Clear Sky Science · en
Mechanism of SARS-CoV-2 resistance to nucleotide analog-based antivirals
How the Virus Outsmarts Key Antiviral Drugs
The COVID-19 virus belongs to a rare group of RNA viruses that can proofread its genetic material, making it unusually good at resisting some of our best antiviral drugs. This study digs into the atomic‑level details of how SARS‑CoV‑2 detects and removes a widely used class of medicines called nucleotide analogs, explaining why drugs that work well against other viruses often underperform against coronavirus infections.
A Tug‑of‑War Over the Viral RNA
SARS‑CoV‑2 copies its ~30,000‑letter RNA genome using a large molecular machine called the replication–transcription complex. At its heart sits the RNA‑dependent RNA polymerase (RdRp), which builds new RNA strands, and a separate unit, the exonuclease (ExoN), which proofreads and trims mistakes. Many antiviral pills, including drugs originally developed for hepatitis C, mimic the natural RNA building blocks closely enough to be inserted by RdRp, but with subtle tweaks that stall copying or introduce errors. Unfortunately, coronaviruses have ExoN, which can recognize these impostors once they are in the RNA chain and cut them back out, rescuing viral replication.
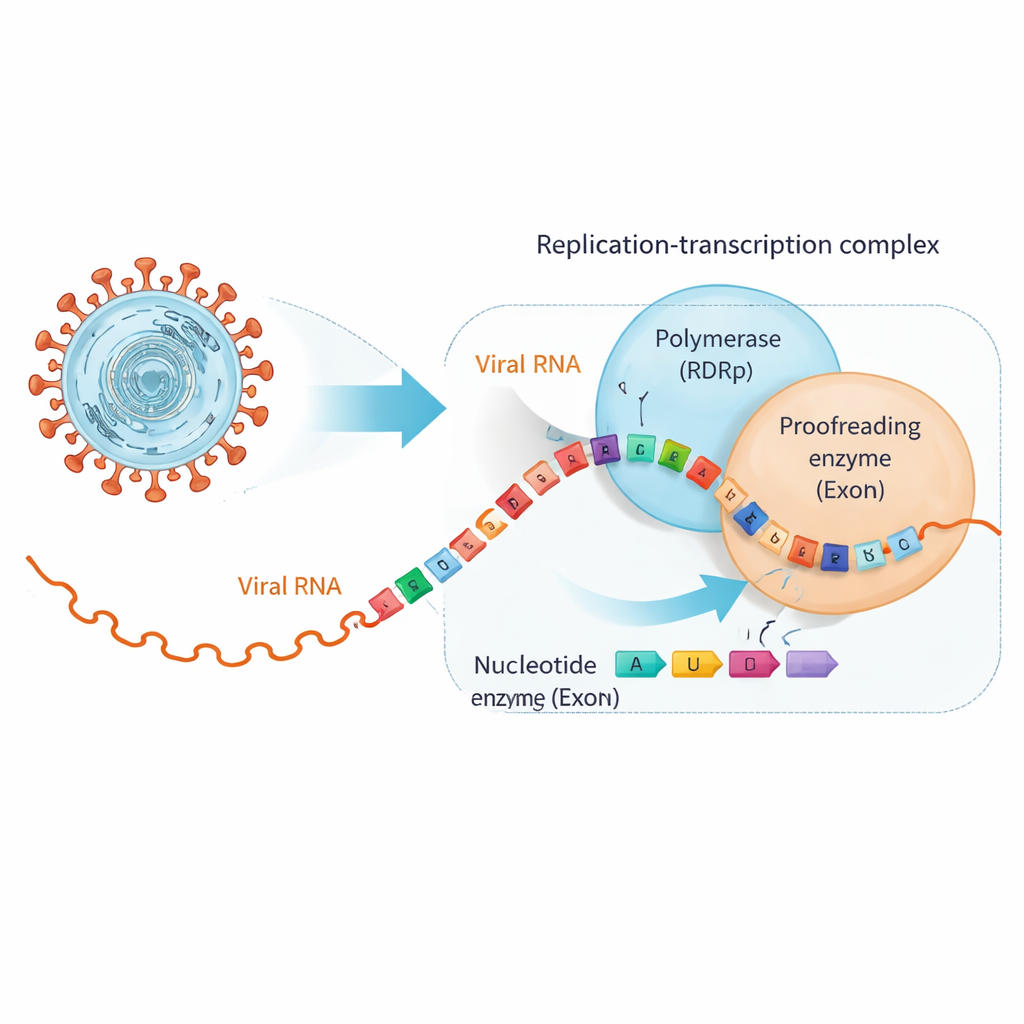
When Helpful Drugs Make RNA Switch Partners
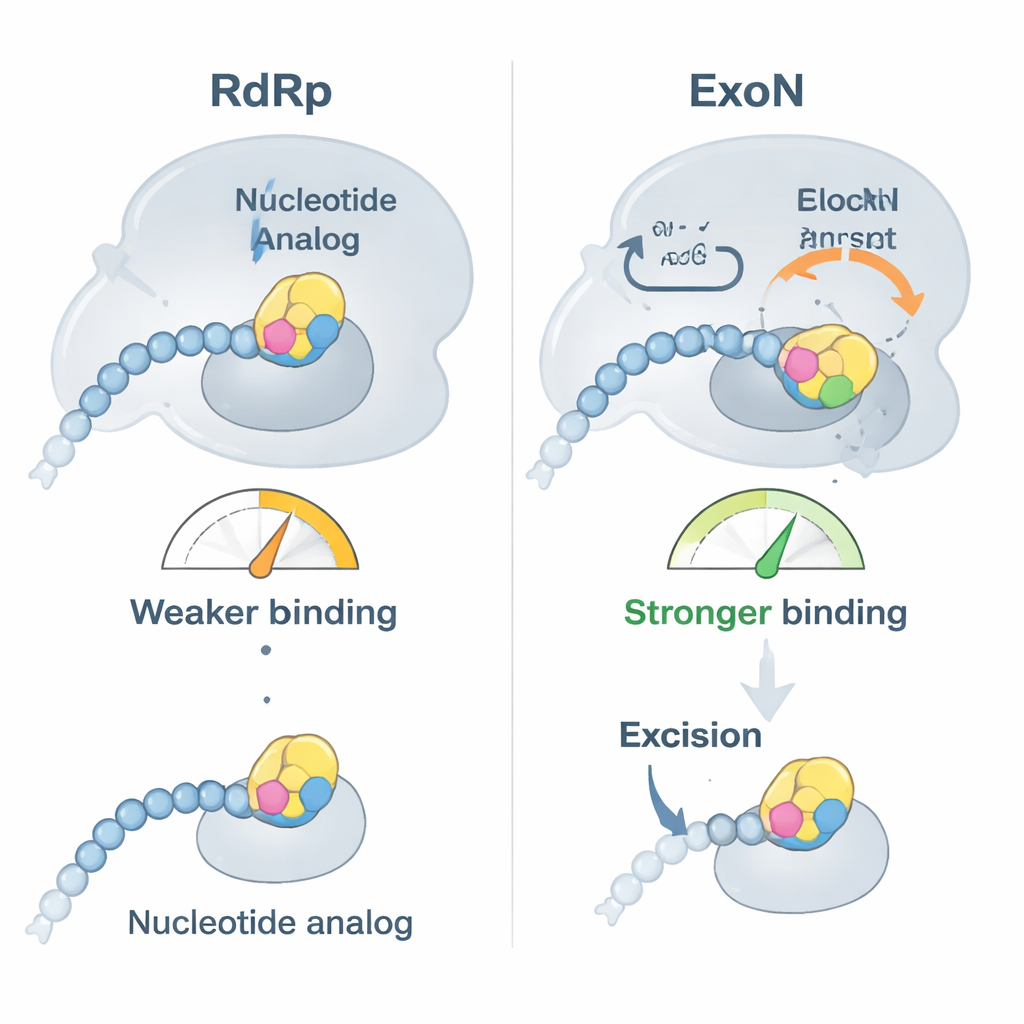
The researchers focused on two clinically important antivirals, bemnifosbuvir and sofosbuvir, both designed to act as defective RNA building blocks. They first showed that SARS‑CoV‑2’s polymerase readily inserts these analogs at the end of an RNA strand and then largely stops extending the chain, as intended. However, binding tests revealed a twist: once an analog sits at the RNA’s tip, the RNA no longer attaches as firmly to the polymerase but instead binds more tightly to ExoN. In effect, the drug‑modified RNA is nudged away from the copy machine and handed over to the proofreader, which then has the chance to snip off the problematic nucleotide and allow replication to continue.
