Clear Sky Science · en
cfGWAS reveal genetic basis of cell-free DNA end motifs
DNA Clues Floating in Our Blood
Every moment, tiny fragments of DNA drift through our blood, released from dying and renewing cells. Doctors already use this “cell-free DNA” to screen pregnancies and search for early signs of cancer, but we still know little about how these fragments are produced and cleared. This study uses genetic sleuthing on an unprecedented scale to uncover which human genes shape the fine-grained patterns at the tips of these DNA fragments—information that could sharpen liquid biopsies and point to new treatments for immune and inflammatory diseases. 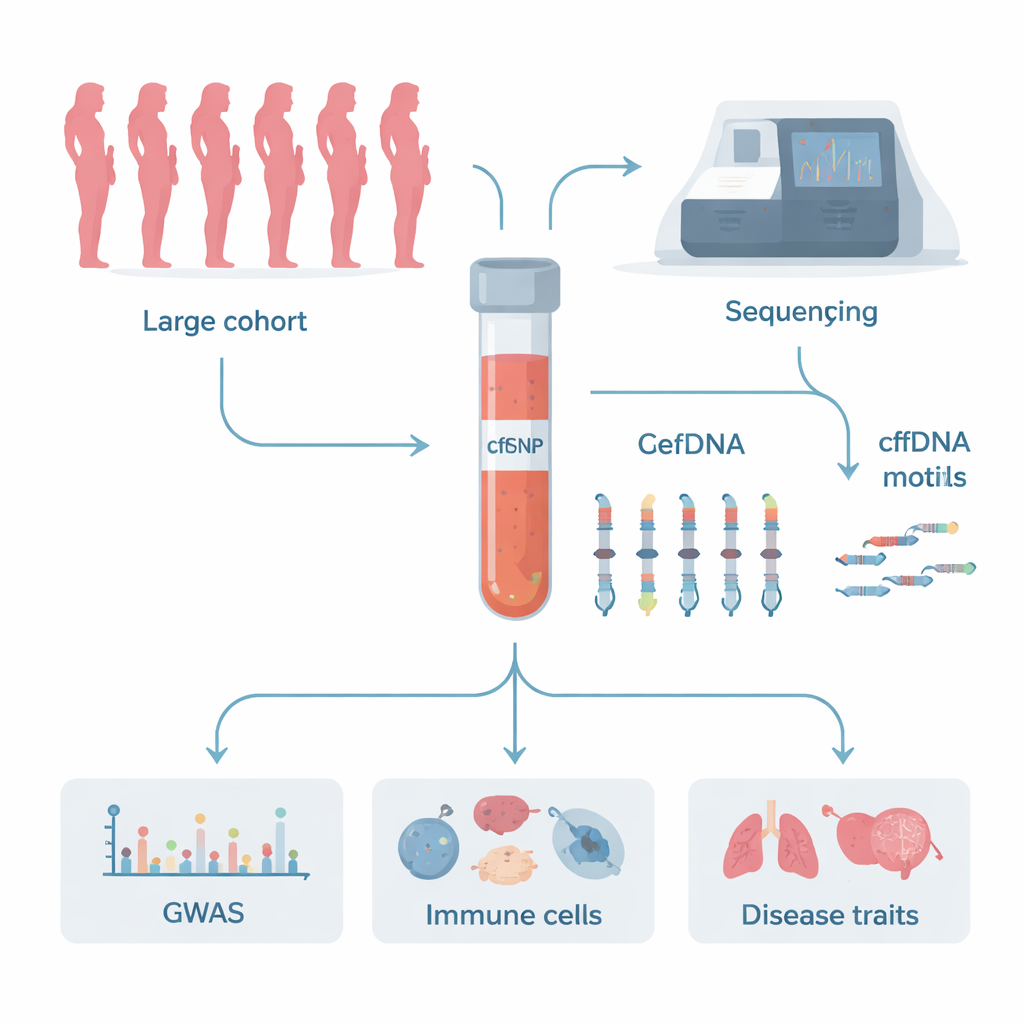
What Are These Free-Floating DNA Fragments?
Cell-free DNA (cfDNA) consists of short pieces of genetic material shed into body fluids like blood, saliva, and urine as cells die or actively release their contents. In pregnancy, fetal cfDNA in a mother’s blood enables noninvasive prenatal testing. In cancer, tumor-derived cfDNA can flag disease long before a lump is visible. Beyond the sheer amount of cfDNA, researchers now pay attention to its “fragmentomics”: how long each piece is, how it is cut, and which short letter patterns, or “end motifs,” appear at the starting ends of the fragments. These end motifs act like molecular fingerprints of how and where DNA was chopped up, reflecting the actions of enzymes and the types of cells that contributed the DNA.
A Massive Genetic Scan of DNA Fragment Patterns
To find out which genes influence these end motifs, the authors carried out a new kind of genome-wide association study they call cfGWAS. They analyzed blood samples from 28,016 pregnant women in China who underwent routine prenatal screening. From the very same low-coverage sequencing data, they extracted both the women’s genetic variants and the detailed frequencies of all 256 possible four-letter end motifs at the tips of cfDNA fragments. Using statistical models that accounted for age, gestational week, and technical factors, they scanned nearly 2.9 million common genetic variants to see which ones tracked with shifts in these motif patterns across individuals.
