Clear Sky Science · en
Sequence-based generative AI design of versatile tryptophan synthases
Teaching Enzymes New Tricks with AI
Modern society runs on molecules—medicines, materials, and specialty chemicals—that are often made with energy‑hungry, polluting processes. Nature’s catalysts, enzymes, can do similar jobs cleanly and efficiently, but finding or building the right enzyme for a new industrial task is slow and uncertain. This study shows that generative artificial intelligence, the same class of technology behind text‑writing chatbots, can be used to design brand‑new enzymes that not only work well in the lab but sometimes outperform the best that evolution and years of engineering have already produced.
Why Enzymes Matter for Everyday Life
Enzymes are tiny protein machines that speed up chemical reactions in living cells. Chemists have learned to repurpose them to make drugs, food ingredients, and other valuable products using less energy and fewer toxic reagents than traditional chemistry. The catch is that each new application usually demands an enzyme with just‑right performance—able to accept particular starting materials, survive processing conditions, and produce high yields. Conventional “directed evolution” improves enzymes by making and testing many mutated versions, generation after generation. This works well but depends on a decent starting enzyme and can take months or years of trial and error, leaving many useful reactions unexplored.
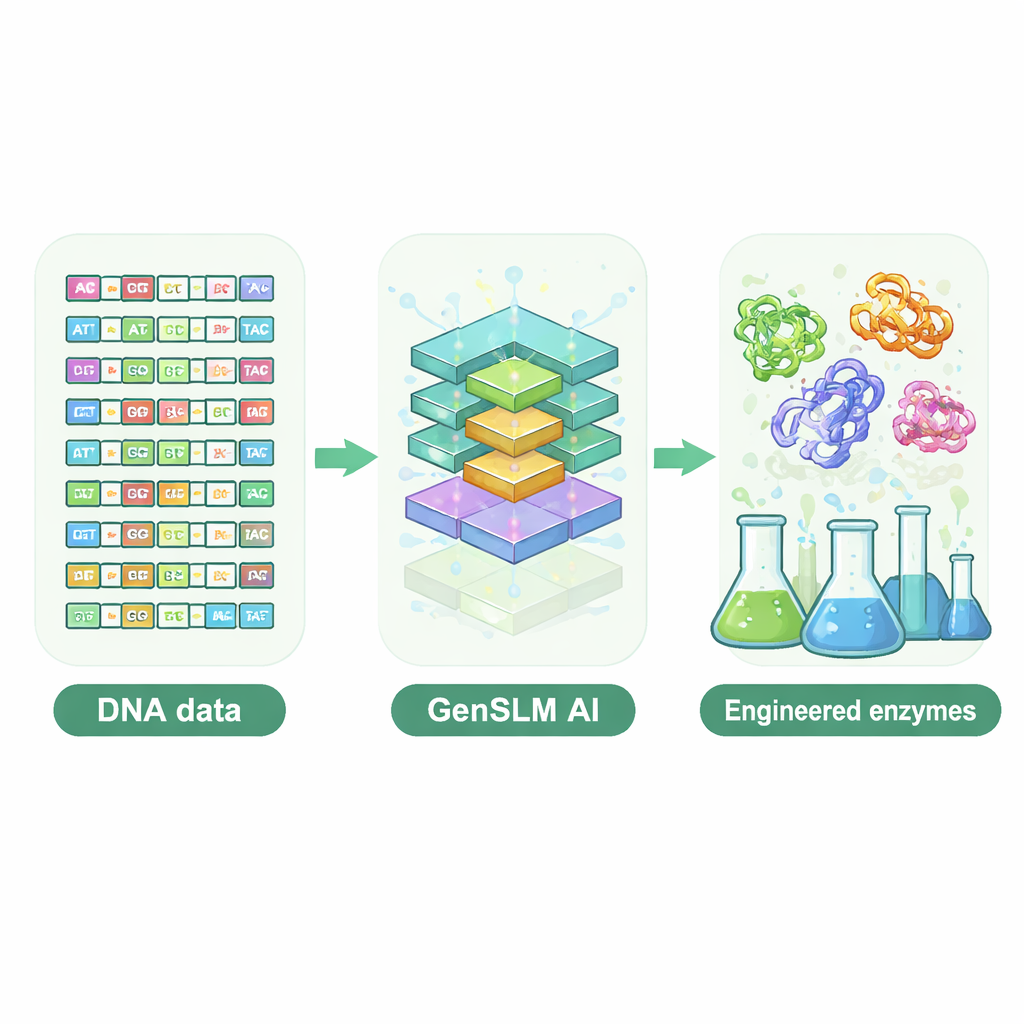
Letting a Language Model Write DNA
The researchers turned to a genome‑scale language model called GenSLM, which learns patterns in DNA the way a language model learns grammar and style in text. Instead of working with finished protein sequences, GenSLM reads and writes DNA in three‑letter codons, mirroring how cells translate genes into proteins. The team first fine‑tuned GenSLM on tens of thousands of natural genes for one particularly complex enzyme subunit, called TrpB, which helps build the amino acid tryptophan. They then asked the model to generate thousands of entirely new trpB genes. Simple computational filters weeded out sequences that were too short or too long, unlikely to fold properly, or nearly identical to known natural enzymes, leaving 105 diverse candidates for experimental testing in bacteria.
