Clear Sky Science · en
Cryo-EM structure of the human COP1-DET1 ubiquitin ligase complex
How Cells Decide Which Proteins to Destroy
Inside every cell, thousands of proteins must be constantly built, remodeled, or destroyed at the right time. This paper looks at one of the cell’s key “quality-control” machines, a molecular shredder that decides when to remove powerful gene regulators linked to cancer and development. By revealing its three‑dimensional structure in near‑atomic detail, the authors show how this machine can switch between an inactive, stacked state and an active, substrate-processing form—insight that may eventually help design drugs to tune it up or shut it down.
A Molecular Recycling Crew With Many Parts
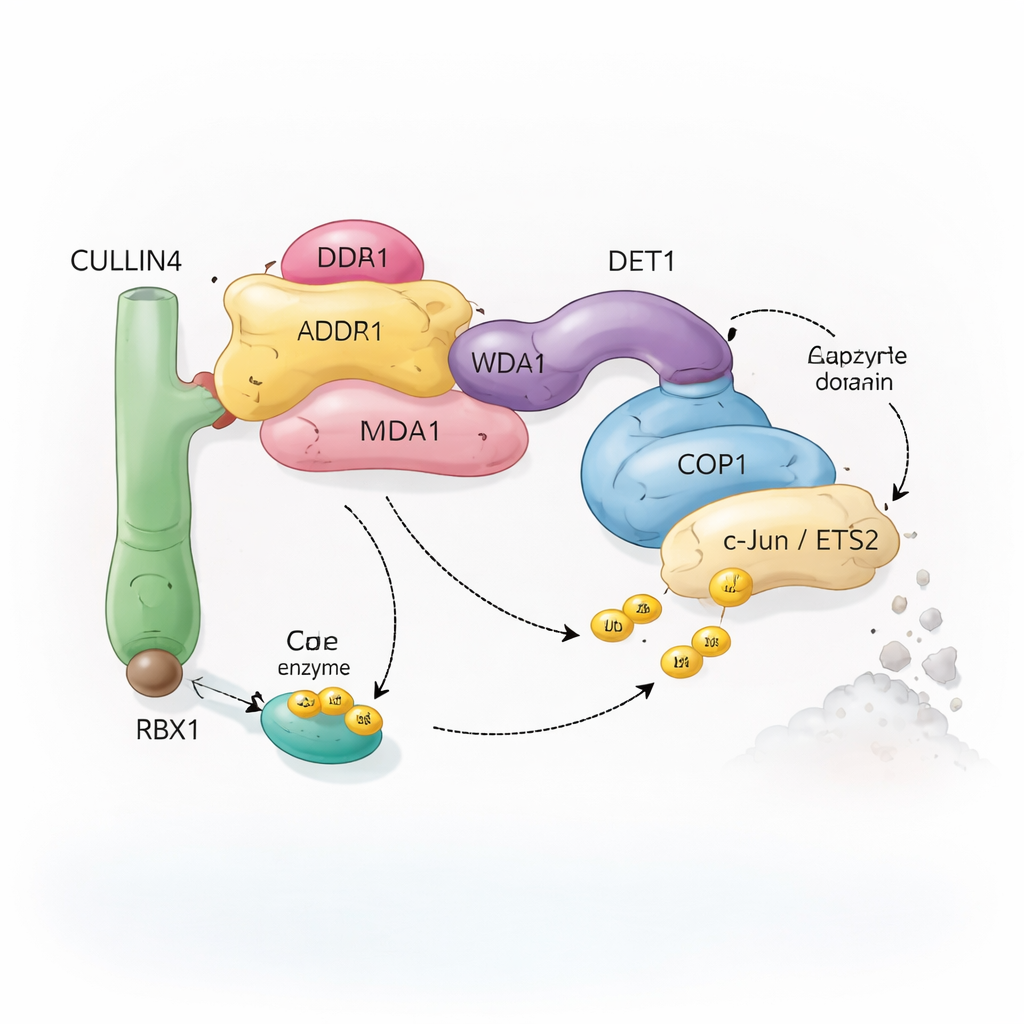
Cells rely on a system called ubiquitination to mark proteins for recycling. Small molecules of ubiquitin act like removable flags that tell the cell whether a protein should be altered, relocated, or destroyed. Special enzyme complexes, known as ubiquitin ligases, attach these flags to selected targets. The complex studied here centers on COP1, a protein found from plants to humans, which helps tag important transcription factors—proteins that control which genes are turned on or off. COP1 does not work alone: it can plug into a larger assembly built around a scaffold protein called CULLIN4, plus several adaptors named DDB1, DDA1, and DET1. Together, these components form a flexible “recycling crew” that recognizes specific transcription factors such as c‑Jun and ETS2, many of which are involved in cell growth and cancer.
Capturing the Complex in 3D
To understand how this crew operates, the researchers used single‑particle cryo‑electron microscopy, a technique that images flash‑frozen proteins and reconstructs their shapes in high resolution. They re‑created the human COP1–DET1 system in cultured cells, purified the complexes, and visualized several distinct structural states. First, they solved the structure of the DDB1–DDA1–DET1 module, which acts as a central hub. DET1 adopts an unusual, partly flexible shape rather than a rigid disk, with a protruding “claw” that can grip partner enzymes. Then, by adding COP1 and a class of helper enzymes known as E2s, they captured larger assemblies that show how all the pieces fit together, revealing both compact stacks and more open dimeric (two‑COP1) arrangements.
