Clear Sky Science · en
DNA end configurations dictate synaptic complex formation during NHEJ-mediated end bridging
When DNA Breaks, Cells Must Think Fast
Every day, the DNA in our cells is nicked, cut, and battered by normal metabolism and environmental stresses. Among the most dangerous injuries are double-strand breaks, where both strands of the DNA helix snap. If these breaks are not repaired quickly and accurately, cells can die or, worse, drift toward cancer. This study explores how the exact shape and chemistry of broken DNA ends help determine whether a key repair pathway, called non-homologous end joining (NHEJ), pulls those ends together efficiently and with as few mistakes as possible.
Two Ways to Bring Broken DNA Together
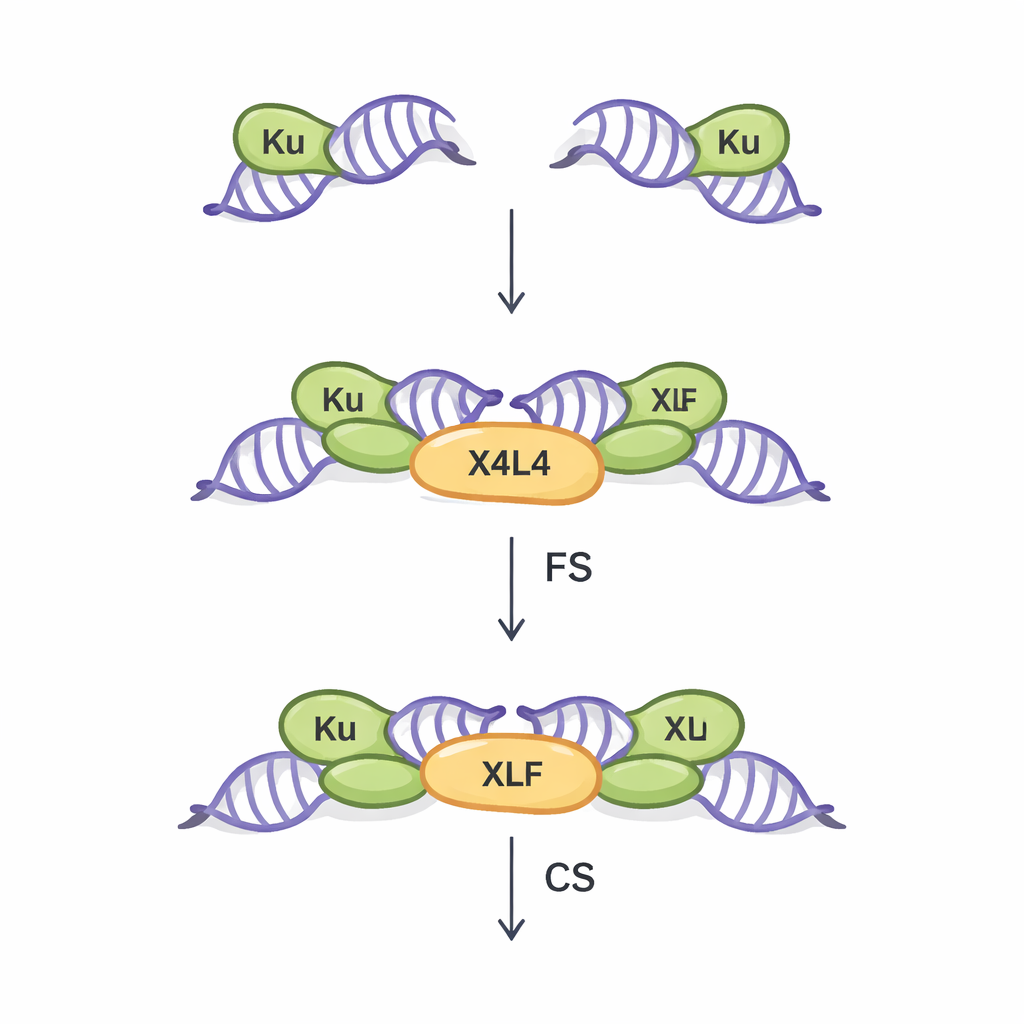
NHEJ is the main repair route for double-strand breaks in most of our body’s cells. It works without needing a matching DNA template, making it fast but somewhat error-prone. The first crucial step in this pathway is called synapsis: physically bridging the two broken DNA ends so they can later be joined. Earlier work, mostly using idealized blunt DNA ends, revealed two kinds of synaptic assemblies. In the “flexible synaptic” (FS) state, the ends are held near each other and can move around, probing for ways to pair. In the “close synaptic” (CS) state, the ends line up end-to-end and are ready for sealing by a ligating enzyme. Three core proteins—Ku, XRCC4–Ligase IV (together called X4L4), and XLF—have been known to coordinate this process, but how real-world, irregular DNA ends influence which synaptic state forms has remained poorly understood.
End Shape and Short Matches Steer the Repair Route
The researchers used single-molecule FRET, a fluorescence-based technique that tracks distances at the nanometer scale, to watch individual DNA molecules as they were brought together by NHEJ proteins. They compared simple blunt ends with more realistic ends carrying short overhangs—little single-stranded tails that can potentially pair with each other. They found that when overhangs are complementary, even by just a few bases, they dramatically increase the chance of forming a close synaptic complex. In fact, three matching bases of “microhomology” at the break were enough for Ku and X4L4 alone to create a stable CS complex, without needing XLF. This shows that the DNA itself can supply part of the energy and guidance normally provided by helper proteins.
