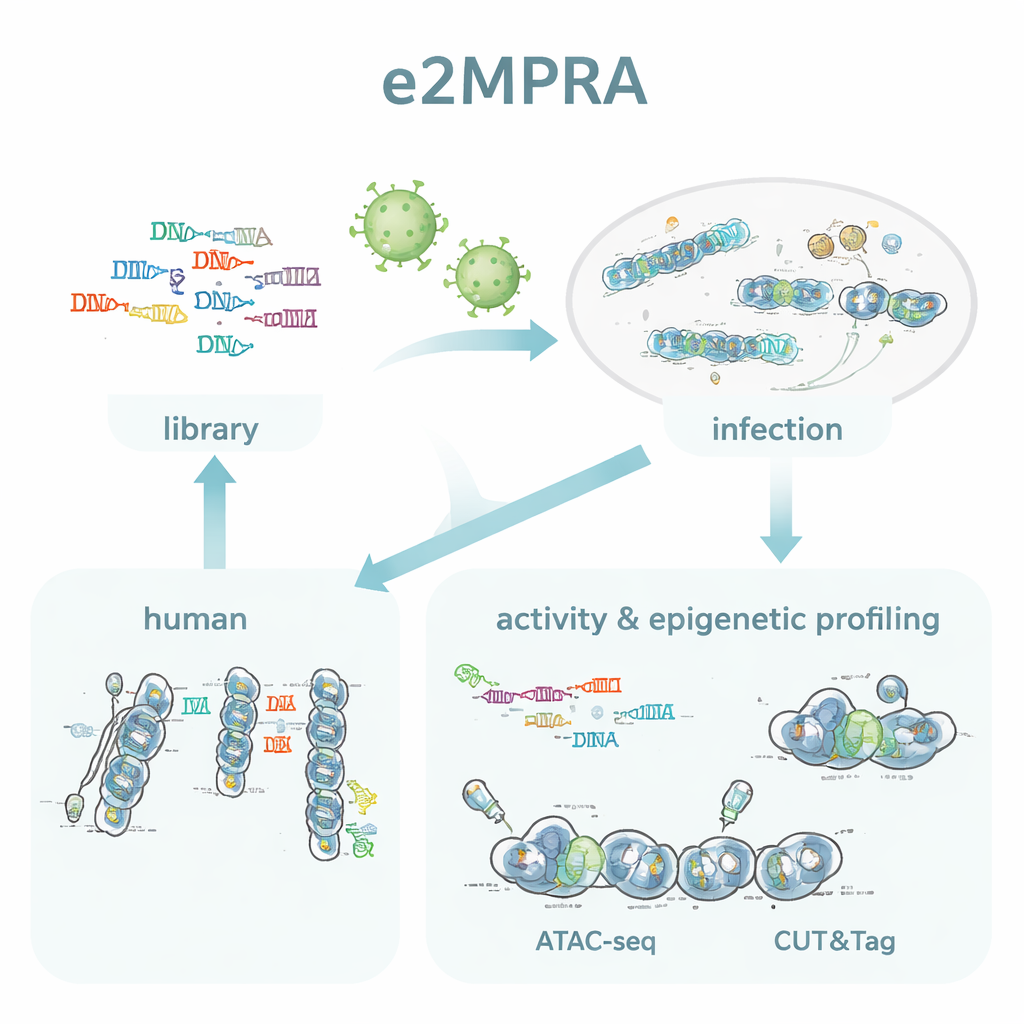
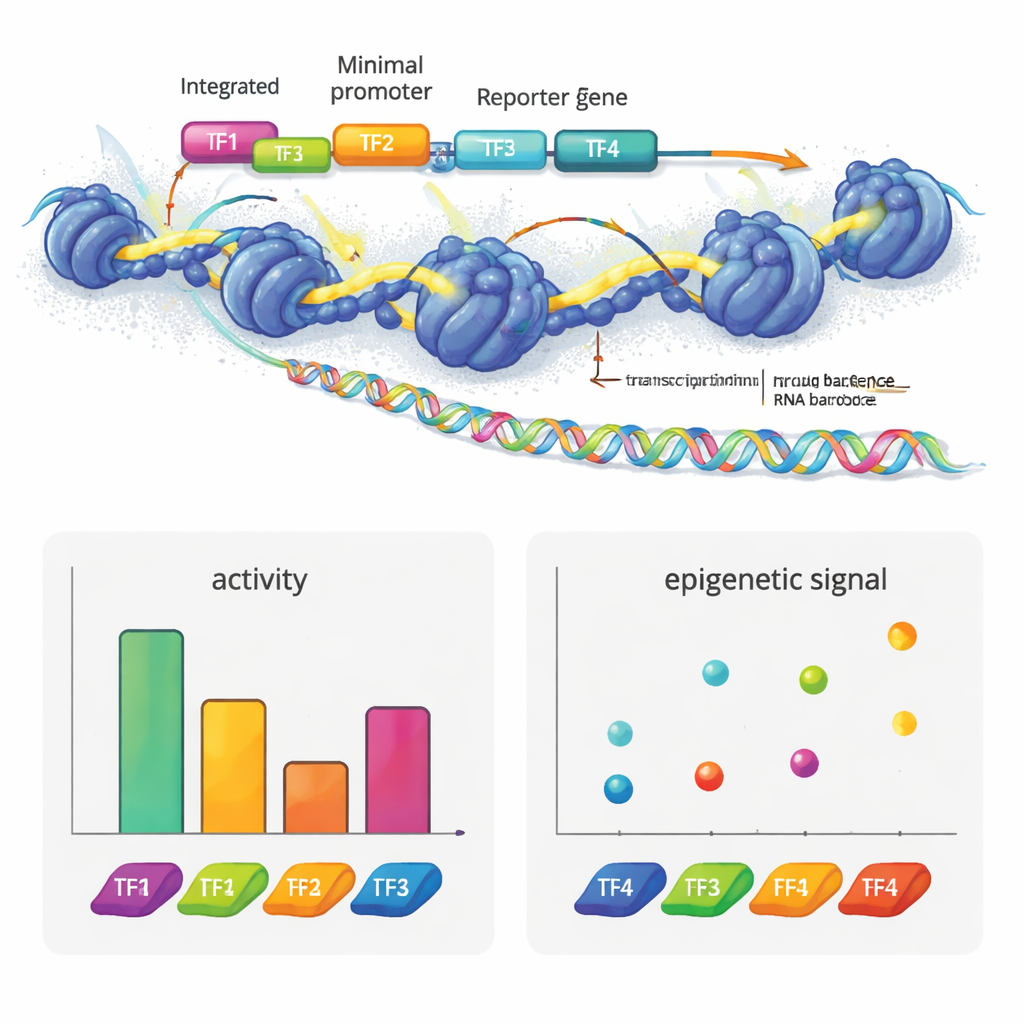
The authors created e2MPRA by blending a lentiviral MPRA with two epigenomic techniques, ATAC-seq and CUT&Tag. First, they build a DNA library in which each candidate regulatory sequence is linked to a short barcode and placed next to a minimal promoter and reporter gene. This library is then packaged into lentiviruses and integrated many times into the genomes of cultured human cells. From the same pool of cells, they read barcodes in RNA to measure regulatory activity, and they use ATAC-seq and CUT&Tag to measure chromatin openness and a key histone mark (H3K27ac) specifically at the inserted sequences. By normalizing these signals to how often each sequence actually integrates, they obtain side-by-side measurements of "on/off strength" and epigenetic state for thousands of elements in a single experiment.
Decoding How Transcription Factor Motifs Work Together
To see what this technology can reveal, the team built synthetic enhancers from known transcription factor motifs used in liver cells. They arranged these motifs in different numbers and orders on neutral DNA templates. Some factors, such as HNF1A and XBP1, behaved as classic activators: adding more copies increased reporter output. Others, like HNF1A and ONECUT1, mainly boosted chromatin accessibility, fitting their role as "pioneer" factors that open closed DNA. PPARA showed a different pattern, strongly altering chromatin and histone marks but not transcription on its own, yet cooperating with other factors to raise gene activity. REST, a known repressor, dampened activity when placed next to activators. Strikingly, simply reordering four motifs could substantially change enhancer strength, and activators tended to work best when closer to the promoter, revealing a sort of grammatical structure in how these motifs are arranged.
Pinpointing Sensitive Bases in Pluripotency Enhancers
