Clear Sky Science · en
HEXIM1 inter-monomer autoinhibition governs 7SK RNA binding specificity and P-TEFb inactivation
Why this tiny RNA matters for gene control
Every cell must decide when to turn genes on and off, and it does so by carefully controlling an enzyme that drives RNA production. This paper uncovers how a small protein called HEXIM1, working together with a non‑coding RNA named 7SK, acts like a safety catch on that enzyme. Understanding this molecular safety system helps explain how our cells prevent runaway gene activity, and it also sheds light on how viruses like HIV hijack the same machinery.
A molecular brake on gene expression
In human cells, a key step in gene expression is controlled by a factor called P‑TEFb, which helps RNA polymerase II move from a paused state into full-speed transcription. HEXIM proteins, especially HEXIM1, are central to keeping P‑TEFb in check. They do this inside a large assembly called the 7SK RNP, built around the 7SK RNA. When HEXIM1 and 7SK team up, they lock P‑TEFb away so it cannot drive transcription. Until now, the missing piece was how HEXIM1 itself avoids grabbing P‑TEFb too soon, and how 7SK RNA flips HEXIM1 from an inactive to an active inhibitor at the right time.
A self‑hugging protein held in check
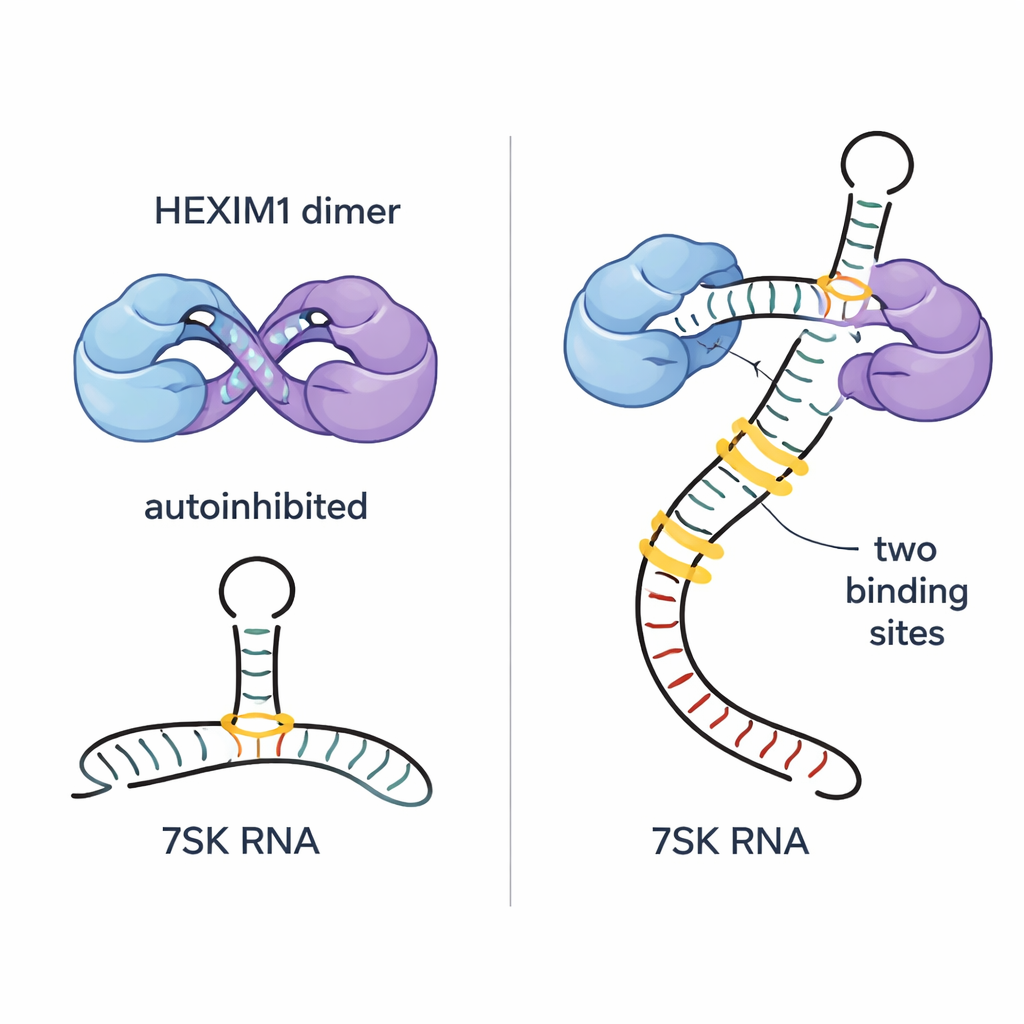
The authors show that HEXIM1 naturally forms a dimer—two identical protein chains paired together—and that the flexible central parts of each chain wrap around each other in a “self‑hugging” arrangement. In this state, a crucial short segment called the PYNT motif, which normally binds and shuts down the P‑TEFb kinase, is tucked away at the interface between the two chains. Using sensitive NMR and biophysical methods, the team mapped how basic (positively charged) and acidic (negatively charged) regions of the two monomers interact to create this autoinhibited dimer. This built‑in restraint keeps HEXIM1 from randomly clamping onto P‑TEFb when 7SK RNA is not present.
