Clear Sky Science · en
Central amygdala single-nucleus atlas reveals chromatin and gene transcription dynamics in human alcohol use disorder
Why this research matters to everyday life
Alcohol use disorder (AUD) affects millions of people and their families, yet we still know surprisingly little about what long-term heavy drinking actually does to the human brain at the level of individual cells. This study zooms in on one small but critical region—the central amygdala, a hub for fear, stress, and motivation—to map how alcohol reshapes brain cells and their genetic control systems. By pairing cutting‑edge "single‑cell" sequencing with genetic studies of addiction risk, the authors begin to explain why some people are more vulnerable to AUD and how alcohol leaves lasting molecular scars in the brain.
Looking closely at a tiny brain hub
The central amygdala is a key output station in the brain’s emotion and stress network and is strongly linked to anxiety relief, craving, and compulsive alcohol use. In this work, researchers analyzed about 175,000 individual cell nuclei taken postmortem from the central amygdala of 50 people—22 with a history of alcohol use disorder and 28 without AUD. Using a technology that reads both gene activity (RNA) and chromatin accessibility (how "open" or "closed" DNA is) in the same cell, they built a detailed atlas of all major brain cell types in this region, including several kinds of neurons and support cells such as astrocytes and microglia. They found that inhibitory neurons—which dampen or sculpt brain activity—are especially abundant in the central amygdala and appear particularly affected in people with AUD. 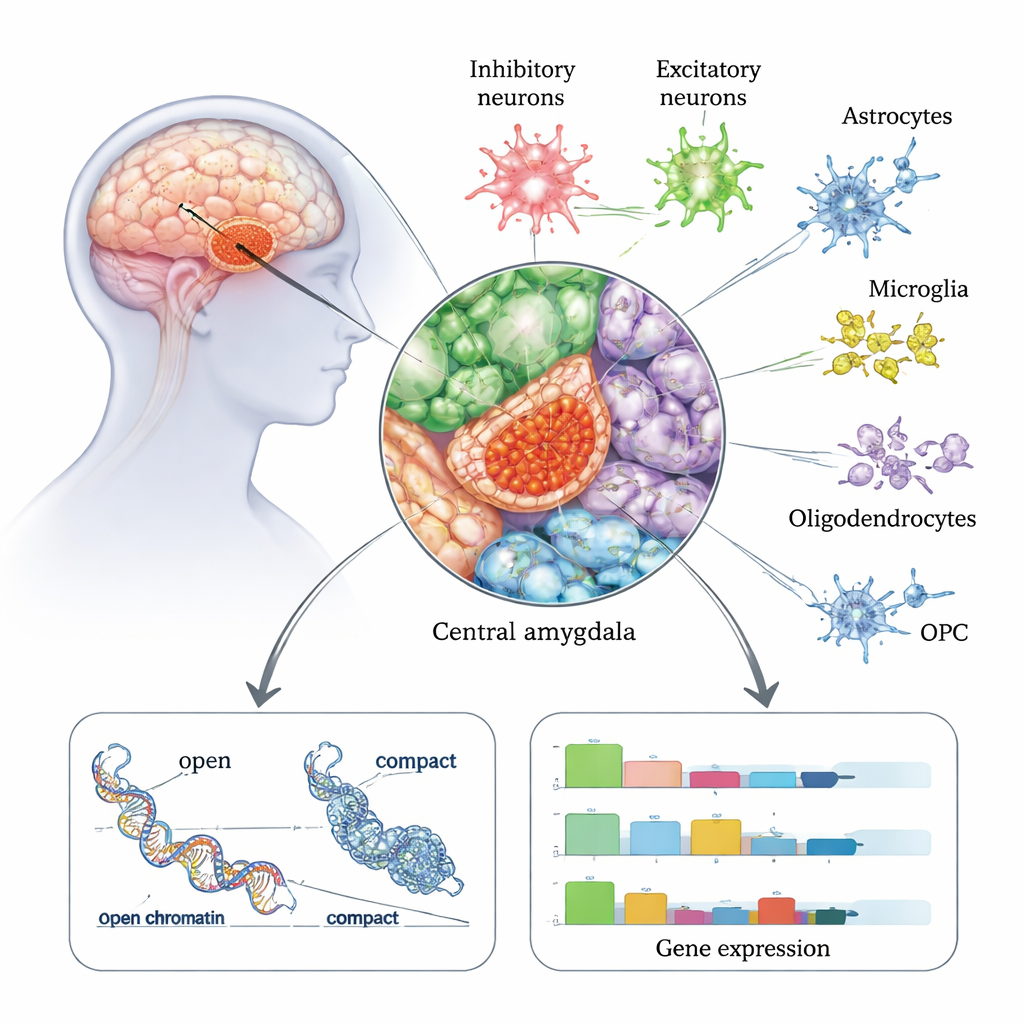
Which cells and genes change in alcohol use disorder?