From Computer Designs to Working Catalysts
When these 105 AI‑designed TrpB enzymes were produced in E. coli, many folded well and were made in high amounts. Dozens could carry out their main job: converting indole and the natural partner amino acid, serine, into tryptophan. Some worked robustly even at elevated temperatures, despite no explicit design for heat resistance. In side‑by‑side tests, a subset of GenSLM‑TrpBs matched or beat a benchmark enzyme that had been painstakingly evolved in the lab for years to function on its own at 75 °C. One standout design, labelled 230, produced more tryptophan than this industrially used benchmark both at room temperature and at high temperature, showing that a model trained only on sequence data can jump directly to top‑tier performance.
New Flexibility Beyond What Nature Built
The team then challenged the enzymes with a panel of non‑natural substrates—indole derivatives, a different alcohol‑like partner, and a fluorinated compound used in drug manufacture. Natural versions of TrpB are usually picky: they strongly favor their native substrates and show little activity on such alternatives. Remarkably, the AI‑generated enzymes were often more adventurous. For every non‑natural substrate tested, at least one GenSLM design showed measurable activity, and many performed better than natural enzymes. Again, variant 230 stood out, converting all seven alternative substrates with yields ranging from modest to nearly complete, a breadth of “promiscuity” not previously seen in this enzyme family. Yet when researchers compared 230 to its closest natural relative—differing at only 78 out of 400 amino acid positions—they found that the natural enzyme lacked this versatility even though its overall structure and key active‑site residues were nearly identical.
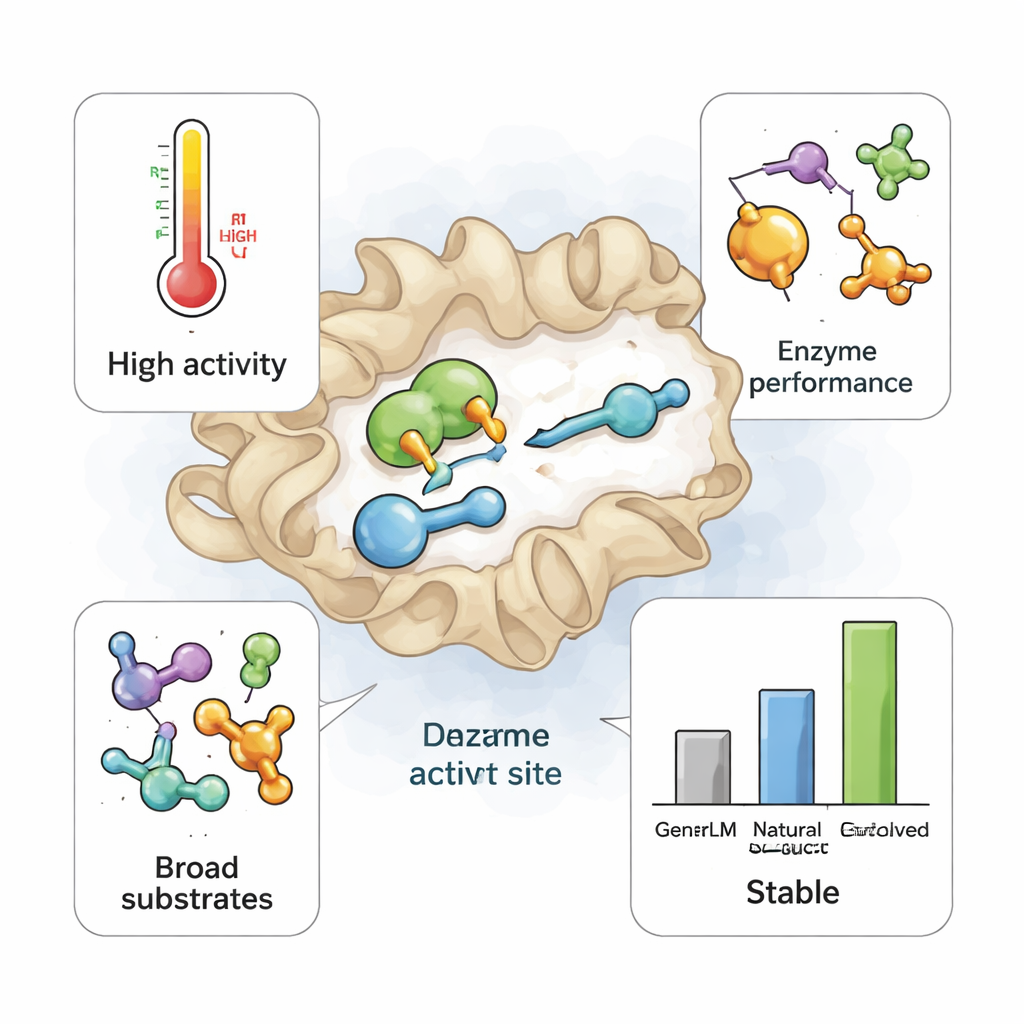
What This Means for Future Green Chemistry
To a non‑specialist, the key message is that an AI model trained only on existing DNA sequences can imagine realistic new enzymes that nature never tried, some of which are better tools for chemistry than the ones we currently use. These AI‑designed TrpB variants keep the essential shape and function of their natural cousins but gain an unusual ability to handle many different starting materials. That flexibility could dramatically reduce the amount of lab work needed to discover enzyme‑based routes to new medicines and other products. As design, DNA synthesis, and testing become faster and cheaper, similar generative models may turn enzyme discovery from a slow treasure hunt into a rapid, routine design task, helping shift more industrial chemistry toward cleaner, enzyme‑powered processes.
Citation: Lambert, T., Tavakoli, A., Dharuman, G. et al. Sequence-based generative AI design of versatile tryptophan synthases. Nat Commun 17, 1680 (2026). https://doi.org/10.1038/s41467-026-68384-6
Keywords: enzyme engineering, generative AI, protein design, tryptophan synthase, biocatalysis