Clear Sky Science · en
Optical super-resolution histology of formalin-fixed paraffin-embedded tissue samples: challenges and opportunities
Seeing More in Saved Tissue Samples
Hospitals around the world keep tiny slices of patients’ organs in wax blocks so that diseases like cancer or kidney damage can be studied for years to come. This review article explains how new kinds of light microscopes are turning those routine samples into powerful windows on disease, revealing details far too small for standard hospital microscopes to see. These advances could help doctors spot illnesses earlier, understand how they progress, and tailor treatments more precisely.
How Wax-Preserved Tissues Power Modern Medicine
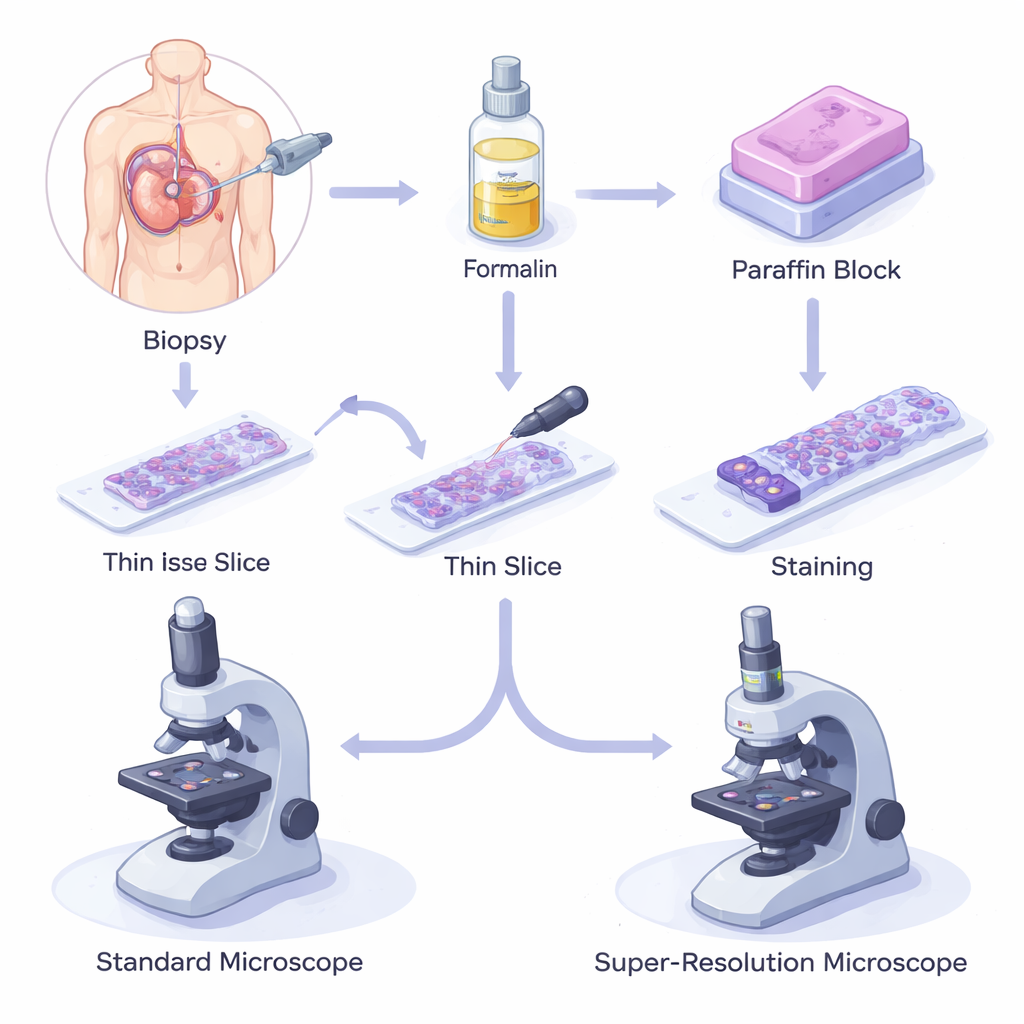
When a patient has a biopsy, a small piece of tissue is removed, fixed in a chemical called formalin to stop decay, and then embedded in paraffin wax. The block is sliced into hair-thin sections, placed on glass slides, stained, and examined under a light microscope. This formalin-fixed paraffin-embedded (FFPE) method is inexpensive, reliable, and can preserve samples for decades at room temperature. As a result, millions of FFPE samples are stored in biobanks worldwide, supporting everything from basic biology to advanced genetic and protein studies, and forming the backbone of modern diagnosis and prognosis.