Proofreading That Is Slowed but Not Stopped
Biochemical time‑course experiments showed that ExoN does remove bemnifosbuvir and sofosbuvir from RNA, but more slowly than it removes normal nucleotides. In cell‑free reactions, both analogs strongly blocked further RNA synthesis—yet when an active ExoN was added, a significant fraction of stalled RNA chains were rescued and extended. A disabled ExoN mutant could not do this, confirming that the proofreading activity is responsible for undoing much of the drugs’ effect. The presence of polymerase alongside ExoN actually sped up removal of the analogs, suggesting that the two viral enzymes work in concert to clear out obstacles and keep replication running.
Watching the Proofreader at Atomic Resolution
To understand how ExoN recognizes and responds to these drugs, the team used high‑resolution cryo‑electron microscopy to solve structures of the ExoN complex bound to RNA strands ending in either bemnifosbuvir or sofosbuvir. These structures, sharp enough to see individual chemical groups, revealed that the modified sugar ring on each drug fits into a hydrophobic pocket formed by a short loop of the ExoN protein. This extra snug fit explains the stronger binding of drug‑containing RNA to ExoN. But it also has an unexpected side effect: by tugging on that loop, the drugs disturb the precise arrangement of a nearby catalytic loop that holds a key histidine residue in place. When this histidine swings away from the cutting site, the active center of ExoN becomes partially deactivated, slowing but not fully preventing excision of the analog.
A Built‑In Switch That Tunes Viral Proofreading
Mutating individual amino acids within the sensitive loop confirmed its importance. Changes to four conserved residues greatly reduced ExoN’s ability to cut both standard RNA ends and drug‑terminated ends, and also altered which final nucleotide ExoN prefers to remove. This identifies the loop as an allosteric regulator—a built‑in mechanical switch that senses what kind of nucleotide is at the RNA tip and adjusts the enzyme’s activity. The structures further show that ExoN recognizes different bases (A, U, C, or G) through flexible hydrogen bonding, but has the hardest time accommodating guanine‑like bases, which is relevant because bemnifosbuvir mimics guanosine.
What This Means for Future COVID‑19 Pills
For non‑specialists, the key message is that SARS‑CoV‑2 carries a sophisticated “spell‑checker” that can pull certain drugs out of its genome after they have done their job of stalling replication. Bemnifosbuvir and sofosbuvir do weaken the virus’s copy machine, but at the same time they steer RNA toward the proofreader that can partially undo their effects. By revealing exactly how the viral proofreader grips these drugs, and how a tiny regulatory loop switches the enzyme between active and less‑active states, this work offers a roadmap for designing new antiviral nucleotides that either bind poorly to ExoN, lock it in an inactive conformation, or exploit its difficulty in handling guanine‑like structures. Such next‑generation molecules could be far harder for the virus to “erase,” improving our ability to treat COVID‑19 and future coronavirus outbreaks.
Citation: Liu, C., Li, Y., Cao, X. et al. Mechanism of SARS-CoV-2 resistance to nucleotide analog-based antivirals. Nat Commun 17, 1601 (2026). https://doi.org/10.1038/s41467-026-68304-8
Keywords: SARS-CoV-2, antiviral resistance, nucleotide analogs, RNA proofreading, coronavirus replication