Key Genes Behind How DNA Gets Cut
The cfGWAS uncovered 15 particularly robust genetic regions linked to 176 different end motifs. Some hits confirmed the starring role of known DNA-cutting enzymes, including DFFB and DNASE1L3, which help chop up DNA during programmed cell death, and a related enzyme gene, DNASE1L1. More surprisingly, the strongest signal came from PANX1, a gene that encodes a channel in the cell membrane involved in cell communication, inflammation, and cell death. People with different versions of these genes showed distinct patterns in which end motifs were common or rare in their blood. The researchers replicated most of these signals in three independent cohorts, including another large pregnancy group in China, a Dutch pregnancy study, and a smaller group of healthy non-pregnant adults, showing that the findings are robust and not limited to one hospital or country. 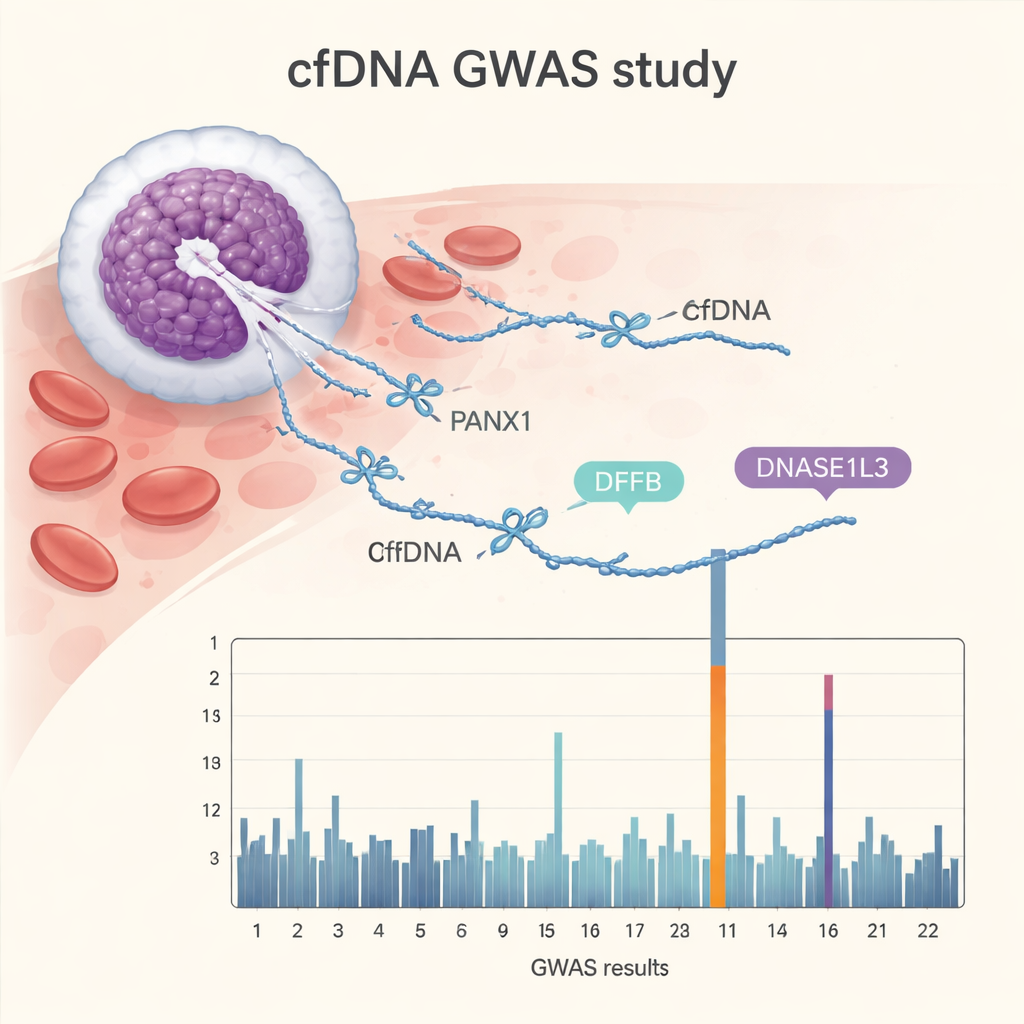
From Genes to Immune Cells and Body Traits
Genetic signals rarely act in isolation, so the team asked how cfDNA motifs relate to other traits measured in the same pregnant women. By comparing their cfDNA results with genome-wide scans of 104 clinical measures—such as blood counts, body weight, and liver function—they found shared genetic roots between certain end motifs and traits like body mass index, white blood cell counts, and especially neutrophil numbers. More detailed causal analyses suggested that changes in immune cells, rather than cfDNA itself, drive shifts in motif patterns. Additional pathway and tissue analyses pointed to a central role for blood and immune cells—particularly neutrophils, which can spew webs of DNA during immune responses—in producing and clearing cfDNA. Experimental work in mice and cultured cells in which PANX1 was switched off confirmed that this gene directly alters both the diversity of end motifs and the overall amount of cfDNA released.
Why This Matters for Future Medicine
To a non-specialist, the idea of tracking four-letter patterns at the tips of DNA fragments may sound esoteric. But this work shows that these patterns are not random noise: they are heritable, shaped by specific genes, tightly linked to immune cell behavior, and sensitive to body traits like weight. As millions of people worldwide have already had cfDNA sequencing for prenatal screening and cancer testing, the same data could be reanalyzed using cfGWAS to discover more genes that govern DNA fragmentation and clearance. In the long term, this knowledge could refine liquid biopsy tests, help distinguish which tissues are releasing abnormal DNA, and even suggest drug targets to either boost DNA clearance—as in autoimmune disease—or preserve fragile tumor DNA to improve early detection.
Citation: Zhu, H., Zhang, Y., Li, L. et al. cfGWAS reveal genetic basis of cell-free DNA end motifs. Nat Commun 17, 1714 (2026). https://doi.org/10.1038/s41467-025-67940-w
Keywords: cell-free DNA, genome-wide association, liquid biopsy, immune cells, pregnancy