Why Ordinary Microscopes Are No Longer Enough
Traditional optical microscopes are limited by the physics of light: details smaller than about 250 nanometers blur together. Many disease-related structures—such as the fine filters in kidneys, the connections between nerve cells, or tiny changes in the architecture of DNA inside the nucleus—fall below this limit. Today’s workflow often combines standard light microscopy for a broad overview with electron microscopy for nanometer details, but that approach is slow, expensive, and requires completely different sample preparation. Clinicians and researchers are therefore looking for a single, flexible system that can scan large areas quickly yet also zoom in to see nanoscale features in the same FFPE sections they already use.
New Ways of Sharpening the View
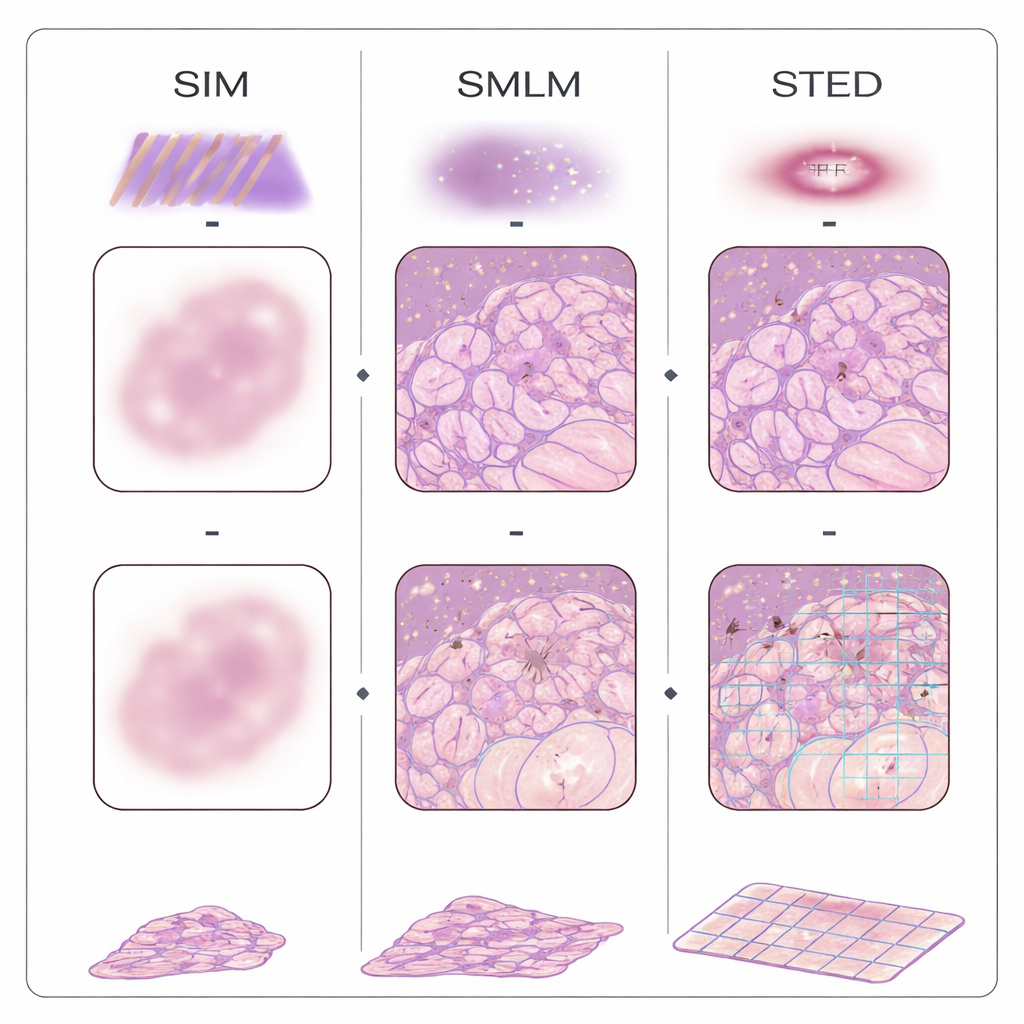
Over the past two decades, several families of “super‑resolution” microscopes have broken the traditional resolution barrier using fluorescent labels and clever optics. Single‑molecule localization methods switch molecules on and off and pinpoint their positions one by one; structured illumination projects striped light patterns onto the tissue and uses computation to recover finer details; stimulated emission depletion carves out a tiny glowing spot using a doughnut‑shaped beam; and fluctuation‑based approaches analyze subtle flickering in the image to infer structure. A different strategy, called expansion microscopy, avoids complex optics by physically swelling the tissue in a gel, so that ordinary microscopes can see formerly invisible detail. These methods have already been applied to FFPE samples from breast, colon, pancreas, kidney, brain, skin, placenta, and other organs, revealing how mitochondria change in tumors, how kidney filters fail, and how protein clumps form in Alzheimer’s disease.
Hurdles on the Road to Everyday Use
Despite their promise, these techniques are not yet ready to become routine tools in most pathology labs. Many super‑resolution systems are slow, cover only tiny fields of view, or require tens of thousands of images to build a single high‑detail picture, which is impractical when a pathologist must assess large tissue areas. FFPE tissues