Clear Sky Science · en
A long road ahead to reliable and complete medicinal plant genomes
Why plant DNA maps matter for human health
Many of today’s most powerful medicines—from cancer drugs like paclitaxel to painkillers like morphine and the antimalarial artemisinin—come from plants. Yet for most medicinal plants, scientists still lack a complete “instruction manual” of their DNA. This review explains how new genome technologies are transforming our ability to read those manuals, why current plant genomes are often still incomplete or flawed, and how truly accurate genomes could unlock better drugs, more sustainable production, and improved conservation of valuable species.
The promise of reading medicinal plants’ blueprints
For millennia, people have relied on herbal remedies, and modern pharmacology continues to draw heavily from plant natural products. These specialized molecules—alkaloids, terpenoids, phenolic compounds and many others—are made through intricate metabolic pathways encoded in plant DNA. Until recently, scientists had to piece these pathways together using slow, labor‑intensive tools such as isotope tracing and one‑gene‑at‑a‑time cloning. The arrival of affordable, high‑throughput DNA sequencing changed the landscape. By February 2025, genomes for 431 medicinal plants (across 203 species) had been sequenced, giving researchers a systematic way to hunt for pathway genes, understand how valuable compounds are regulated, and explore how these chemistries evolved.
A boom in sequencing, but many imperfect genomes
Long‑read sequencing technologies from PacBio and Oxford Nanopore, paired with short‑read Illumina data and chromosome‑level mapping methods such as Hi‑C, have dramatically improved plant genome quality. Nearly half of all medicinal plant assemblies were released in just the past three years, and most recent genomes are now built at chromosome scale. However, the review shows that quantity has outpaced quality. Over half of the genomes exist only as an initial version, many remain draft‑level, and only 11 medicinal plants have “telomere‑to‑telomere” (T2T) gapless assemblies that fully capture centromeres and other repetitive regions. Standard metrics like N50 (a measure of contiguity) and BUSCO scores (a measure of conserved genes) look encouraging overall, but they can mask critical gaps precisely where key biosynthetic genes reside.
Hidden gaps where the medicine genes should be
To test how useful current genomes really are, the authors examined known, experimentally validated pathway genes in nine well‑studied medicinal plants. Even in some chromosome‑level assemblies, important enzymes for compounds such as ginsenosides in ginseng or artemisinin in Artemisia annua were either completely missing or only partially captured. In other cases, genes were present in the raw genome sequence but absent or truncated in the official gene annotations, making them hard to find. A striking example comes from the coumarin‑producing herb Peucedanum praeruptorum: an older chromosome‑level genome broke one key gene and missed two others; a newer T2T assembly not only restored these genes but also revealed that several of them sit together in a tightly packed biosynthetic gene cluster. This kind of cluster map is exactly what researchers need to engineer plants or microbes to produce medicines more efficiently.
Why plant genomes are so hard to assemble
Medicinal plants pose special challenges that go beyond those of many crop species. Their genomes often carry high levels of heterozygosity (many DNA differences between the two copies of each chromosome), frequent polyploidy (multiple chromosome sets), and large fractions of repetitive DNA—features that confuse assembly algorithms and lead to breaks or mis‑joins. About a third of sequenced medicinal plants have genomes with more than 70% repetitive content, and over a quarter show very high heterozygosity. Breeding highly inbred lines or isolating haploid tissue can help, but this is slow, expensive, or biologically difficult for many species. New strategies that assemble each parental haplotype separately, and more powerful algorithms tuned to repeat‑rich, polyploid genomes, are beginning to ease these hurdles but are not yet routine.
