Clear Sky Science · en
The role of low-complexity repeats in RNA–RNA interactions and a deep learning framework for duplex prediction
Sticky RNA Sequences That Shape Cell Behavior
Inside every cell, RNA molecules constantly bump into each other, forming fleeting partnerships that help control which genes are turned on, how proteins are made, and how cells develop. This study reveals that many of these RNA–RNA encounters are not random: they are guided by short, simple, highly repetitive sequences that act like molecular Velcro. The researchers also build an artificial intelligence tool that can spot where such RNA pairs are likely to form, opening new ways to explore how cells work in health and disease.
Simple Repeats with Powerful Effects
RNA is often described as a messenger that carries genetic information from DNA to proteins, but it also serves as a scaffold, a regulator, and a guide. Much of this activity depends on two RNA strands binding to each other. By combining data from several large experimental surveys in human and mouse cells, the authors show that the regions of RNA that actually engage in such pairing are strongly enriched in what they call low-complexity repeats. These are stretches built from short motifs—like runs of G and C bases—repeated again and again. Rather than being genomic “junk,” these repetitive pieces turn out to be prime docking sites where one RNA can latch onto many others, forming dense interaction hubs across the transcriptome. 
RNA Hubs for Development and Regulation
When the team examined which genes carry these repeat-rich contact sites, a striking pattern emerged: many of them encode proteins that control development and cell identity, such as transcription factors. Even in cancer cell lines that are not actively differentiating, RNAs linked to developmental programs were heavily involved in repeat-based contacts. The authors also zoomed in on specific long non-coding RNAs (lncRNAs), which are RNA molecules that do not code for proteins but often regulate them. For example, targets of the lncRNA TINCR and of another lncRNA important for motor neuron formation, Lhx1os, both showed an overabundance of complementary repeats. In these cases, simple repeats on the lncRNA are matched by complementary repeats in their partner RNAs, enabling stable pairings that can help tune the levels or translation of key developmental genes.
Where Proteins and Editing Enzymes Join In
These repeat-driven RNA contacts rarely act alone. The authors overlaid protein-binding maps onto their interaction data and found that many repeat-bearing contact sites are also recognized by RNA-binding proteins involved in translation control, RNA decay, and the formation of cytoplasmic granules such as P-bodies and stress granules. One protein in particular, STAU1, which can trigger the destruction of its RNA targets, frequently binds duplexes formed through low-complexity repeats. Knocking down STAU1 led to higher levels of RNAs involved in these duplexes, especially those carrying repeats, suggesting that repeat-mediated RNA pairing can flag transcripts for controlled degradation. The same repeat-rich regions also attract RNA editing enzymes such as ADAR1, which chemically modify specific bases within double-stranded RNA, hinting that low-complexity repeats help position editing sites that fine-tune RNA behavior.
