Clear Sky Science · en
Biochemical principles of miRNA targeting in flies
Tiny RNA Messages That Steer Fly Cells
Inside every fruit fly cell, tiny molecules called microRNAs act like quality-control inspectors, deciding which genetic messages get turned into protein and which get silenced. This study asks a deceptively simple question: exactly how do these microRNAs recognize their targets in flies? By answering it, the authors move us closer to predicting, from sequence alone, which genes will be dialed up or down during development, behavior, and disease.
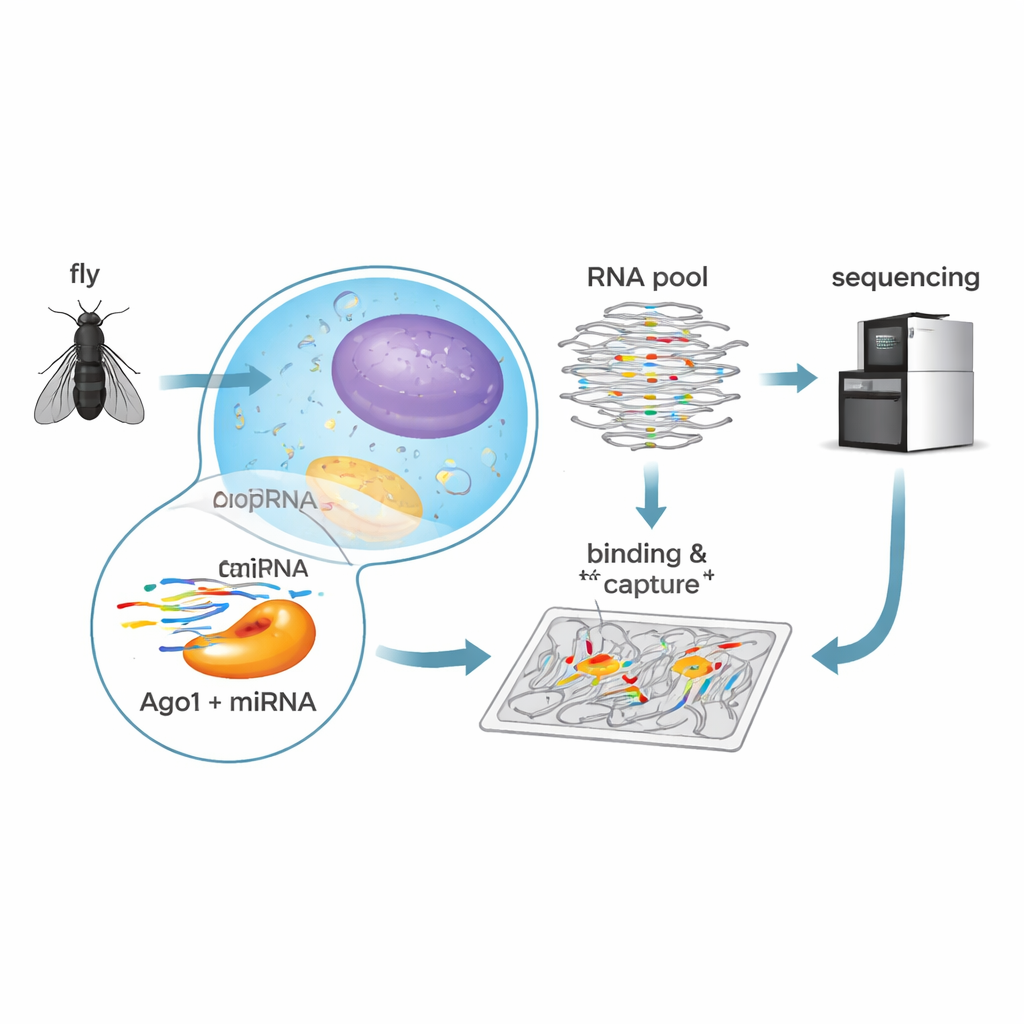
How Small RNAs Turn Down Genes
MicroRNAs are short pieces of RNA, about 22 building blocks long, that partner with a protein called Argonaute 1 (Ago1) in fruit flies. Together they scan much longer messenger RNAs (mRNAs), which carry instructions for making proteins. When a microRNA finds a partially matching stretch on an mRNA, the Ago1 complex can either cut the mRNA or block its translation into protein, reducing the gene’s output. In mammals, scientists have mapped these recognition rules in great detail, revealing a surprising variety of ways microRNAs can latch onto their targets. By contrast, the rules in flies have remained less clear, even though microRNAs there control key processes such as growth, timing of development, and daily sleep–wake cycles.
A High-Throughput Binding Test
To decode these rules in fruit flies, the researchers used a biochemical method called RNA Bind-n-Seq. They loaded purified fly Ago1 with one of five abundant microRNAs—let-7, bantam, miR-184, miR-11, or miR-124—each with known roles in fly development and brain function. They then mixed each Ago1–microRNA complex with a massive library of synthetic RNAs containing random sequences. After allowing binding to occur, they separated bound from unbound RNAs, sequenced the bound molecules, and used statistical modeling to calculate how tightly each sequence type was recognized. This approach provided quantitative binding strengths for hundreds of distinct target patterns in a single series of experiments.