How Tiny DNA Flaps Help Stabilize the Bridge
By analyzing distinct FRET signatures, the team uncovered at least two structural flavors of the close synaptic state for overhanging ends. In one, the very tips of the overhangs pair with each other, making a nicked or gapped joint. In the other, the junctions where double-stranded DNA meets single-stranded overhangs press together in-line, while the overhangs flip outward like small flaps. These flaps still allow base pairing and keep the ends in very close proximity, which likely stabilizes the complex and gives enzymes room to trim or extend the DNA as needed before final sealing. Intriguingly, common oxidative damage in the overhangs, such as 8-oxoguanine, only modestly affected this process, highlighting how robust NHEJ is to certain forms of damage.
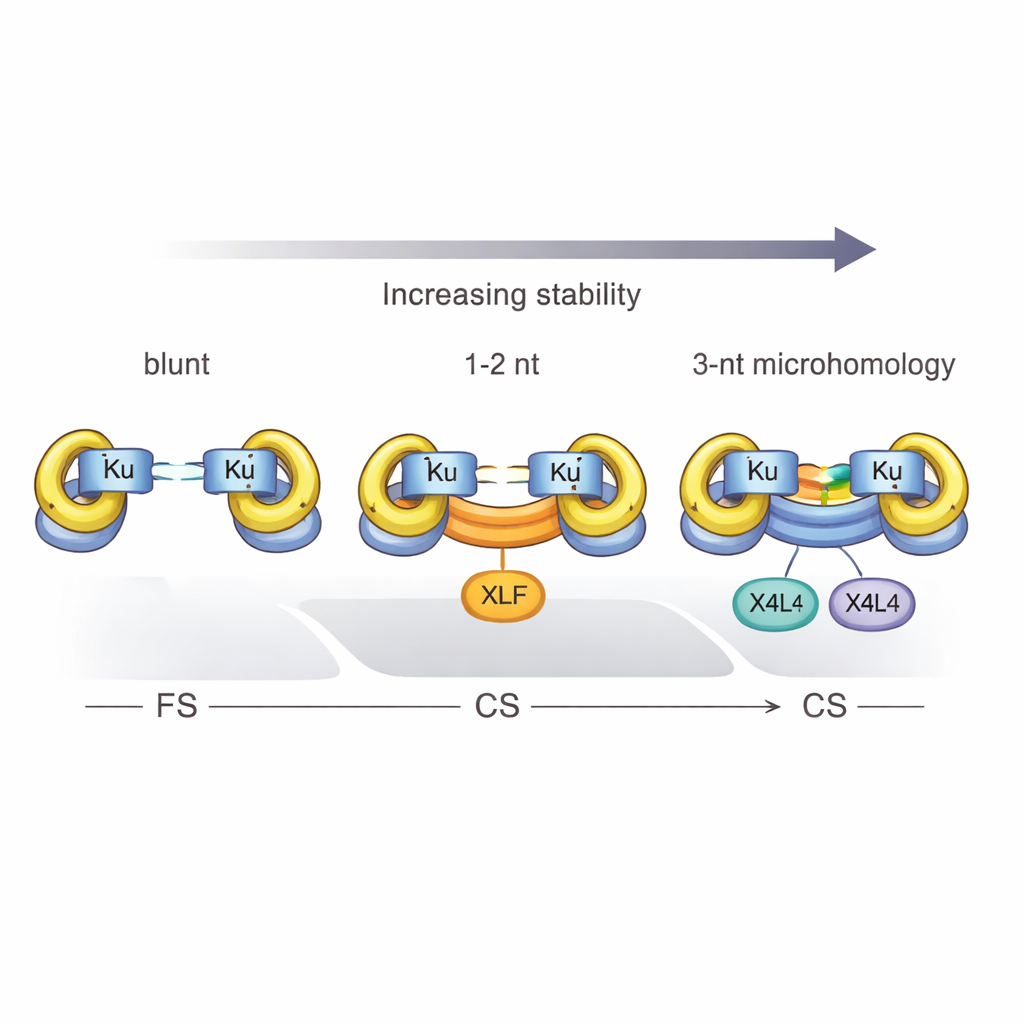
Proteins, Phosphates, and the Power of Three
The study also defined a hard threshold: at least three complementary base pairs are required for Ku and X4L4 to reliably convert a flexible complex into a close one. With only one or two matches, the system mostly stalls in the flexible state unless XLF joins in to help clamp the ends together. Chemical details also matter. When the broken DNA carries a 5′ phosphate group—a normal feature of many physiological breaks—synapsis becomes more efficient, and close complexes are more likely to be truly ligated, turning a temporary bridge into a permanent repair. However, this phosphate boost cannot fully replace the stabilizing effect of base pairing at the break.
Why This Matters for Health and Genome Editing
In everyday language, this work shows that broken DNA ends are not passive victims waiting to be fixed. Their exact shapes, tiny patches of matching sequence, and chemical tags actively guide how repair proteins grab, align, and reconnect them. When three or more matching bases are present, the DNA ends themselves help lock the break into a repair-ready configuration, sometimes making certain helper proteins optional. When such matches are scarce, extra protein factors become essential. These insights help explain why some breaks are mended neatly while others lead to small mutations or rearrangements. They also offer clues for improving genome-editing tools like CRISPR, where deliberately created DNA breaks are repaired by the same NHEJ machinery. By tuning the ends of those breaks, scientists may better control how the genome is reshaped.
Citation: Zhang, C., Jin, W., Jiang, Y. et al. DNA end configurations dictate synaptic complex formation during NHEJ-mediated end bridging. Nat Commun 17, 1720 (2026). https://doi.org/10.1038/s41467-026-68417-0
Keywords: DNA double-strand break repair, non-homologous end joining, DNA synapsis, microhomology, genome stability