themselves are optically tricky: they scatter light, glow on their own, and can hide the very molecules that need to be labeled, all of which can cause blurry images or false structures. Some methods demand special dyes, custom buffers, or multi‑step chemical treatments that do not fit easily into standard histology workflows. On top of this, commercial instruments are costly and complex to operate, and the resulting data demand heavy computational processing and storage.
Bringing Smart Optics and Smart Software Together
To overcome these obstacles, researchers are combining improved optics with label‑free imaging and artificial intelligence. High‑throughput, diffraction‑limited methods such as Fourier ptychography and non‑linear optical microscopy can rapidly scan large, even unstained, FFPE sections and provide quantitative maps of tissue structure. Digital pathology platforms now capture entire slides at high resolution and use machine learning to detect tumors and score biomarkers like HER2, Ki‑67, and PD‑L1. Deep‑learning models can even turn low‑resolution images into super‑resolved views, potentially reducing the need for some specialized hardware while making better use of existing biopsy archives.
What This Means for Future Patients
The authors conclude that true “super‑resolution histology” of FFPE samples—combining nanometer‑scale detail, high speed, reasonable cost, and compatibility with current lab routines—has not yet been fully realized, but it is within reach. As optical methods become faster and more robust, and as AI‑based analysis matures, these tools could reveal subtle early changes in tissues that now go unnoticed, improve the accuracy of diagnoses, and guide more personalized treatment decisions. In the long run, turning every archived wax block into a high‑definition map of disease could transform both research and clinical care.
Citation: Villegas-Hernández, L.E., Dubey, V.K., Acharya, G. et al. Optical super-resolution histology of formalin-fixed paraffin-embedded tissue samples: challenges and opportunities. Nat Commun 16, 9760 (2025). https://doi.org/10.1038/s41467-025-64626-1
Keywords: super-resolution microscopy, FFPE tissue, digital pathology, optical imaging, cancer diagnostics