Clear Sky Science · en
Transcriptomic signature-guided depletion of intermediate alveolar epithelial cells ameliorates pulmonary fibrosis in mice
Why scarring lungs matter
Pulmonary fibrosis is a serious lung disease in which delicate air sacs gradually turn into stiff scar tissue, making every breath feel like hard work. Doctors know that this scarring begins when normal repair processes go off track, but it has been unclear which specific cells are to blame, and how to remove them without harming healthy lung tissue. This study explores a new way to “read” the molecular messages inside lung cells, use that information to identify harmful transitional cells, and selectively eliminate them to reduce scarring in mice.
Cells caught in the act of faulty repair
After lung injury, special surfactant-producing cells called type II cells normally multiply and then mature into thin type I cells that line the air sacs and allow oxygen to pass into the blood. In fibrosis, many of these cells get stuck in an in-between, or transitional, state instead of completing this journey. Previous single-cell RNA sequencing studies had found these intermediate cells in both mice and humans, but it was not clear whether they were merely bystanders or key drivers of disease. The authors focused on a mouse transitional cell type called Krt8+ alveolar differentiation intermediates, and their human counterparts known as aberrant basaloid cells, which appear in high numbers in scarred lungs.
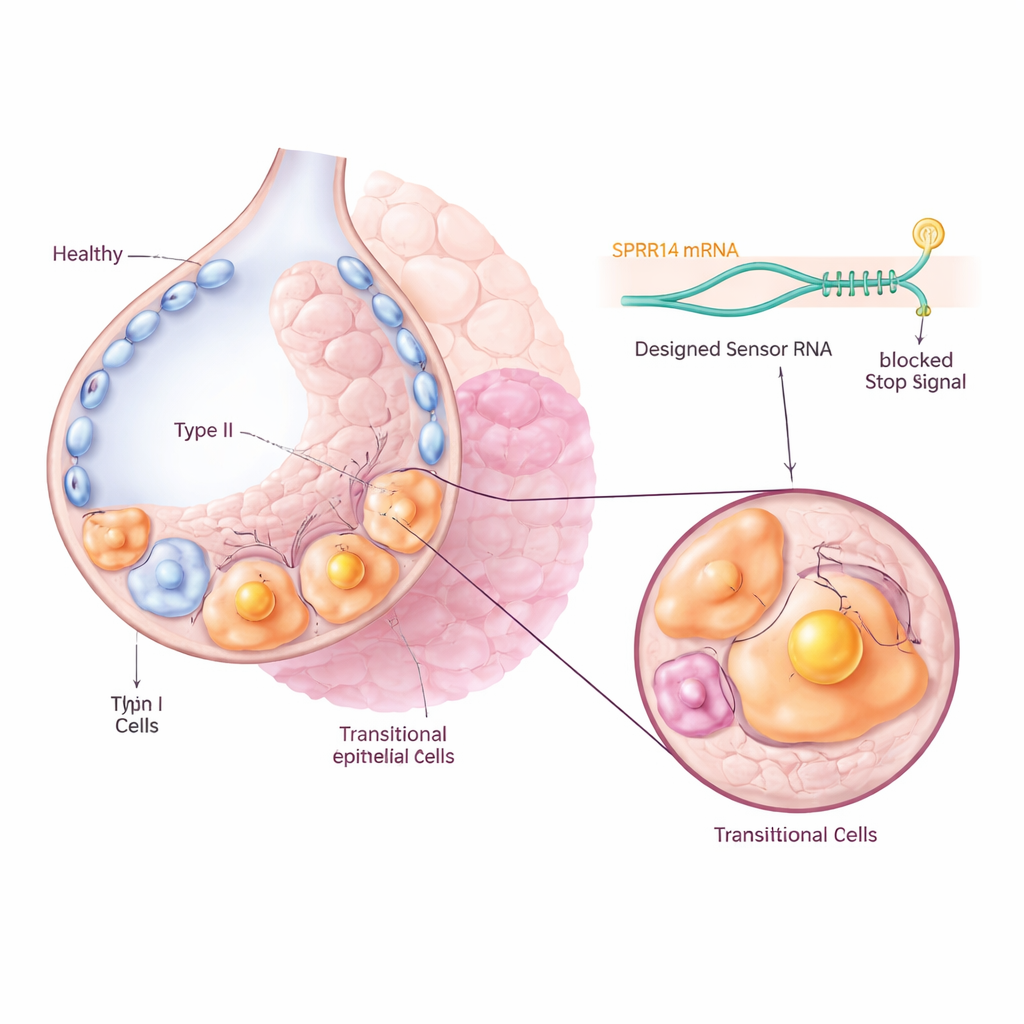
Finding a unique molecular name tag
To target these problem cells without touching their healthy neighbors, the team first searched large gene expression datasets for a marker that was turned on strongly and specifically in the transitional population. They identified a molecule called SPRR1A as a standout “name tag”: it was highly enriched in Krt8+ intermediate cells in fibrotic mouse lungs, and in KRT5-/KRT17+ aberrant basaloid cells from patients with idiopathic pulmonary fibrosis, but largely absent from normal lung cells. Microscopy of both mouse and human lung tissue confirmed that SPRR1A appeared mainly in injured, remodeled regions and not in healthy air sacs, suggesting it could serve as a precise handle for identifying the culprit cells.