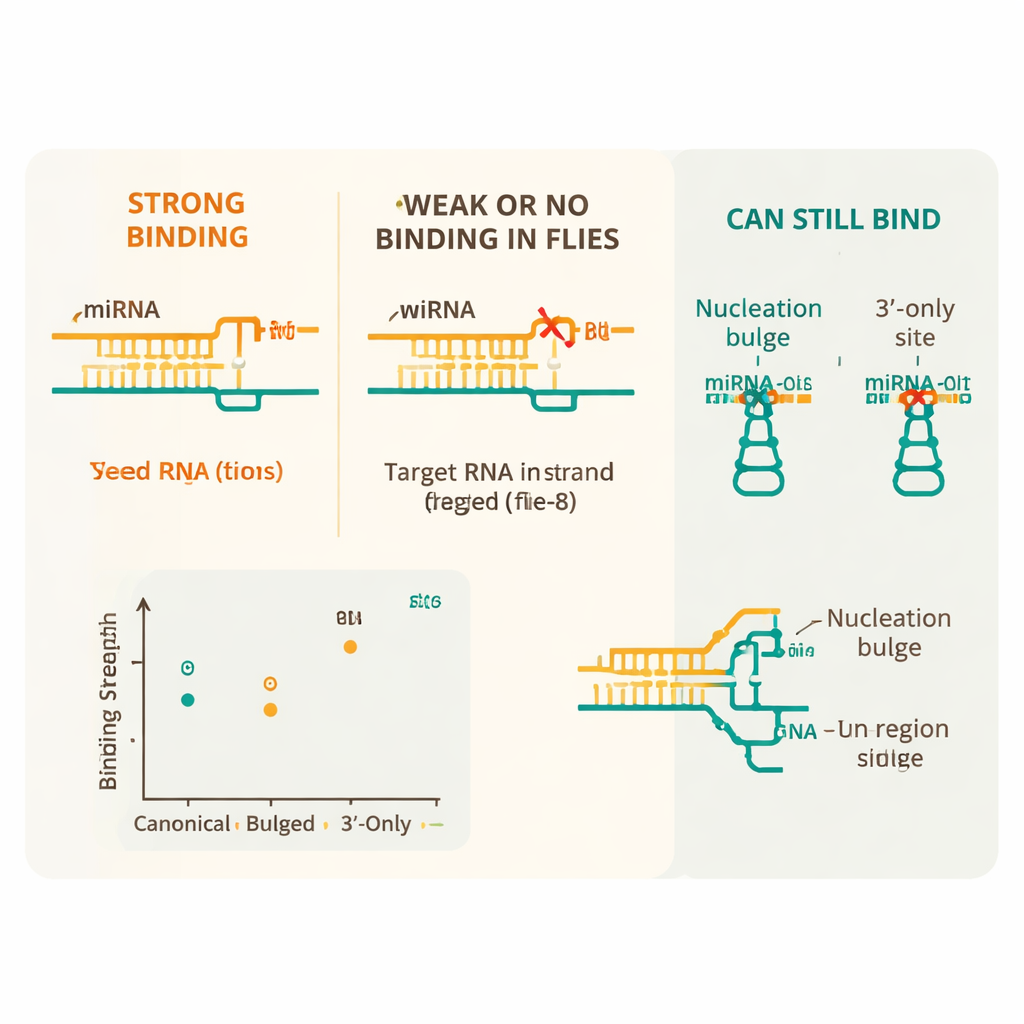
Simple Rules With a Few Clever Exceptions
The results reveal that fly microRNAs follow a stricter rulebook than their mammalian counterparts. The most important feature is a “seed” region—positions 2 through 8 of the microRNA—that must pair almost perfectly with the mRNA for strong binding. Canonical seed-matched sites, especially those with eight matching bases and a particular neighboring nucleotide, bound with the highest affinity. In contrast, even a single wobble pairing of the wrong type (a so-called G:U pair) within this seed sharply reduced binding, and two or more such imperfections made the interaction indistinguishable from background. Mismatches in the middle of the seed were especially damaging, emphasizing how sensitively Ago1 reads this core segment.
Hidden Flexibility Beyond the Core Match
Despite this overall rigidity, the study uncovered several important escape valves that let some imperfect sites still be recognized. Extra pairing between the microRNA’s tail end and the mRNA could compensate for a single flaw in the seed, restoring strong binding. Certain special arrangements, called nucleation bulges—where an extra nucleotide sticks out near the seed—also bound nearly as well as standard sites. The team further showed that Ago1 can bind “3′-only” sites, where the seed is not involved but the tail of the microRNA pairs strongly, and that it can efficiently cut targets with long central matches. Finally, they found that the surrounding sequence matters: sites flanked by A- and U-rich regions, which tend to keep RNA unstructured and accessible, were bound more strongly than the same sites buried in more rigid sequence contexts.
Why These Rules Matter for Fly Biology
Taken together, these findings show that fly microRNAs generally demand near-perfect matches in their seed region, with only a limited menu of tolerated exceptions. This simpler and tighter rule set contrasts with the broader flexibility observed in mammals. By providing hard numbers on how strongly different target patterns are bound, the work lays the groundwork for next-generation computer tools that can more accurately predict which fly genes will be controlled by which microRNAs. For non-specialists, the takeaway is that gene regulation in flies, though guided by tiny RNAs, follows clear biochemical principles—principles that can now be used to understand and eventually manipulate complex traits such as development, behavior, and disease resistance.
Citation: Vega-Badillo, J., Zamore, P.D. & Jouravleva, K. Biochemical principles of miRNA targeting in flies. Nat Commun 17, 1641 (2026). https://doi.org/10.1038/s41467-026-68360-0
Keywords: microRNA, Drosophila, Argonaute, RNA binding, gene regulation