The researchers then turned to enhancers important for stem cell identity, focusing on regions bound by the pluripotency factors POU5F1 (also called OCT4) and SOX2. They created dense mutation libraries in which every base of 100-base-pair enhancers was systematically altered, along with small six-base windows that were randomly scrambled. Using e2MPRA in induced pluripotent stem cells, they could see which mutations weakened or strengthened reporter activity and how they affected chromatin openness and acetylation. Disrupting the POU5F1::SOX2 binding motif often reduced both gene activity and epigenetic marks, confirming its central role. In a well-studied enhancer near the POU5F1 gene, they also found that changing a YY1 motif increased transcription but reduced chromatin opening, hinting that some factors can promote an open state while still restraining expression.
What This Means for Genes and Disease
e2MPRA does not perfectly reproduce every nuance of the genome’s natural 3D wiring, but it offers a practical way to compare thousands of regulatory sequences and variants under the same conditions. By coupling a functional readout (how strongly a DNA segment drives a gene) with epigenetic readouts (how that DNA is packaged and marked), this method helps reveal why certain transcription factor motifs, combinations, and positions are so crucial. In the long run, tools like e2MPRA can make it easier to interpret noncoding genetic variants linked to disease, design synthetic enhancers for gene therapy, and build a more complete "regulatory code" that connects DNA sequence to cell behavior.
Citation: Zhang, Z., Georgakopoulos-Soares, I., Bourque, G. et al. Simultaneous epigenomic profiling and regulatory activity measurement using e2MPRA. Nat Commun 17, 1724 (2026). https://doi.org/10.1038/s41467-026-68422-3
Keywords: gene regulation, enhancers, epigenetics, transcription factors, functional genomics