How 7SK RNA unlocks HEXIM1
The 7SK RNA contains several structural features, including a long stem‑loop region called SL1. By dissecting this RNA into smaller fragments and tracking subtle changes in NMR signals, the researchers pinpointed five possible HEXIM1 binding sites along SL1. Two of these sites stand out as high‑affinity docking points; both share an A–U–rich double‑helix section and a nearby U‑rich bulge that can form special three‑base interactions. The team found that each half of the HEXIM1 dimer binds one of these two strong sites on linear 7SK RNA. When both sites are engaged, the central region of HEXIM1 rearranges, breaking the self‑hugging contacts and exposing the hidden PYNT motifs, effectively priming the protein to capture P‑TEFb.
Choosing the right RNA shape—and avoiding the wrong ones
7SK RNA does not exist in just one shape; it can switch between a linear form, which presents both high‑affinity sites, and a circular‑like form with only one suitable site. The authors show that HEXIM1’s autoinhibition makes it surprisingly picky: the dimer binds weakly to single sites on their own, but much more tightly and cooperatively to the full linear SL1 segment that offers both strong sites. This dual‑site requirement prevents HEXIM1 from getting stuck on random RNAs elsewhere in the cell and makes its binding highly specific for the linear 7SK conformation that assembles the active repressive complex. They also demonstrate that adding extra negative charges near the RNA‑binding region (mimicking phosphorylation at one key serine) weakens RNA binding without fully disrupting autoinhibition, a potential way for cells to release P‑TEFb when needed.
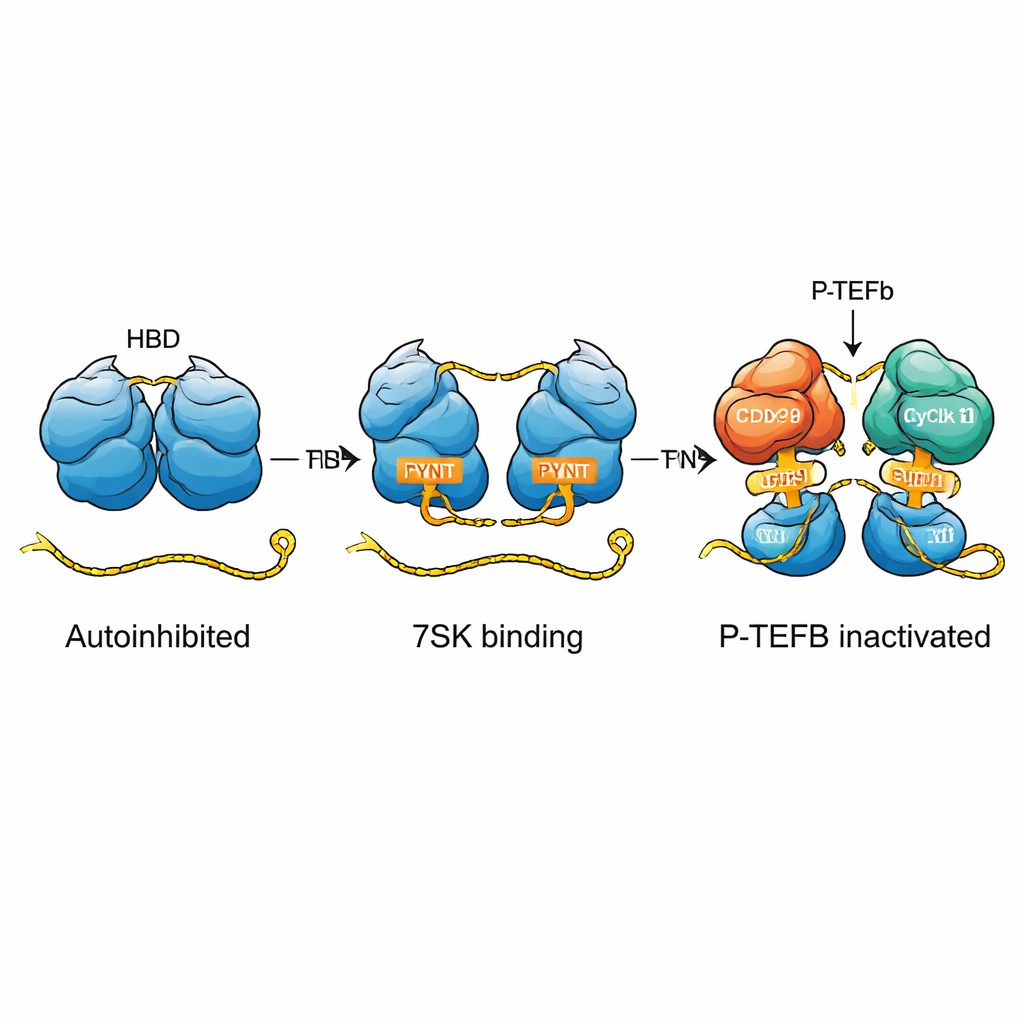
From molecular switch to cellular impact
Overall, the study reveals HEXIM1 as a finely tuned molecular switch. In its resting, self‑hugging state, it keeps its P‑TEFb‑blocking PYNT segments hidden. Only when it encounters linear 7SK RNA with two properly arranged binding sites does the protein snap into an “on” state, exposing these segments so it can capture and inactivate P‑TEFb—often two copies at once. This mechanism explains how cells use a small RNA and a flexible protein to impose precise control over transcription, and it provides a clearer framework for understanding how cellular signals, chemical modifications, or viral proteins like HIV Tat can tip the balance between paused and active gene expression.
Citation: Yang, Y., Murrali, M.G., Galvan, S. et al. HEXIM1 inter-monomer autoinhibition governs 7SK RNA binding specificity and P-TEFb inactivation. Nat Commun 17, 1570 (2026). https://doi.org/10.1038/s41467-026-68285-8
Keywords: transcription regulation, 7SK RNA, HEXIM1, P-TEFb, HIV Tat