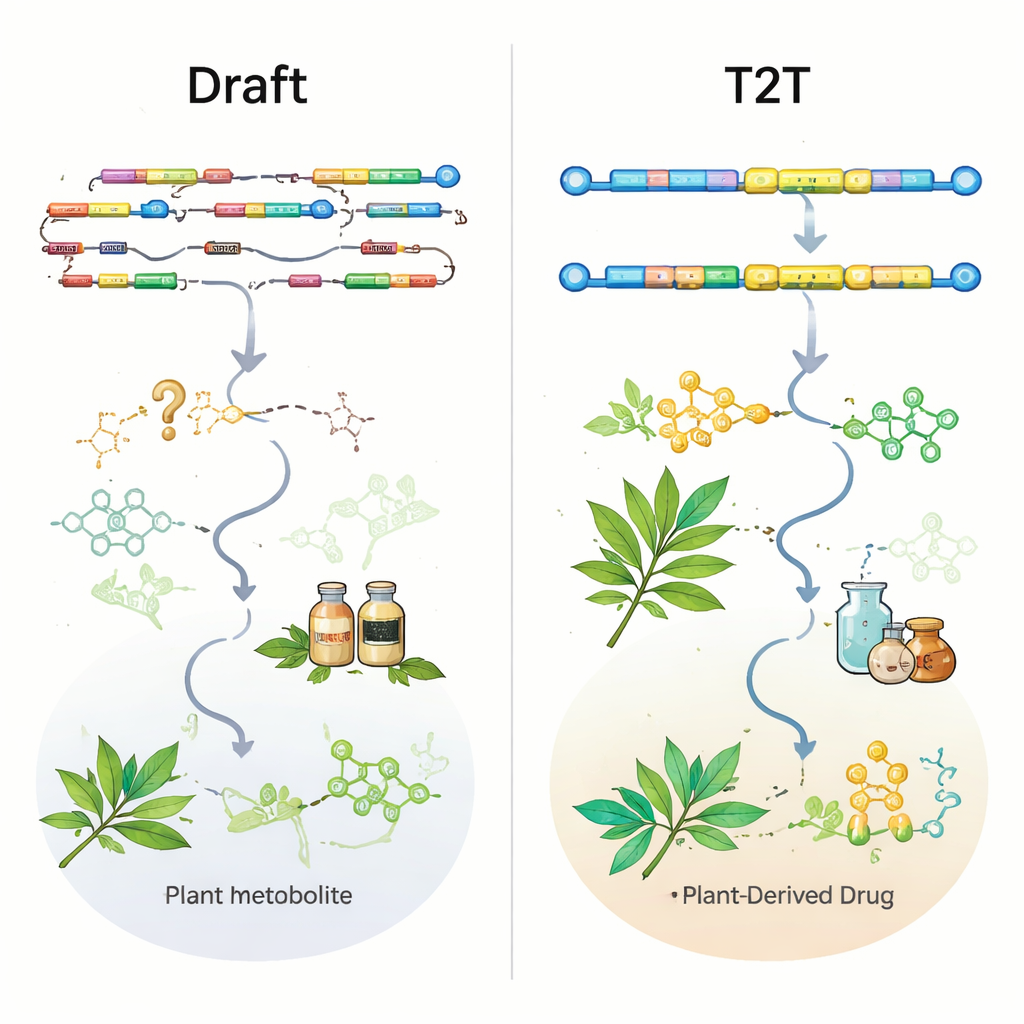
From genomes to new medicines and future directions
When genomes are good enough, they become powerful engines for discovery. Researchers can combine whole‑genome data with transcriptomics, metabolomics, and synthetic biology to pinpoint enzymes, regulatory genes, and biosynthetic gene clusters that control production of high‑value compounds. These insights have already enabled reconstruction of complex plant pathways—such as those for vinblastine, paclitaxel, and many other drugs—in yeast or model plants, opening a path toward stable, large‑scale biomanufacturing. Looking ahead, the authors argue for a shift from “one rough genome per species” to multiple, high‑quality, T2T and haplotype‑resolved assemblies that capture intraspecies diversity, much like pan‑genomes in crop research. Coupling these reference genomes with large‑scale resequencing, advanced phenotyping, and emerging single‑cell and spatial transcriptomics should illuminate how environment, cell type, and gene networks interact to shape medicinal chemistry.
What this means for patients and the planet
The review’s central message is that reliable, complete medicinal plant genomes are not a luxury; they are the foundation for turning centuries of herbal knowledge into precise, modern therapies. Better genomes will help scientists find missing steps in drug pathways, engineer safer and more abundant supplies of critical medicines, and identify alternative species that can produce the same compounds. They will also guide conservation and sustainable use of threatened medicinal plants, most of which still lack any genomic resource. In short, finishing the job of accurately mapping these genomes could speed drug discovery, stabilize supply chains, and preserve botanical diversity—all of which ultimately benefit human health.
Citation: Cheng, LT., Wang, ZL., Zhu, QH. et al. A long road ahead to reliable and complete medicinal plant genomes. Nat Commun 16, 2150 (2025). https://doi.org/10.1038/s41467-025-57448-8
Keywords: medicinal plant genomics, biosynthetic gene clusters, telomere-to-telomere genomes, natural product biosynthesis, synthetic biology