Teaching a Neural Network to Read RNA Contacts
Standard computer programs try to predict RNA–RNA binding based mainly on thermodynamic stability—how much energy it would take to form or break a duplex. While useful, these models often miss real interactions observed in cells, especially between long RNAs. To move beyond simple energy rules, the authors trained a deep learning model called RIME that uses “language model” style embeddings: numerical representations of RNA sequences that encode patterns learned from huge collections of nucleic acid data. RIME is shown pairs of RNA segments and learns to classify whether they interact, using many real pairings from psoralen-based crosslinking experiments as positive examples and carefully constructed non-interacting pairs as negatives. 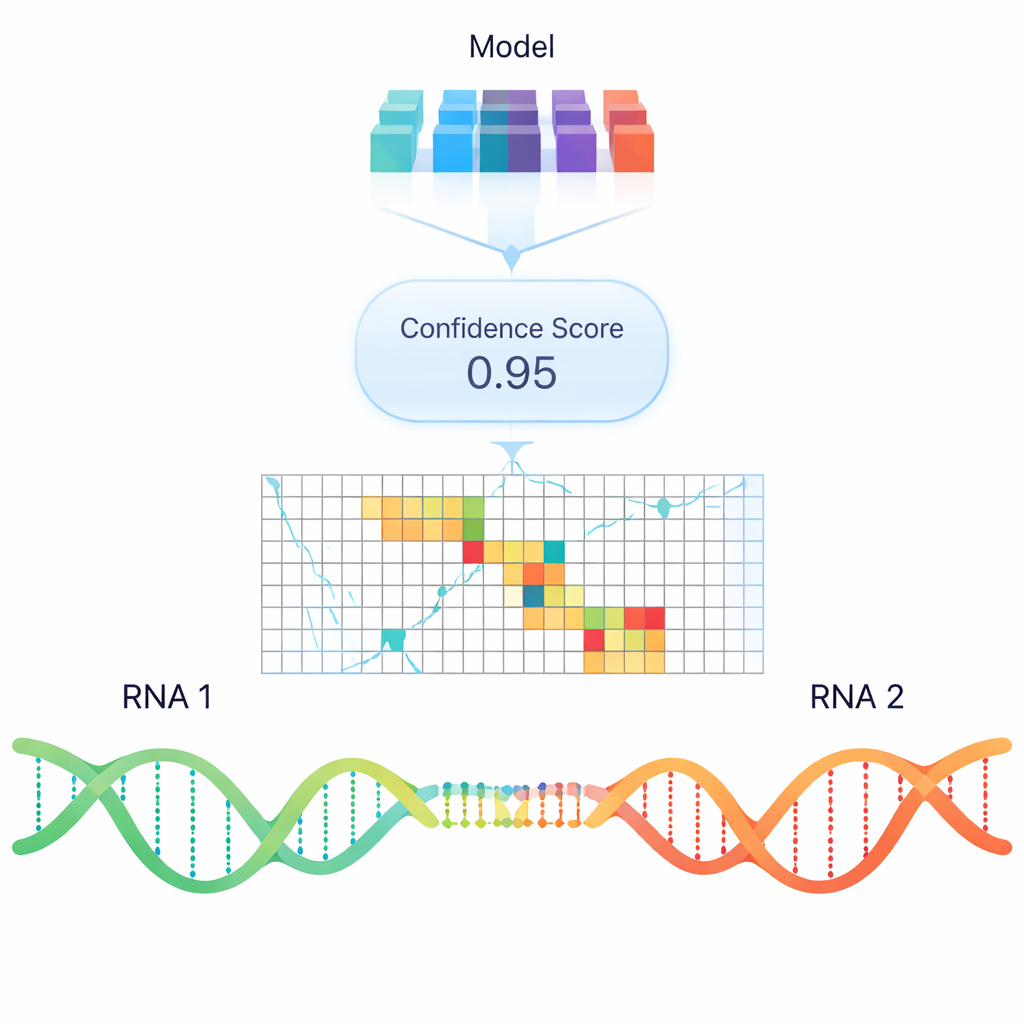
Smarter Predictions and New Biological Clues
When benchmarked against leading thermodynamics-based tools and another neural network method, RIME consistently performs better at distinguishing true RNA–RNA contacts from decoys, especially for high-confidence experimental interactions. It not only predicts whether two RNAs will pair, but also tends to highlight the exact regions involved, and it naturally learns that low-complexity repeats are strong predictors of contact. Remarkably, the same model, trained only on interactions between different RNAs, also works well for predicting internal base-pairing within a single RNA molecule, aligning with both structural experiments and classic folding algorithms. For non-coding regulators like TINCR, NORAD, and SMaRT, RIME successfully rediscovers known functional interaction sites and suggests additional candidate regions.
Why This Matters
For a lay reader, the key message is that short, repetitive stretches in RNA—once easy to dismiss as useless noise—act as central connection points in the cell’s RNA wiring diagram. They help bring RNAs together, invite in regulatory proteins and editing enzymes, and are heavily used in pathways that control how cells develop and respond to stress. The new RIME model gives researchers a powerful way to scan genomes for these RNA–RNA partnerships, including those that may go wrong in neurological and other diseases tied to repeat expansions. In essence, this work shows that understanding—and predicting—how simple RNA repeats stick together can reveal hidden layers of gene regulation.
Citation: Setti, A., Bini, G., Pellegrini, F. et al. The role of low-complexity repeats in RNA–RNA interactions and a deep learning framework for duplex prediction. Nat Commun 17, 1637 (2026). https://doi.org/10.1038/s41467-026-68356-w
Keywords: RNA–RNA interactions, low-complexity repeats, long non-coding RNA, deep learning, gene regulation