Clear Sky Science · en
K48-ubiquitin-dependent proteases cut-up post-ER proteins
How Cells Decide Which Proteins to Destroy
Our cells constantly remove worn-out or damaged proteins, especially those embedded in membranes that act as gates and sensors. This paper reveals that cells use a tiny tag called ubiquitin not just as a generic “trash” label, but as a code that tells membrane proteins exactly how they will be broken down. Understanding this code helps explain how cells stay healthy and may guide future therapies that deliberately destroy disease-causing proteins.
A Cellular Barcode for Protein Disposal
Cells decorate unwanted proteins with chains of a small protein called ubiquitin. These chains can be linked together in different ways, rather like beads strung through different holes, and each pattern can signal a different outcome. The authors focused on membrane proteins that have already left the endoplasmic reticulum (ER) and travel through compartments such as the Golgi and endosomes. They asked whether two common chain types, called K48-linked and K63-linked ubiquitin, are truly interchangeable labels or whether they direct cargo into different destruction routes.
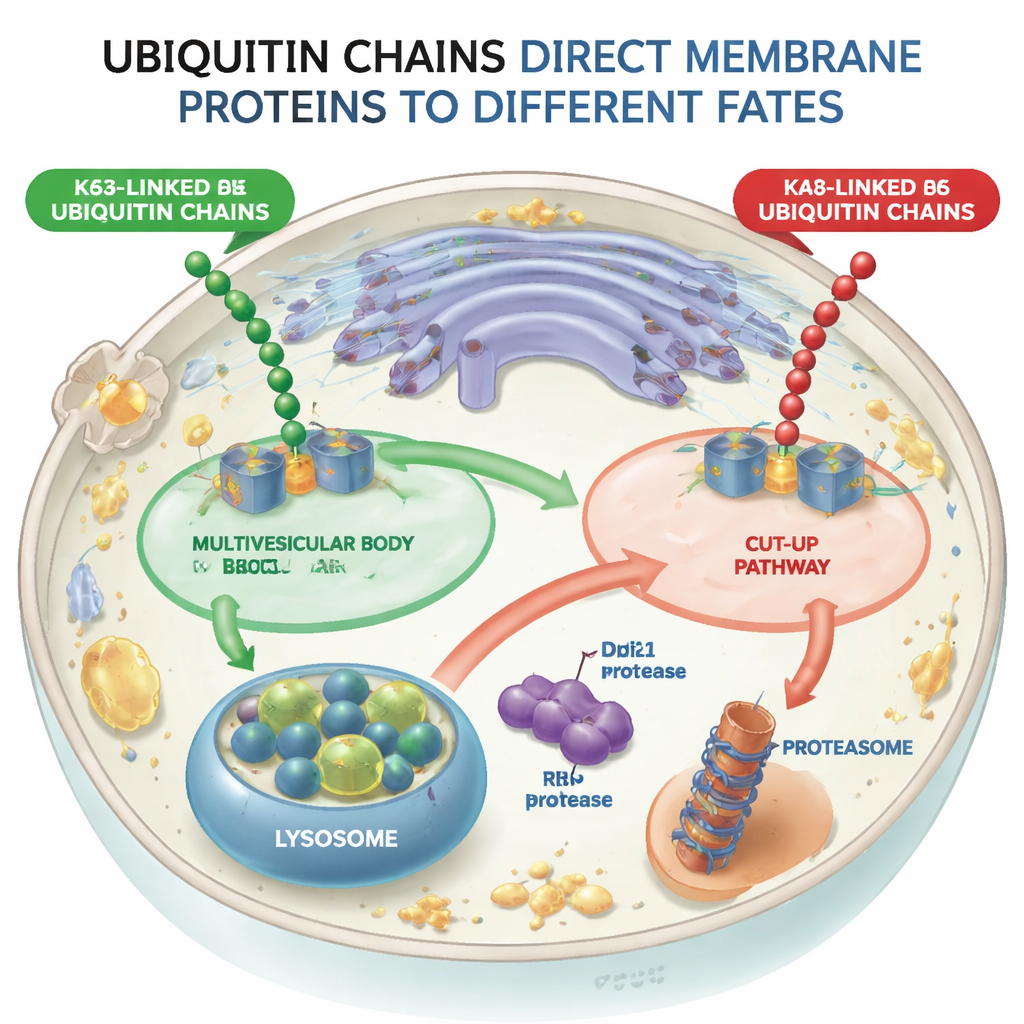
Two Diverging Paths for Tagged Proteins
Using yeast cells as a model, the researchers engineered molecular machines (ubiquitin ligases) that could attach either K48-linked or K63-linked chains to the same test membrane protein, a sorting receptor called Vps10. When Vps10 received K63-linked chains, it was sent into internal vesicles inside endosomes—structures known as multivesicular bodies—which eventually fuse with lysosomes, the cell’s digestive compartments. In contrast, when Vps10 carried K48-linked chains, it did not follow this multivesicular route. Instead, it was broken apart in a completely different way, showing that the cell distinguishes between these two chain types and that the multivesicular pathway effectively rejects K48-tagged cargo.
A “Cut-Up” Pathway that Slices Proteins in Two
The team discovered that K48-linked chains trigger a protein-shearing pathway they call CUT-UP, short for Cleavage of Ubiquitinated Targets by Ubiquitin-dependent Proteases. Rather than pulling the entire protein out of the membrane at once, CUT-UP chops it into pieces that are then finished off by existing cellular disposal systems. One key enzyme, Ddi1, floats in the cytosol and cuts off the portion of the membrane protein that faces the cytoplasm. That fragment is then digested by the proteasome, the cell’s main protein shredder. At the same time, another enzyme, Rbd2, sits within the membrane itself and cuts the part of the protein that faces the interior of the compartment, releasing it into the space that eventually connects to the lysosome. Blocking Ddi1, Rbd2, the proteasome, and lysosomal enzymes together was required to fully protect K48-tagged Vps10 from destruction, confirming that all of these components cooperate in the CUT-UP pathway.