Programming cells with an RNA sensor
The researchers then used a recently developed technology called CellREADR, which acts like an internal molecular sensor. They designed short RNA sequences that can base-pair with SPRR1A RNA inside a cell. When the sensor detects SPRR1A, it triggers the production of an attached “effector” protein, such as a fluorescent tag or a receptor that makes the cell sensitive to a drug. In mice, viruses were used to deliver these sensor–effector constructs to lung cells. Only cells actively producing SPRR1A switched on the fluorescent signal, allowing the team to track and isolate transitional cells directly from fibrotic lungs. Single-cell sequencing showed that the labeled cells closely matched the known transitional gene signature, confirming that the RNA sensor was accurately homing in on the intended population.
Turning harmful intermediates off
Next, the effector was switched from a harmless fluorescent protein to the diphtheria toxin receptor, so that SPRR1A-positive cells could be selectively killed by delivering diphtheria toxin. Timing the treatment to coincide with the peak abundance of transitional cells after chemical lung injury, the authors were able to remove roughly one third of this population. This targeted depletion led to strikingly less lung scarring: there was reduced collagen build-up, lower levels of fibrotic proteins, and more normal-looking air sac structures. Detailed analyses showed that most SPRR1A-positive cells carried signs of a stressed, aging-like state, with a small subset capable of high proliferation, suggesting that these intermediates both fail to repair the tissue properly and help sustain the fibrotic process.
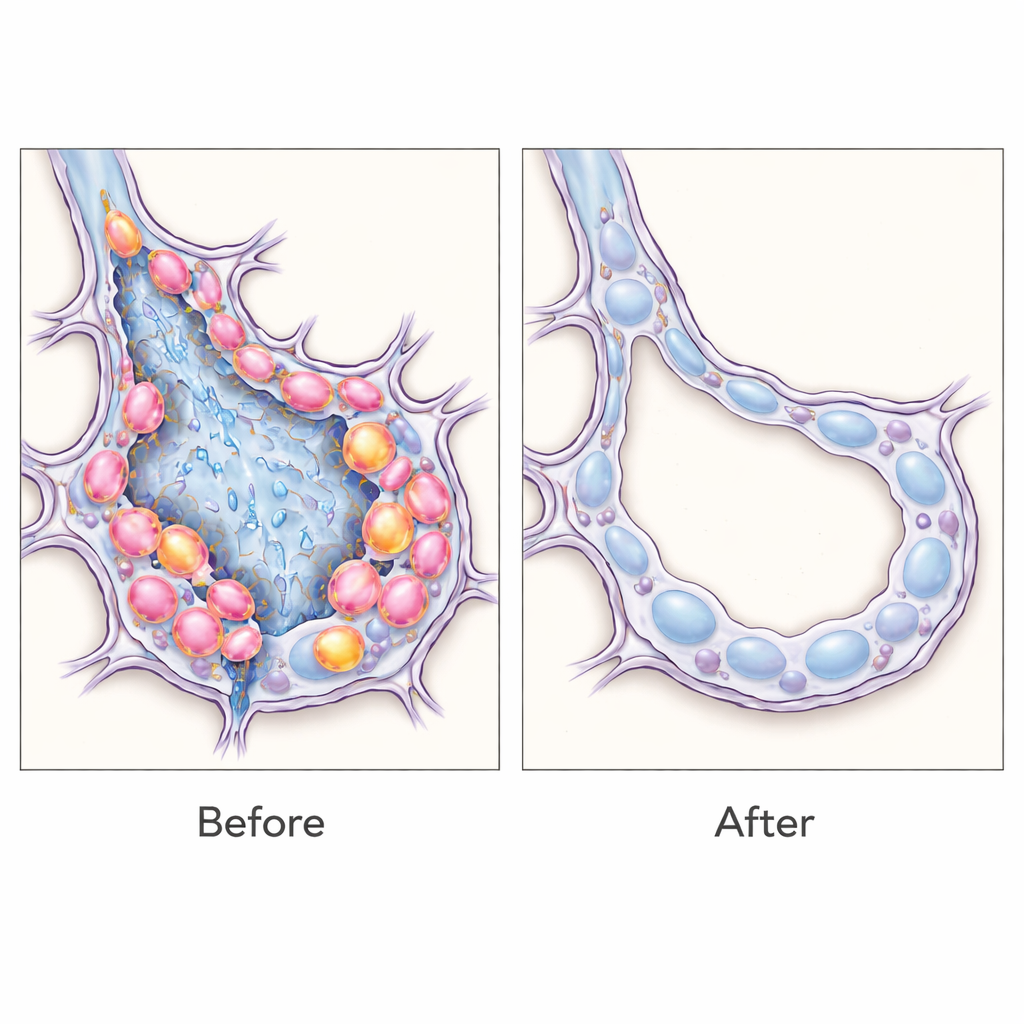
What this means for future treatments
These findings argue that transitional epithelial cells are not just markers but active drivers of pulmonary fibrosis, and that selectively removing them can tip the balance back toward healthier lung architecture in mice. More broadly, the work demonstrates a flexible strategy: by “reading” cell-specific RNA signatures, researchers can label, study, and even eliminate narrowly defined cell populations in living tissue without creating custom animal lines each time. While much work remains before such RNA-sensing tools could be safely adapted for people, this approach opens a path toward precision therapies that target the exact cell types that derail healing in chronic lung scarring and potentially other fibrotic diseases.
Citation: Peng, F., Jiang, Cs., Zheng, Z. et al. Transcriptomic signature-guided depletion of intermediate alveolar epithelial cells ameliorates pulmonary fibrosis in mice. Nat Commun 17, 1636 (2026). https://doi.org/10.1038/s41467-026-68354-y
Keywords: pulmonary fibrosis, alveolar epithelial cells, single-cell RNA sequencing, RNA sensing technology, cellular senescence