An Inactive Stack and an Active Dimer
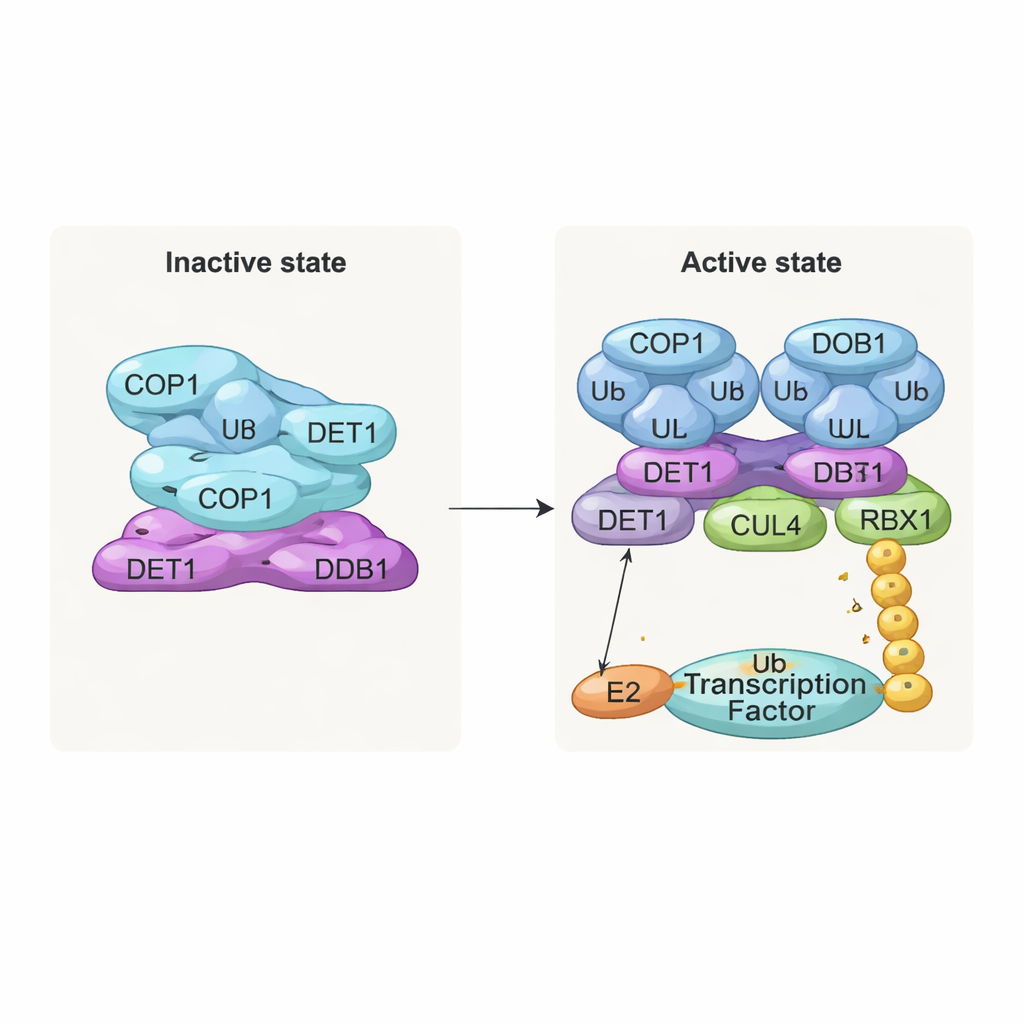
One of the most striking findings is a stacked, filament‑like structure made of repeated layers of the complex. In each layer, eight COP1 molecules intertwine through their coiled‑coil segments, forming a diamond‑shaped ring. Their substrate‑recognition regions, called WD40 domains, all face in one direction and are partly shielded by neighboring layers. This tight packing likely represents an “off” state in which access for target proteins is limited. In contrast, when the team examined complexes co‑expressed with COP1 substrates such as c‑Jun or ETS2, they saw a different, dimeric form: two COP1 molecules arranged side by side, with their WD40 domains facing outward and available to bind the short “VP” motifs found in many transcription factors. Biochemical tests confirmed that only this dimeric state efficiently supports the building of long ubiquitin chains on c‑Jun.
DET1 as a Flexible Bridge and Traffic Director
DET1 emerges as the key organizer that links all parts of the machine. Its N‑terminal region nests into DDB1, anchoring the complex to the CULLIN4 scaffold, while its claw region wraps around a family of E2 enzymes called Ube2e. Detailed mutational experiments show that DET1, not COP1, is what directly grips these E2s, positioning them close to COP1’s catalytic RING region. In this arrangement, Ube2e appears to help place the very first ubiquitin tags on the substrate. A second E2 enzyme, Ube2d3, is then recruited via the CULLIN4–RBX1 arm to extend these tags into longer chains that signal full degradation. In other words, DET1 acts as a flexible bridge that presents one E2 to COP1, while the larger scaffold brings in a second E2 to finish the job.
Why This Matters for Health and Disease
By mapping these structural states and their activities, the study explains how the same set of proteins can alternate between an idle, stacked form and an active dimer that targets transcription factors for destruction. This switch is especially important for factors like c‑Jun, which drive cell division and are tightly linked to cancer when misregulated. Understanding how COP1 and DET1 cooperate—how DET1’s claw selects specific E2 enzymes, how COP1’s WD40 surface recognizes VP‑containing substrates, and how the active dimer forms—provides a blueprint for future work. In the long term, drugs that stabilize the inactive stack or favor the active dimer could give researchers new ways to tune gene‑control pathways in cancer and other diseases where protein degradation goes awry.
Citation: Wang, S., Teng, F., Stjepanovic, G. et al. Cryo-EM structure of the human COP1-DET1 ubiquitin ligase complex. Nat Commun 17, 543 (2026). https://doi.org/10.1038/s41467-026-68375-7
Keywords: ubiquitin ligase, COP1, protein degradation, cryo-EM structure, c-Jun