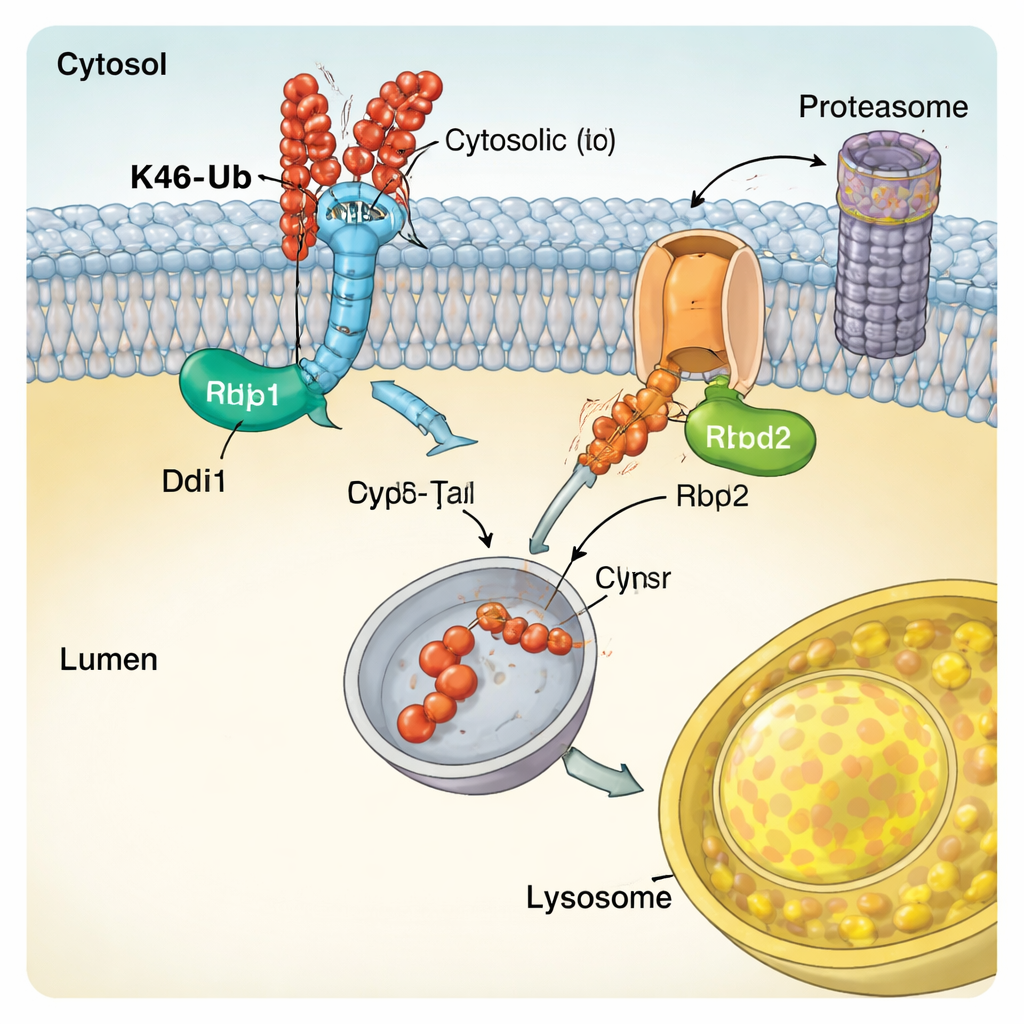
Specialized Enzymes that Read the Ubiquitin Code
To understand how CUT-UP recognizes its targets, the authors examined the molecular features of Ddi1. They showed that Ddi1’s central catalytic core not only cuts proteins but can also bind ubiquitin directly, and that its activity is tuned by extra regions that latch onto ubiquitin chains. Ddi1 showed a particular dependence on K48-linked chains, matching its role in CUT-UP. Rbd2, on the other hand, is a rhomboid protease that spans the membrane. The study found that Rbd2’s active site is essential for cutting the lumen-facing portions of Vps10 and that the enzyme travels through several secretory compartments, allowing it to act at multiple cellular locations. Similar enzymes exist in human cells and have been implicated in conditions ranging from neurodegeneration to cancer, suggesting that CUT-UP-like mechanisms may be broadly important.
Why This Matters for Health and Therapy
By showing that different ubiquitin chain types send the same membrane protein into sharply different fates—either multivesicular body sorting or CUT-UP-mediated slicing—this work provides strong evidence that cells use a genuine “ubiquitin code” to fine-tune protein disposal. The discovery of CUT-UP clarifies how stubborn membrane proteins can be dismantled piece by piece and highlights Ddi1 and Rbd2 as key players that read and enforce this code. For non-specialists, the takeaway is that cells do not simply mark proteins as garbage; they specify how that garbage will be taken out. As drug developers design tools that tag harmful proteins with ubiquitin to remove them, understanding pathways like CUT-UP will be essential to predict what fragments are produced and how that might affect cell behavior.
Citation: Minard, A.Y., Winistorfer, S., Yu, L. et al. K48-ubiquitin-dependent proteases cut-up post-ER proteins. Nat Commun 17, 1669 (2026). https://doi.org/10.1038/s41467-026-68367-7
Keywords: ubiquitin code, protein degradation, membrane proteins, proteasome, lysosome