The team systematically compared gene activity in people with and without AUD, cell type by cell type. They discovered more than 1,800 genes whose activity was altered in specific cell populations, with the largest shifts occurring in inhibitory neurons, followed by excitatory neurons and astrocytes. One subtype of inhibitory neuron marked by the peptide proenkephalin (PENK) showed especially strong changes. Many of the affected genes are involved in communication between brain cells, particularly the balance between the excitatory signal glutamate and the inhibitory signal GABA. For example, the gene GABRA2, which helps build a major type of GABA receptor, was increased in certain inhibitory neurons, while GRM8, a glutamate receptor, and NCAM1, a cell-adhesion molecule important for synapses, changed in opposite directions across neuron subtypes. These patterns suggest that the fine tuning of excitation and inhibition in the central amygdala is disturbed in AUD.
The brain’s control switches: chromatin and regulatory elements
Genes do not act alone; they are controlled by stretches of DNA that function like switches and dimmers. By looking at chromatin—the way DNA is packaged—the researchers identified more than half a million candidate regulatory elements and linked them to nearby genes in a cell-type-specific way. Nearly half of these elements were unique to a single cell type, highlighting how specialized different brain cells are. Many regulatory elements that changed in AUD were connected to genes involved in calcium handling and glutamate signaling, such as CALN1, a calcium-binding gene strongly active in neurons. In inhibitory neurons, CALN1 showed both increased expression and dense clusters of regulatory loops, implying that alcohol exposure reshapes the local DNA architecture controlling this gene. 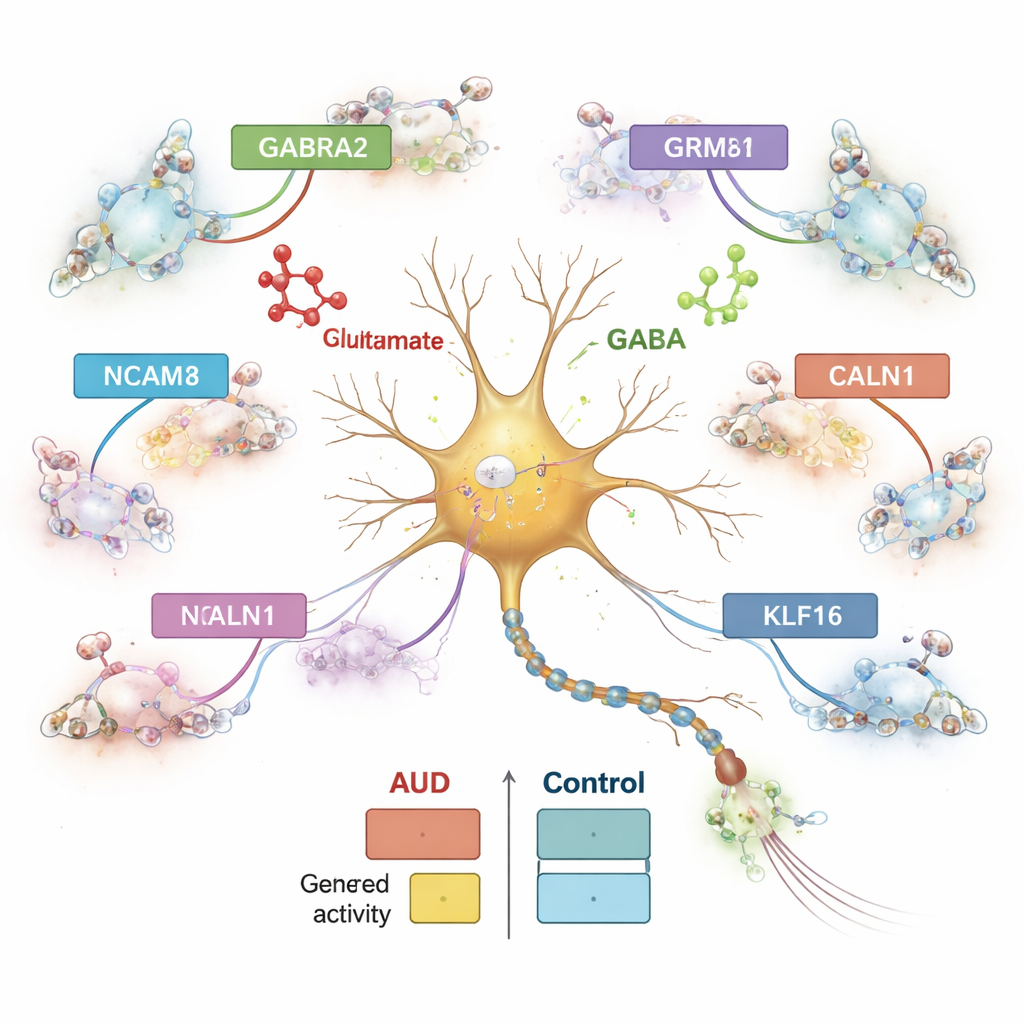
From risk genes to real brain changes
Genetic studies have identified many DNA variants that raise the risk of developing alcohol problems, but where and how they act in the brain has been unclear. By overlaying their single‑cell maps with large genome‑wide association studies of problematic alcohol use, the authors found that risk genes are disproportionately active and dysregulated in neurons of the central amygdala, especially inhibitory ones. They pinpointed over 200 likely causal risk variants sitting inside regulatory elements that are open in specific cell types. Some fell in or near genes already tied to addiction, such as CACNA1C and DRD2, which help control neuronal excitability and dopamine signaling. Others, including SEMA6D and NF1, emerged as new candidates. The team also identified a family of transcription factors called Kruppel-like factors (notably KLF16) that appear to coordinate wide-ranging changes in calcium and glutamate-related genes, and they confirmed parts of this regulatory network in a mouse model of acute alcohol exposure.
What this means for understanding and treating AUD
In plain terms, this study shows that alcohol use disorder is not just a diffuse brain problem but a highly organized disturbance concentrated in particular cell types and molecular pathways within a critical emotional hub. Inhibitory neurons of the central amygdala, along with astrocytes and other support cells, show coordinated changes in gene activity and in the DNA switches that control those genes—especially in systems that manage glutamate, GABA, and calcium signals. By connecting these cellular changes to genetic risk variants, the work offers a roadmap linking inherited vulnerability to concrete molecular events in the brain. Over time, such cell‑specific maps may guide the development of more precise treatments that restore the balance of excitation and inhibition, rather than acting broadly on the whole brain.
Citation: Lee, C.Y., Hwang, A., McRiley, D. et al. Central amygdala single-nucleus atlas reveals chromatin and gene transcription dynamics in human alcohol use disorder. Nat Commun 17, 1634 (2026). https://doi.org/10.1038/s41467-026-68351-1
Keywords: alcohol use disorder, central amygdala, single-cell genomics, inhibitory neurons, gene regulation